 |
 |
Abstract
An in vitro gas production technique was used in this study to elucidate the effect of two strains of active live yeast on methane (CH4) production in the large intestinal content of pigs to provide an insight to whether active live yeast could suppress CH4 production in the hindgut of pigs. Treatments used in this study include blank (no substrate and no live yeast cells), control (no live yeast cells) and yeast (YST) supplementation groups (supplemented with live yeast cells, YST1 or YST2). The yeast cultures contained 1.8×1010 cells per g, which were added at the rates of 0.2 mg and 0.4 mg per ml of the fermented inoculum. Large intestinal contents were collected from 2 Duroc×Landrace×Yorkshire pigs, mixed with a phosphate buffer (1:2), and incubated anaerobically at 39°C for 24 h using 500 mg substrate (dry matter (DM) basis). Total gas and CH4 production decreased (p<0.05) with supplementation of yeast. The methane production reduction potential (MRP) was calculated by assuming net methane concentration for the control as 100%. The MRP of yeast 2 was more than 25%. Compared with the control group, in vitro DM digestibility (IVDMD) and total volatile fatty acids (VFA) concentration increased (p<0.05) in 0.4 mg/ml YST1 and 0.2 mg/ml YST2 supplementation groups. Proportion of propionate, butyrate and valerate increased (p<0.05), but that of acetate decreased (p<0.05), which led to a decreased (p<0.05) acetate: propionate (A: P) ratio in the both YST2 treatments and the 0.4 mg/ml YST 1 supplementation groups. Hydrogen recovery decreased (p<0.05) with yeast supplementation. Quantity of methanogenic archaea per milliliter of inoculum decreased (p<0.05) with yeast supplementation after 24 h of incubation. Our results suggest that live yeast cells suppressed in vitro CH4 production when inoculated into the large intestinal contents of pigs and shifted the fermentation pattern to favor propionate production together with an increased population of acetogenic bacteria, both of which serve as a competitive pathway for the available H2 resulting in the reduction of methanogenic archaea.
With the growing concern over the use of antibiotics, feed additives, such as active dry yeasts, have been widely used in livestock industry to improve animal performance (van der Peet-Schwering et al., 2007; Desnoyers et al., 2009; Robinson and Erasmus, 2009) and to enhance their immunity (Podzorski et al., 1990; Davis et al., 2004). Active dry yeasts have also been reported to be effective in reducing methane (CH4) production through increasing the propionate concentration and homoacetogenic bacteria in ruminants (Mutsvangwa et al., 1992; Chaucheyras et al., 1995; Lila et al., 2004). Production of propionate serves as a competitive pathway for hydrogen (H2) in the rumen (Boadi et al., 2004) while homoacetogenic bacteria can utilize CO2 (carbon dioxide) and H2 to produce acetate, playing a major role in re-utilization of fermentative H2 in some non-ruminant digestive ecosystems (Bellier et al., 1996) and in gut of termites (Breznak and Switzer, 1986). Fermentation of undigested carbohydrates in the large intestine of pigs results predominantly in production of short chain fatty acids such as acetate, propionate and butyrate; gasses such as CO2, H2 and CH4 (Bach Knudsen and Jørgensen, 2001). From the energetic efficiency and environmental points of view, CH4 and H2 are important as they represent a loss of energy and are greenhouse gases that contributing to the global warming (Scheehle and Kruger, 2006). Although enteric CH4 production in pigs is much lower than that of ruminants, the large population of the former, particularly in China, warrants investigations to the possibility of reducing CH4 production from pigs. The objective of this study was to elucidate the effect of live yeasts on CH4 production in large intestinal content of pigs using in vitro production technique to provide an insight to whether active live yeast could suppress enteric CH4 production in the hindgut of pigs.
Two Duroc×Landrace×Yorkshire male finisher pigs (60±1 kg BW) were used as donor animals of the inoculums for this study. The animals were individually housed and fed a corn-soybean basal diet (Table 1) formulated to meet the nutritional requirements recommended by the National Research Council (NRC, 1998) for 50 to 80 kg pigs. Pigs were fed twice daily, at 0800 and 1600 h, and managed according to the protocols approved by the Animal Experimental Committee of Guangdong Institute of Animal Science. Clean drinking water is provided at all time.
