 |
 |
| Anim Biosci > Volume 32(2); 2019 > Article |
|
Abstract
Methods
Representative rumen contents were obtained from three ruminally cannulated Holstein steers (793±8 kg). Pre-reduced media were used for the growth and isolation of methanogens. Optimum growth temperature, pH, and sodium chloride (NaCl) concentration as well as substrate utilization and antibiotic tolerance were investigated to determine the physiological characteristics of the isolated strain. Furthermore, the isolate was microscopically studied for its morphology. Polymerase chain reaction of 16S rRNA and mcrA gene-based amplicons was used for identification.
Results
One strain designated as KOR-2 was isolated and found to be a non-motile irregular coccus with a diameter of 0.2 to 0.5 μm. KOR-2 utilized H2+CO2 and formate but was unable to metabolize acetate, methanol, trimethylamine, 2-propanol, and isobutanol for growth and methane production. The optimum temperature and pH for the growth of KOR-2 were 38°C and 6.8 to 7.0, respectively, while the optimum NaCl concentration essential for KOR-2 growth was 1.0% (w/v). KOR-2 tolerated ampicillin, penicillin G, kanamycin, spectromycin, and tetracycline. In contrast, the cell growth was inhibited by chloramphenicol. Phylogenetic analysis of 16S rRNA and mcrA genes revealed the relatedness between KOR-2 and Methanoculleus bourgensis.
In the past decade, ruminal methanogens have attracted much research interest to mitigate ruminal methane (CH4) emission, as rumen CH4 emission accounts for about 17% of the global CH4 emission [1]. In addition, about 2% to 12% of the ingested feed energy is lost in the form of CH4 [2]. Studies related to ruminal methanogens are directed to understand their diversity and community structure, relationship with other ruminal microbes and feed efficiency, CH4 emission, and responses to dietary interventions.
The rumen provides a unique environment characterized by a relatively rapid passage rate and readily available carbon dioxide (CO2) and hydrogen (H2). This environment, therefore, facilitates the assembly of a community of archaea and is different from other anoxic habitats. Methanogens are the most dominant archaea in the rumen, and most of them are hydrogenotrophic rather than acetoclastic in spite of the high concentrations of acetates in the rumen [3]. H2 and CO2 produced from the fermentative pathways of other ruminal microbes are scavenged by rumen methanogens that also utilize formic acid and methyl-amines as substrates [4]. The interspecies H2 transfer in the rumen ecosystem prevents H2 accumulation and feedback inhibition. Most methanogens live freely in the rumen fluid or as members of the biofilm adhering to feed particles, whereas small portions of the ruminal methanogens are symbionts and may be either ectosymbionts or endosymbionts [5].
Given that methanogens are difficult to study through culture-based methods, many researchers have instead used culture-independent techniques such as real-time polymerase chain reaction (qPCR), denaturing gradient gel electrophoresis, and sequencing approaches, all of which have been valuable tools for the study of the biodiversity of complex microbial communities such as those in the rumen [6,7]. The diversity in the rumen methanogens is much smaller than that of rumen bacteria, and methanogens account for only 6.8% of total ruminal small subunit ribosomal ribonucleic acid (SSU rRNA) [8]. Archaea represent <3.3% of the total rRNA (both 16S and 18S) in the rumen [3]. The 16S rRNA gene sequences from cultured methanogens only account for approximately 0.7% of the total archaeal sequences of rumen origin, and several taxa have no single cultured representative [3]. In comparison with other anaerobic habitats from where methanogens have been isolated and classified into 28 genera and more than 100 species [9], the diversity and species richness of ruminal methanogens are quite low, reflecting the highly selective ruminal environment for methanogens [10]. To date, only 10 species of ruminal methanogens have been isolated as pure cultures [11,12]. Methanobacterium formicicum, Methanobacterium bryantii, Methanobrevibacter ruminantium, Methanobrevibacter millerae, Methanobrevibacter olleyae, Methanomicrobium mobile, Methanoculleus olentangyi (a later heterotypic synonym of M. bourgensis), Methanosarcina barkery, Methanobrevibacter boviskoreani, Methanobacterium beijingense, M. marisnigri, and Methanosarcina mazei (based on the RDP database). Here, we have isolated and studied the characteristics of a strain of M. bourgensis from the rumen of Holstein steers in Korea.