After 4 wks of feeding, the pigs were slaughtered in the morning and the large intestinal content was collected as inoculum for the study. The contents were quantitatively transferred into plastic bags, sealed and taken to the laboratory within 2 h in a water bath pre-heated to 39°C. In the laboratory, the intestinal contents were mixed with sodium and ammonia bicarbonate buffer solution (35 g NaHCO3 plus 4 g NH4HCO3 per L) in a ratio of 1:3 (w/v) according to Ly et al. (1997). The intestinal content-buffer mixture was stirred for 60 s in a kitchen-blender after which the solution was squeezed through 4 layers of surgical gauze, and then mixed with the buffer mineral solution (Menke and Steingass, 1988) in a 1:2 ratio (v/v) at 39°C under continuous flushing with CO2.
The corn-soybean basal diet was used as substrate for the in vitro fermentation. The samples were ground to pass through 1-mm sieve, and 500±1 mg of the sample was weighed into calibrated glass syringes of 100-ml capacity (HÄBERLE, Germany). Two yeast samples; namely S. cerevisiae YST1, isolated from a commercial yeast supplement for pig production in China and S. cerevisiae YST2, isolated from bread were used as inocula for the in vitro gas production study.
The two strains of yeasts were isolated using plates of Yeast Peptone Dextrose (YPD) agar medium (Sigma, USA) from 2 different sources; the first from a commercial probiotic (Angela yeast Co., Guangzhou, China) sold as feed supplement to pig farms in China, and second from bread sample (purchased from bakery at Kuala Lumpur, Malaysia). Before placing them in the agar medium, they were diluted in saline solution (0.9% NaCl) in ratio of 1:100 w/v. Serial dilutions of the diluted samples (10−2 to 10−6) were prepared in sterilized saline solution. About 100 μl of samples from dilutions of 10−2 to 10−6 were transferred onto the YPD plates and incubated at 30°C for 72 h. At the end of incubation, 20 single colonies were selected from different dilutions of the commercial probiotic and bread inoculums and subcultured for 3 times in the YPD agar to get the pure culture samples. The isolated microorganisms were subcultured in YPD broth and incubated at 30°C for 48 h. After incubation, the samples were centrifuged at 9,600 g for 10 min, the supernatant was removed and the cells were washed by double distilled H2O (ddH2O) twice and centrifuged again. After centrifugation, the pellets were collected and froze into dry powder, which contained 1.8×1010 live organisms/g.
The harvested cells from 4 ml culture of each isolates were used for 18S rRNA identification. DNA was extracted from each yeast cells using the i-genomic BYF DNA Extraction Mini Kit (iNtRON biotechnology, Inc) according the manufacturer procedure. The primers of ITS 1 (5′-TCC GTA GGT GAA CCT TGC GG-3)′ and ITS 2 (5′-GCT GCG TTC TTC TTG ATC GAT GC-3′) were used for amplification of 18S rRNA from isolated samples (White et al., 1990). The PCR reaction system contained 2 μl of DNA extract from each sample, 2.0 μl of each prime, 4.0 μl of dNTPs, 1.0 μl of EX Taq, 5.0 μl of 10×buffer (Promega, Madison, WI), and 34.0 μl of ddH2O. The PCR conditions were 35 cycles of denaturing at 95°C for 1 min; annealing at 60°C for 45 s and extension at 72°C for 45 s; and final extension at 72°C for 10 min. PCR products (50 μl) were further purified using the E.Z.N.A Gel Extraction kit (omega, USA). The purified PCR products were sent for sequencing. The sequences were compared to the GenBank database over the Internet by using the NCBI BLAST server (http://blast.ncbi.nlm.nih.gov/) NCBI blast online system was used for identification study.
In vitro gas production was performed as described by Menke and Steingass (1988) and adapted for use in pigs by Ly et al. (1997). About 30 ml of the inoculum was added to each 100-ml glass syringe (HÄBERLE, Germany) with or without 0.5 g of substrate. The YST was supplemented at 0.2 and 0.4 mg per ml of fermented inoculum. The syringe was gently shaken and pushing the piston to remove air bubbles in the syringe before closing the plastic clip on the silicon tube attached to the tip of the syringe. The position of the piston of the lubricated syringes was recorded and the syringes were incubated at 39°C for 24 h, shaking at 60 g. According to the change in the piston position, the volume of accumulated gas was recorded at 2 h intervals.