This study was approved by the Institutional Animal Care and Use Committee at the Chung-Ang University, Seoul, Korea (No. 2013-0047).
Representative rumen contents were obtained from three ruminally cannulated Holstein steers (793±8 kg) 2 h after morning feeding. Holstein steers were offered typified commercial concentrates (Table 1) and rice straw at a ratio of 40:60. The steers had a free access to the diet and water. Fresh rumen contents (900 mL) were collected into bottles previously kept warm and filled with O2 free-CO2 gas and then filtered through four layers of gauze. The filtered rumen fluid was used as an inoculum to isolate methanogens. The concentrates were oven-dried at 60°C for 3 days, milled to pass through a 1-mm sieve, and analyzed for chemical composition using the appropriate AOAC [13] and Van Soest method [14].
Methanogens are extremely sensitive to oxygen and need strict anoxic conditions; therefore, pre-reduced media are essential for their growth and isolation [6,7]. The methods used for the preparation of media and substrate solutions and culture techniques were those described by Hungate [15], as modified by Balch et al [16]. For enrichment culture and isolation, the medium was prepared based on the modification by Sowers and Schreier [17] and contained the following compounds (per liter distilled water): KH2PO4, 0.51 g; K2HPO4, 0.24 g; MgSO4·7H2O, 0.204 g; (NH4)2SO4, 0.255 g; NaCl, 1.02 g; CaCl2·2H2O, 0.134 g; vitamin solution, 10.0 mL; yeast extract, 1.0 g; sodium acetate, 50 mM; methanol, 50 mM; sodium formate, 50 mM; rumen fluid, 300 mL; NaHCO3, 5.0 g; resazurin, 0.001 g; 0.2 M cysteine hydrochloride, 15 mL; and 0.2 M Na2S·9H2O, 4.0 mL. Clarified rumen fluid was prepared by filtering the rumen fluid collected from the rumen of Holstein steers, followed by centrifugation at 13,000×g for 30 min. The rumen fluids were autoclaved at 121°C for 20 min and recentrifuged to obtain a clear yellow solution. The yellow solution (5%, v/v) was used to supply unknown growth factors. The medium was prepared under a H2/CO2 (80:20, v/v) gas phase at 173 kPa (25 psi) and its pH was adjusted between 6.8 and 7.0. The medium was sterilized by autoclaving at 121°C for 30 min.
Enrichments were performed in the medium with pH 7.0 adjusted under H2/CO2 (80:20, v/v) gas phase. A 5-mL inoculum was added to vials containing medium (45 mL). To inhibit the growth of bacteria, streptomycin sulfate (7×104 IU/L) and benzyl penicillin (2×1010 IU/L) were added to the medium. The inoculated medium was incubated at 38°C in the dark for 2 weeks. After detection of high levels of methane in the culture, 5 mL of the culture was anaerobically transferred into a new vial of sterile medium. Roll tubes containing the medium with 1.8% agar were prepared, followed by four successive transfers. Well-isolated colonies were withdrawn with Pasteur pipettes and transferred to culture tubes containing the medium under anaerobic condition. The culture tubes were sealed with butyl-rubber stoppers and repressurized with sterile filtered H2/CO2 (80:20, v/v) at 173 kPa (25 psi). The organism was re-isolated with the solid medium from the liquid cultures and inoculated on the bacterial growth medium to check for the purity of methanogen cultures. The bacterial growth medium contained peptone (2.5 g/L), yeast extract (2.5 g/L), d-glucose (0.5 g/L), d-cellobiose (0.5 g/L), and d-xylose (0.5 g/L). An antibiotic mixture containing four antibiotics (benzyl penicillin, 0.5 mg/mL; streptomycin sulfate, 0.5 mg/mL; vancomycin-HCl, 0.2 mg/mL; and ampicillin, 0.2 mg/mL) was prepared for the further purification of the isolated strain KOR-2. After purification, the isolate was incubated in the media without antibiotics.