Two 24-h identical incubation runs were carried out independently in two fermenters, which received same treatments, with 3 replicates per treatment in each run. The experimental treatments were as follows: blank (containing no substrate, but only the inoculum), control (containing the substrate and inoculum) and YST supplementation groups (containing substrate and inoculum, and mixed with 2 strains of Saccharomyces cerevisiae live cells, YST1 and YST2 respectively, at the rate of 0.2 and 0.4 mg per ml fermented inoculum). The lower YST inclusion rate was estimated based on the recommendation of the manufacturer (Angela Yeast Inc.) that 100 g of S. cerevisiae (YST1) was to add to per 100 kg of feed and we further assumed that the volume of the large intestine of a finisher pig is 10 L, and the average daily feed intake of 2 kg per head. From the above assumption, the estimated concentration for the live yeast in the large intestine would be approximately 0.2 mg/ml.
At the end of the fermentation, about 50 ml gas samples was removed from each syringe using a gas tight syringe (Hamilton, Reno, NV, USA) and transferred into a gas-sampling bag for CH4 analysis. Contents of the syringe were transferred separately to previously weighted tubes and put in icebox to stop further fermentation. The tubes were centrifuged at 3,500×g for 10 min and the supernatant were collected in Eppendorf tubes and stored at −80°C for subsequent volatile fatty acids (VFA) analysis and bacterial quantification. The residue of incubation was washed with 30 ml of distilled water twice and dried at 60°C till constant weight and weighed to calculate in vitro dry matter disappearance (IVDMD).
Concentration of CH4 was determined by injection of 500-μl of the gas collected from each gas-sampling bag into a gas chromatograph (GC-2010, Shimadzu, Kyoto, Japan) equipped with a HP-Plot Q column (30 m×0.32 mm×0.25 μm) (Agilent Technologies, Wilmington, DE, USA). Temperatures of the injector oven, column oven and detector were 180, 50 and 200°C, respectively, and the flow rates for nitrogen, H2 and air were 50, 40 and 400 ml/min, respectively. Peaks were identified by comparison with a known concentration of pure CH4 (50.10 ml/L).
For VFA concentration analysis, 1 ml of the supernatant was collected into Eppendorf tubes after 24 h of incubation and 200 μl of 25% methaphosphoric acid was added to acidify the samples. The samples were kept at room temperature for 24 h and centrifuged at 5,000×g for 10 min. After centrifugation, 1 μl of the supernatant was injected into the previously mentioned gas chromatograph equipped with a flame ionization detector (FID) and DB-FFAP column (30 m×0.32 mm×0.25 μm) (Agilent Technologies, Wilmington, DE, USA). Temperatures of the injector oven and detector were 220 and 250°C, respectively. Temperatures of the column oven was initially kept at 60°C for 2 min, and then heated up to 150°C at a rate of 30°C/min and kept for 2.5 min and finally heated up to 250°C at a rate of 40°C/min. The gas flow rates for nitrogen, H2 and air were 50, 40 and 400 ml/min, respectively.
To examine the effect of S. cerevisiae live cells on the number of methanogenic archaea, 1.5 ml of the supernatant was used for microbial quantification by real-time quantitative PCR. Whole genomic community DNA was extracted using the QIAamp DNA Stool Mini Kit (Qiagen Inc., Valencia, CA, USA) according to the manufacturer’s protocol. Integrity of the extracted DNA was verified by electrophoresis on a 0.7% agarose gel stained with ethidium bromide, and DNA in samples was quantified using ultraviolet-clear microplates (Corning, NY, USA) at an optical density of 260 and 280 nm. The OD260/OD280 ratio of all samples was 1.90. The extracted DNA was stored at −20°C and used as a template for real-time quantitative PCR. Real-time PCR was performed with the BioRad CFX96 Touch (BioRad, USA) using optical grade plates. Paired primers LuF/LuR (amplify a 464 bp gene fragment) targeting specific to the mcrA gene of methanogenic archaea was designed as 5′-GGTGGTGTMGGATTCACACARTAYGCWACAGC-3′ (forward); 5′-TTCATTGCRTAGTTWGGRTAGTT-3′ (reverse). The qPCR reaction