Optimum growth temperature, pH, and NaCl concentration as well as substrate utilization and antibiotic tolerance were investigated to determine the physiological characteristics of the KOR-2 strain. Growth was confirmed by observing optical density (OD) at 660 nm (OD660) with a spectrophotometer (V-530, Jasco, Tokyo, Japan) and measuring the methane concentration in the gas phase using a gas chromatograph (GC-2010A, Shimadzu, Kyoto, Japan). All experiments for physiological studies were repeated twice.
The medium without acetate, formate, and methanol was prepared under N2 and used for substrate utilization studies. Anaerobic stocks of the filter-sterilized substrates (sodium formate, sodium acetate, trimethylamine, methanol, 2-propanol, and isobutanol) were prepared and separately added at a final concentration of 50 mM. Freshly grown cultures of the isolate were inoculated at 10% (v/v) and vials were incubated at 38°C for 20 days. The medium under H2/CO2 served as the control.
The optimum growth temperature in the medium was determined at optimum pH. Vials inoculated with 10% (v/v) culture were incubated at temperatures ranging from 20°C to 50°C. The vials were pressurized every other day with H2/CO2 to ensure an adequate supply of substrate.
The optimum growth pH in the medium was determined at the optimum temperature, with pH values ranging from 4.0 to 9.0. Media with pH above 4.0 were prepared by adding sterile sodium carbonate (Na2CO3) to media at pH 4.0 until the required pH value was reached. Medium with pH 4.0 was produced by removing sodium bicarbonate (NaHCO3) from the medium and cooling it under a CO2 headspace.
The sensitivity of KOR-2 strain to ampicillin, penicillin G, spectromycin, kanamycin, tetracycline, and chloramphenicol (all at a concentration of 100 μg/mL) was tested. Aliquots (5 mL) of the cultures were inoculated into fresh media containing one of the six antibiotics. KOR-2 strain was incubated for 1 week at 38°C. The tolerance to antibiotics was determined by comparing the growth of cultures containing these antibiotics to that of the control.
The salinity range of the isolate was tested at NaCl concentrations ranging from 0.5% to 3.0% at an interval of 0.5%. Media with various concentrations of NaCl were prepared by adding a sterile anoxic stock solution of 58.44 g/L NaCl.
An Olympus BX41 phase-contrast microscope (Olympus, Tokyo, Japan) was routinely used to observe cells. Motility was determined by the hanging-drop method using a glass cavity slide.
Culture samples of KOR-2 strain grown in the media were used for DNA isolation (FastDNA SPIN kit for soil, MP Biomedicals, Irvine, CA, USA) following the manufacturer’s instructions. DNA integrity was evaluated on 1% agarose gels and DNA concentration was determined using a Nanodrop (ND 2000, Thermo Fisher Scientific, Waltham, MA, USA). The DNA G+C content was analyzed from thermal denaturation profiles [18]. The experiments for DNA extraction and G+C content of KOR-2 strain were conducted several times until clear results were obtained.