was performed in a total volume of 20 μl with SYBR Green I on an i-Cycler iQ (Bio-Rad, Hercules, CA, USA). Final reaction mixtures contained 10 μl SYBR Green Supermix, 1 μl of each Primer, 3 μl of DNA samples and 5 μl H2O. Cycle conditions were conducted following a program of 1 min at 95°C, 15 s at 95°C, 30 s at 58°C, and 25 s at 72°C for 35 cycles followed by 10 min at 72°C. Standard curves were constructed using appropriate serial dilution of the Plasmid (PMD-18T, Takara, Japan) DNA. The copy numbers of standard plasmid and per ml of elution buffer (copy/ml) and copy numbers of target gene per ml of incubate sample were determined by standard method (Ritalahti et al., 2006).
The total gas produced in the control and the YST treatment groups was calculated by subtracting the average gas produced in blank syringes. The percent loss in weight of dry matter (DM) of the substrate was presented as IVDMD. The volume of CH4 produced (ml) was calculated by multiplying gas produced (ml) by its corresponding CH4 concentration in the analyzed sample. Methane production reduction potential (MRP) was calculated by taking net methane values for the control (no yeast supplemented) as 100%:
The molar proportion of acetate, propionate and butyrate (mmol/100 mmol) was calculated as the determined concentration of VFA in samples divided by the determined total VFA concentration and then multiply 100. Recovery of H2 was estimated from the molar proportion of VFA and CH4 (Demeyer and Degraeve, 1991) as shown below:
Where M is methane, A is acetate, P is propionate, B is butyrate and V is valerate.
Data were analyzed by one-way analysis of variance (ANOVA) using the general linear model procedures of the SPSS 17.0 Program (Chicago, IL, USA). The syringe was considered as the experimental unit. The Tukey’s tests were performed to identify differences among group means. The correlation between VFA and methane production was also analyzed. A p values≤0.05 were considered statistically significant, 0.05<p values≤0.1 were considered tendencies.
Analysis of the partial 18s rRNA sequences (411-bp) in the selected 2 yeast strains (one isolated from a commercial yeast product sold as feed supplement for pigs and another was isolated from bread) showed a high homology of 90%. Pylogenetic analysis between the two strains of active dry yeast on the basis of the ClustalW method in Lasergene software (DNASTAR) are shown in Figure 1. Phylogenetic analysis indicated that the two strains were located in the same branch with Saccharomyces cerevisiae (GU256758.1) indicating that the two yeast strains used in our study belong to Saccharomyces cerevisiae.
Effects of active dry yeast on production (ml) of total gas and CH4 and IVDMD (%) after 24 h of incubation are shown in Table 2. Total gas and CH4 production decreased (p<0.05) by the addition of yeast. CH4/total gas ratio for YST 2 treatments was lower (p<0.05) than the control. The MRP of YST 1 and YST 2 were 10.1 and 26.2 (average value of 2 supplemental level). IVDMD increased (p<0.05) in higher level of YST1 and lower level of YST2 treatments.
Propionate, butyrate, valerate, and total VFA concentrations in the 0.4 mg/ml YST 1- and 0.2 mg/ml YST 2-supplementation groups were significantly higher (p<0.05) than the control (Table 3). Concentration of acetate tended (p = 0.087) to be lower with addition of yeasts. Proportion of propionate, butyrate, and valerate increased (p<0.05), but that of acetate decreased (p<0.05), which led to a decrease (p<0.05) in the A: P (acetate: propionate) ratio in the 0.4 mg/ml YST 1 and both the YST 2 supplementation groups. Hydrogen recovery decreased (p<0.05) by yeasts supplementation.
A significantly positive correlation (p<0.05) was found between CH4 production and molar proportion of acetate and the A:P ratio (Table 4). The proportion of propionate tended (p = 0.073) to be negatively correlated with CH4 production.
The standard curve for quantitative PCR was obtained by preparing 10-fold dilutions of plasmid containing 464-bp partial mcrA gene fragment amplified with LuF/LuR primer set and it had a linear scope of detection ranged from 103 to 109 target molecule numbers, with a slop of −3.491 and amplification efficiency of 94.3%. Effects of yeasts on mean numbers of mcrA genes (methanogenic archaea) in inoculum of each treatment are presented in Figure 2. DNA extracted from the fermented inoculum had to be diluted 10 folds in order to minimize PCR inhibition in our study. In all samples, the absolute abundance of total methanogenic archaea was in the magnitude of 106 to 108 per ml of ferment inoculum. The absolute abundance of total methanogenic archaea was significantly lower (p<0.01) in yeast treatment groups than in the control.
In the present study, the 2 strains of Saccharomyces cerevisiae live cells, selected based on their growth rate alone, showed potential to suppress CH4 production in substrate fermented in large intestinal contents of pigs as inoculum (Table 2). This result is in agreement with a previous study that Saccharomyces cerevisiae reduces ruminal CH4 production (Lynch and Martin, 2002), but others found no effect (Lila et al., 2004) or an increase (Martin et al., 1989) in batch cultures with mixed rumen microflora. The discrepancies among studies could be associated with the characteristics of the strain (Chuang et al., 2011), diet composition (Sullivan and Martin, 1999) and dose (Lila et al., 2006).
A significant decrease in total gas production in the yeast treatments was observed in this study (Table 2), which was increased in previous studies (Lila et al., 2004; Lila et al., 2006). This may be partly associated with the decreased production of acetate in the yeast treatments, because CO2 and H2 are byproducts of acetate production during carbohydrate fermentation. Methane production was decreased by yeast supplementation, and the result of MRP parameter also reflects the total methane production reduction potential of the yeast (Table 2). The lower CH4/total gas values in the yeast treatment groups suggest the reduction in CH4 was absolute and not a result of a decrease in the total gas. The increased IVDMD in the higher level of YST 1- and lower level of YST 2-supplementation groups (Table 2) is in agreement with results of Ayala et al. (1992), who found an improvement in digestibility with yeast supplementation because of the improved NDF degradability. Other in vivo studies showed that some yeast cultures increased the number of cellulolytic bacteria in the rumen and, in some cases, increased cellulose degradation (Dawson, 1990; Newbold, 1995).
The increased total VFA concentrations and the greater molar proportion of propionate in the higher level of YST1 and lower level of YST2 treatments were in accordance with the increased substrate degradability (Table 3). The significantly lower A:P and A+B:P ratios in the yeast supplementation groups than the control seem to suggest one possible mechanism of how yeast suppresses CH4 production is by shifting the fermentation product to higher propionate and lower acetate and butyrate productions. CO2 and H2 that served as the main substrates for CH4 synthesis are generated when carbohydrate was fermented to acetate and butyrate (Flint, 1997), while production of propionate serves as a competitive pathway for available H2 (Boadi et al., 2004). The positive correlation between acetate (and butyrate) and CH4 production together with the negative correlation between propionate and CH4 production (Table 4) recorded in our study are in agreement with the above suggestions and further support our hypothesis that the YST suppressed CH4 production by shifting the fermentation pathway to favor propionate production.
Several in vitro studies have shown the beneficial effects of supplementing live yeasts on H2 utilization and acetate production by homoacetogenic bacteria, even in the presence of methanogens (Chaucheyras, et al., 1995a; Chaucheyras-Durand et al., 1997). When methane production is decreased, hydrogen concentration is often increased (Czerkawski, 1986). Homoacetogenic bacteria can utilize CO2 and H2 to produce acetate, which has been reported to play an important role in re-utilization of fermentative H2 in some non-ruminant digestive ecosystems (Bellier and Gidenne, 1996). The presence of homoacetogenic bacteria in pigs has been reported by DeGraeve and Demeyer et al. (1990). Besides, hydrogen sulfide is produced in the large intestine by a dissimilatory sulfate reduction, which has higher affinity for hydrogen and a lower hydrogen threshold than methanogenesis if a sufficient amount of sulfate is supplied (Zinder, 1993). Although we did not measure the sulfate concentration in the digesta but around 10 mM free sulfate was detected in the pig cecal digesta (Ushida et al., 1995), which is assumed to be sufficient for sulfate reduction. Hydrogen recovery value in this study was only between 58 to 65%, which might be because part of the H2 was used for hydrogen sulfide production. Low H2 recovery rate indicates the presence of reductive acetogenesis as a substantial source of acetate in the hindgut of rabbit (Demeyer and Degraeve, 1991). The H2 recovery decreased significantly by yeasts in this study (Table 3), together with the increased acetate production and the lower CH4 yield in yeast supplementation groups seem to suggest the possibility that YST induces higher acetogenesis.
A novel finding of our study is to quantitatively show a reduction of the total methanogenic archaea population in the yeast-supplemented groups (Figure 2). We proposed that the reduction of methanogenic archaea is because YST supplementation shifted the fermentation pattern to enhance propionate production and induced acetogenesis, both of which competes for the H2 available for methanogenic archaea.
In conclusion, the 2 strains of active live yeasts tested in this study showed to be potential agents to mitigate enteric CH4 production. Compared to the control group, the higher level of yeast 1 and the lower level of yeast 2 also improved DM digestibility, total VFA and propionate production and decreased the number of methanogenic archaea. Our results show that live yeast cells mitigate CH4 by modifying the fermentation pathways toward to favor propionate over acetate and butyrate productions as well as increasing homoacetogenesis, thereby sinking the available H2 thus suppressing growth of methanogenic archaea.
ACKNOWLEDGMENTS
This study was financially supported by a grant from the National Science and Technology Major Project of the Ministry of Science and Technology of China (20011ZX08006004).
Figure 1.
ITS2 sequence-based phylogenetic tree of yeast isolates. Neighbor-joining dendrogram with 5,000 bootstraps was based on aligned positions of ITS2 sequences of isolated yeasts (YST1, YST2 and YST3) and adjacent partial sequences of rRNA genes from 18 yeast isolates obtained from Genbank. Accession numbers of isolates were shown in front of them. Branches corresponding to partitions reproduced in less than 50% bootstrap replicates are collapsed. The evolutionary distances were computed using the Kimura 2-parameter method. The analysis involved 22 nucleotide sequences. Encephalitozoon cuniculi was used as out-group isolate. Phylogenetic tree was designed by MEGA5 software.
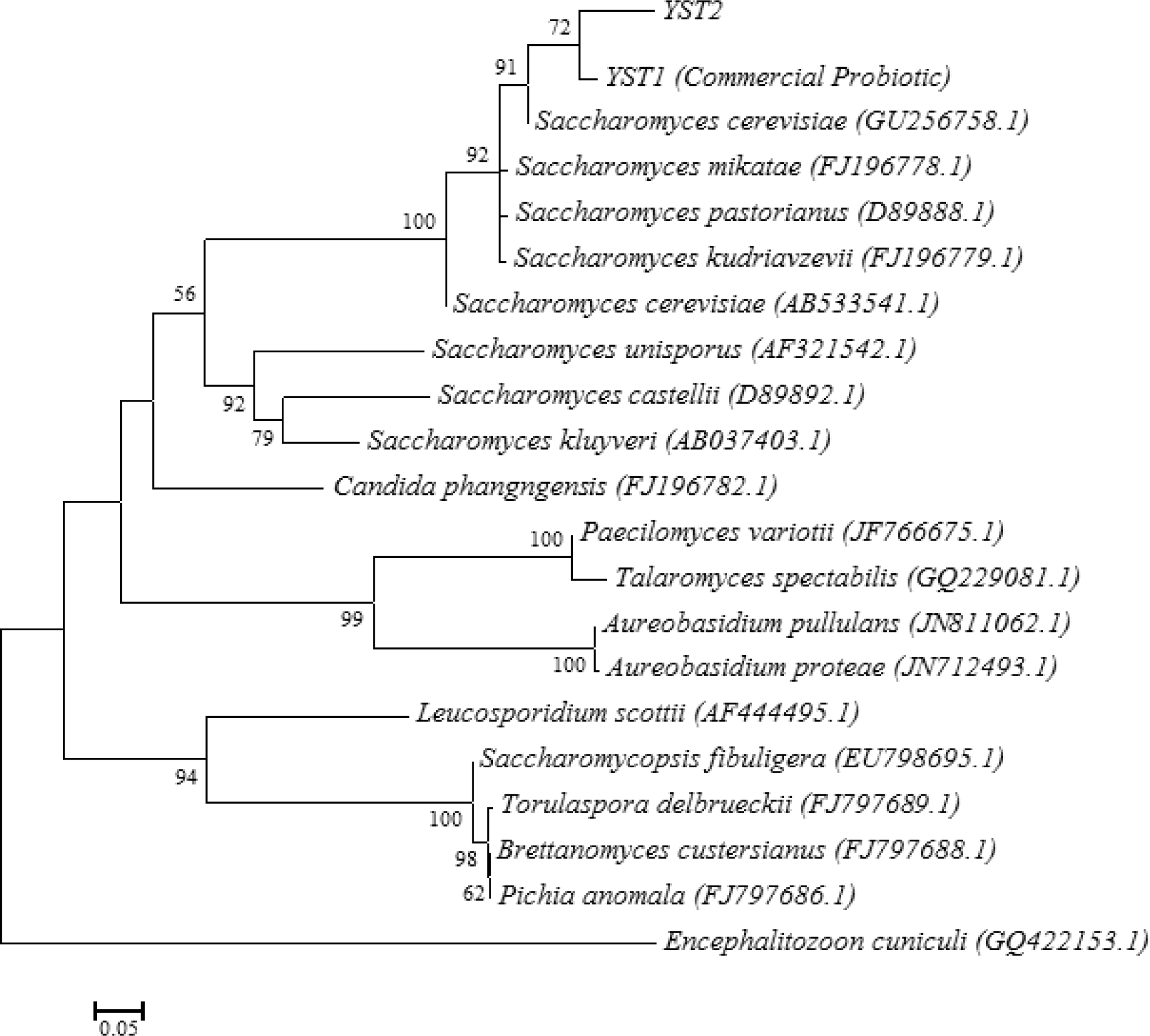
Figure 2.
Mean numbers of mcrA genes (methanogenic archaea) in inocula. Error bars represent standard errors of means. a,b Treatments with different letters are different at p<0.05.
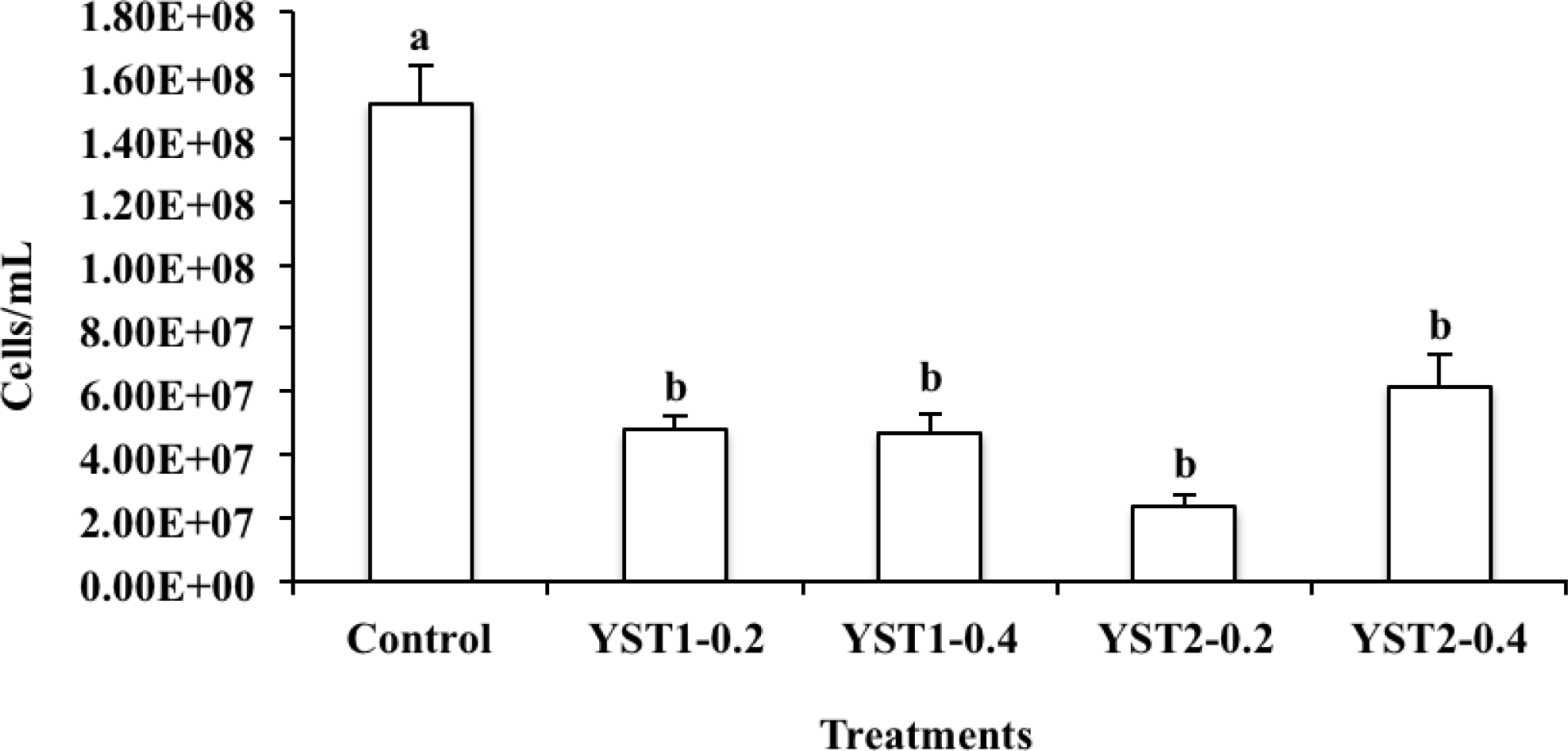
Table 1.
Ingredient composition and nutrient content of the basal diet
| Item | |
|---|---|
| Ingredient (% as-fed) | |
| Corn | 70 |
| Soybean meal | 20 |
| Wheat bran | 7 |
| Powder | 1 |
| CaHPO3 | 0.52 |
| Salt | 0.4 |
| L-lysine-HCl (98%) | 0.08 |
| Premixa | 1 |
| Nutrient content (%)b | |
| Gross energy (Mcal/kg) | 3.2 |
| Crude protein | 15 |
| Crude fiber | 2.97 |
| Ether extract | 3.10 |
| Calcium | 71 |
| Available phosphorus | 20.1 |
| Lysine | 0.77 |
| Methionine+cystine | 0.47 |
a Provided per kg of diet: 3,400 IU; vitamin A, 1,200 IU; vitamin D3, 12 IU; vitamin E, 2.5 mg; vitamin K3, 1.0 mg; vitamin B1, 4.0 mg; vitamin B2, 2.4 mg; vitamin B6, 0.015 mg; vitamin B12, 35 mg; niacin, 16 mg; calcium pantothenate, 0.5 mg; folic acid, 0.05 mg; biotic, 40 mg; manganese, 50 mg; iron, 75 mg; zinc, 3.5 mg; copper, 0.14 mg; iodine, 0.15 mg; selenium, 0.15 mg.
Table 2.
Effects of two strains of Saccharomyces cerevisiae on total gas and methane production and in vitro dry matter disappearance
Table 3.
Effects of two strains of Saccharomyces cerevisiae on concentrations and molar proportion of VFA, and the calculated hydrogen release and recovery
REFERENCES
Ayala OJ, GonzaÂlez SS, Herrera R, Barcena R, Mendoza GD. 1992. Effect of a probiotic and a molasses-urea supplement on fiber digestibility of sesame straw. J. Anim. Sci 70:307
Bellier R, Gidenne T. 1996. Consequences of reduced fibre intake on digestion, rate of passage and caecal microbial activity in the young rabbit. Br J Nutr 75:353–363.


Breznak JA, Switzer JM. 1986. Acetate synthesis from H2 plus CO2 by termite gut microbes. Appl Environ Microbiol 52:623–630.



Boadi D, Benchaar C, Chiquette J, Massé D. 2004. Mitigation strategies to reduce enteric methane emissions from dairy cows: Update review. Can J Anim Sci 84:319–335.

Chaucheyras F, Fonty G, Bertin G, Gouet P. 1995. In vitro H2 utilization by a ruminal acetogenic bacterium cultivated alone or in association with an archaea methanogen is stimulated by a probiotic strain of Saccharomyces cerevisiae. Appl Environ Microbiol 61:3466–3467.



Chaucheyras-Durand F, Fonty G, Bertin G. 1997. Effects of a microbial additive, Levucell SC on growth and metabolism of a ruminal acetogenic bacterial strain in vitro. In : Proceedings of Rumen Function Conference; Chicago, USA. p. 33
Chuang YH, Walker ND, McGinn SM, Beauchemin KA. 2011. Differing effects of 2 active dried yeast (Saccharomyces cerevisiae) strains on ruminal acidosis and methane production in nonlactating dairy cows. J Dairy Sci 94:2431–2439.


Czerkawski WJ. 1986. An introduction to rumen studies. Oxford, UK: Pergamon Press.
Davis ME, Brown DC, Maxwell CV, Johnson ZB, Kegley EB, Dvorak RA. 2004. Effect of phosphorylated mannans and pharmacological additions of zinc oxide on growth and immunocompetence of weaning pigs. J Anim Sci 82:581–587.


Dawson KA. 1990. Designing the yeast culture of tomorrow-mode of action of yeast culture for ruminants and non-ruminants. Biotechnology in the Feed Industry. In : Proc. Alltech’s 6th Annu. Symp; Lexington, KY. Alltech Tech. Publ; Nicholasville, KY: p. 59
Demeyer DI, De Graeve K. 1991. Differences in stoichiometry between rumen and hindgut fermentation. J Anim Physiol Anim Nutr 22:50–61.
Desnoyers M, Giger-Reverdin S, Bertin G, Duvaux-Ponter C, Sauvant D. 2009. Meta-analysis of the influence of Saccharomyces cerevisiae supplementation on ruminal parameters and milk production of ruminants. J Dairy Sci 92:1620–1632.


Flint HJ. 1997. The rumen microbial ecosystem- some recent developments. Trends Microbiol 5:483–488.


Knudsen KE, Jørgensen H. 2001. Intestinal degradation of dietary carbohydrates-from birth to maturity. Lindberg JE, Ogle B, editorsDigestive Physiology of Pigs. CABI Publishing; Wallingford: p. 109–120.

Lila ZA, Mohammed N, Yasui T, Kurokawa Y, Kanda S, Itabashi H. 2004. Effects of a twin strain of Saccharomyces cerevisiae live cells on mixed ruminal microorganism fermentation in vitro. J Anim Sci 82:1847–1854.


Lila ZA, Mohammed N, Takahashi T, Tabata M, Yasui T, Kurihara M, Kanda S, Itabashi H. 2006. Increase of ruminal fiber digestion by cellobiose and a twin strain of Saccharomyces cerevisiae live cells in vitro. J Anim Sci 77:407–413.

Ly J, Lai NV, Rodriguez L, Preston TR. 1997. In vitro gas production and washing losses of tropical crop residues for ruminants and pigs. Livest Res Rural Dev 9:26–37.
Lynch HA, Martin SA. 2002. Effects of Saccharomyces cerevisiae culture and Saccharomyces cerevisiae live cells on in vitro mixed ruminal microorganism fermentation. J Dairy Sci 85:2603–2608.


Martin SA, Nisbet DJ, Dean RG. 1989. Influence of a commercial yeast supplement on the in vitro ruminal fermentation. Nutr Rep Int 40:395–403.
Menke KH, Steingass H. 1988. Estimation of the energetic feed value obtained from chemical analysis and in vitro gas production using rumen fluid. J Anim Res Dev 28:7–55.
Mutsvangwa T, Edwards IE, Topps JH, Paterson GFM. 1992. The effect of dietary inclusion of yeast culture (Yea-Sacc) on patterns of rumen fermentation, food intake and growth of intensively fed bulls. Anim Prod 55:35–40.

National Research Council. 1998. Nutrient Requirements of Swine, 10th. Washington DC: National Academy Press.
Newbold CJ. 1995. Microbial feed additives for ruminants. Biotechnology in Animal Feeds and Animal Feeding. Wallace RJ, Chesson A, editorsVCH; Weinheim, Germany: p. 259–278.

Podzorski RP, Gray GR, Nelson RD. 1990. Different effects of native Candida albicans mannan and mannan-derived oligosaccharides on antigen-stimulated lymphoproliferation in vitro. J Immunol 144:707–716.



Ritalahti KM, Amos BK, Sung Y, Wu Q, Koenigsberg SS, Loffler FE. 2006. Quantitative PCR targeting 16s rRNA and reductive dehalogenase genes simultaneously monitors multiple dehalococcoides strains. Appl Environ Microbiol 72:2765–2774.



Robinson PH, Erasmus LJ. 2009. Effects of analyzable diet components on responses of lactating dairy cows to Saccharomyces cerevisiae based yeast products: A systematic review of the literature. Anim Feed Sci Technol 149:185–198.

Scheehle EA, Kruger D. 2006. Global anthropogenic methane and nitrous oxide emissions. Energy J 3:33–34.

Sullivan HM, Martin SA. 1999. Effects of Saccharomyces cerevisiae culture on in vitro mixed ruminal microorganism fermentation. J Dairy Sci 82:2011–2016.


Ushida K, Ohashi Y, Tokura M, Miyazaki K, Kojima Y. 1995. Sulfate reduction and methanogenesis in the ovine rumen and porcine caecum: a comparison of two microbial ecosystems. Dtsch Tierarztl Wochenschr 102:154–156.

Van der Peet-Schwering CM, Jansman AJ, Smidt H, Yoon I. 2007. Effects of yeast culture on performance, gut integrity, and blood cell composition of weaning pigs. J Anim Sci 85:3099–3109.







 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print