The primer pair Ar109f (5′-ACKGCTCAGTAACACGT-3′) [19] and Ar1383r (5′-CGGTGTGTGCAAGGAGCA-3′) [20] was used to obtain 16S rRNA gene amplicons (~1,350 bp). Reaction mixtures contained the following components in a final volume of 20 μL within a 200-μL PCR tube: 2 μL of PCR buffer (Takara, Tokyo, Japan), 2 μL of dNTP mix (35 mM), 0.5 μL of each primer (10 pmol/μL), 0.1 μL of Taq DNA polymerase (Takara, Japan), 1.0 μL of template DNA sample (100 ng), and 13.9 μL of molecular grade water (Severn Biotech Ltd, Kidderminster, Worcester, UK). PCR was started by immediately placing the reaction tubes into a preheated (94°C) thermal cycler (PCR Thermal Cycler Dice, Takara, Japan). The thermal program was as follows: Initial denaturation step (94°C, 4 min) followed by 30 cycles of denaturation (94°C, 30 s), annealing (55°C, 30 s), and extension (72°C, 90 s). After a final extension step (72°C, 6 min), samples were stored at 4°C until further analysis. Each PCR run included a positive control of DNA extracted from pure cultures and a negative PCR control where molecular grade water was substituted for the DNA template. The DNA product was analyzed using gel electrophoresis (1.2% w/v agarose gel stained and run at 100 V for 30 min in 1× tris-acetate ethylene-diamine-tetraacetic acid [EDTA; TAE] buffer with 5 μL aliquots of each DNA product). TAE buffer comprised 40 mM Tris base, 20 mM acetic acid, and 0.5 M EDTA (pH 8.0). The gel was run with 2.5 μL of Ladder I DNA quantification marker (TNT research, Seoul, Korea) and imaged using the Bio Imaging System (Daihan, Seoul, Korea). Photographs were obtained using Wise Capture II software (Daihan, Korea).
The mcrA gene fragments were amplified using the primer combinations MLf (5′-GGTGGTGTMGGATTC ACACARTA YGCWACAGC-3′) and MLr (5′-TTCATTGCRTAGTTWGG RTAGTT-3′), yielding ~490-bp amplicons [21]; ME1 (5′-GCM ATGCARATHGGWATGTC-3′) and ME2 (5′-TCATKGCRTA GTTDGGRTAGT-3′), yielding ~740-bp amplicons [22]; and MR1 (5′-GACCTCCACTWCGT VAACAACGC-3′) and ME2, yielding amplicons of ~1,100 bp [23]. Denaturation, annealing, and extension were carried out at 96°C (15 s), 55°C (30 s), and 72°C (90 s), respectively, with MLf/MLr primers, and 94°C (40 s), 50°C (45 s), and 72°C (90 s), respectively, with ME1/ME2 and MR1/ME2 primers.
Purification of PCR products was performed with the AccuPrep PCR purification kit (Bioneer, Daejeon, Korea). PCR products were sequenced using the BigDye terminator cycle sequencing kit on ABI 3730XL capillary DNA Sequencer (Applied Biosystems, Thermo Fisher Scientific Inc., Carlsbad, CA, USA). 16S rRNA and mcrA gene sequences from the isolated strain were compared to the similar sequences obtained from GenBank using the BLAST program. Phylogenetic analysis was conducted using MEGA 4.0 [24]. We examined eight additional 16S rRNA sequences (M. bourgensis MS2 [HE964772], M. palmolei [Y16382], M. receptaculi ZC-2 [DQ787476], M. taiwanensis CYW4 [KM111599], Methanofollis tationis [AF 095272], Methanoregula formicica SMSP [CP003167], Methanogenium marinum AK-1 [DQ177344], and Methanoplanus limicola DSM 2279 [CM001436]). Phylogeny was further confirmed with mcrA gene sequences (M. bourgenisis MS2 [AB AB300787], M. bourgensis RC/ER [AB300785], M. bourgensis CB1 [AB300786], M. bourgensis MAB1 [KJ708788], M. bourgensis MAB2 [KJ708789], M. chikugoensis NBRC 101202 [AB 703634], M. chikugensis [AB300779], M. thermophius DSM 2624 [AF313804], and M. thermophiles [AB300783]).
A new methanogen was isolated from the rumen of Holstein steers. The methanogenic enrichment culture was obtained after repeated transfer in the presence of antibiotic mixtures for 2 months. Visible colonies on solid media appeared after 2 weeks of incubation at 38°C. Surface colonies were about 0.5 to 1.0 mm in diameter, yellow, circular, and convex. Only one strain, designated as KOR-2, was characterized in detail.
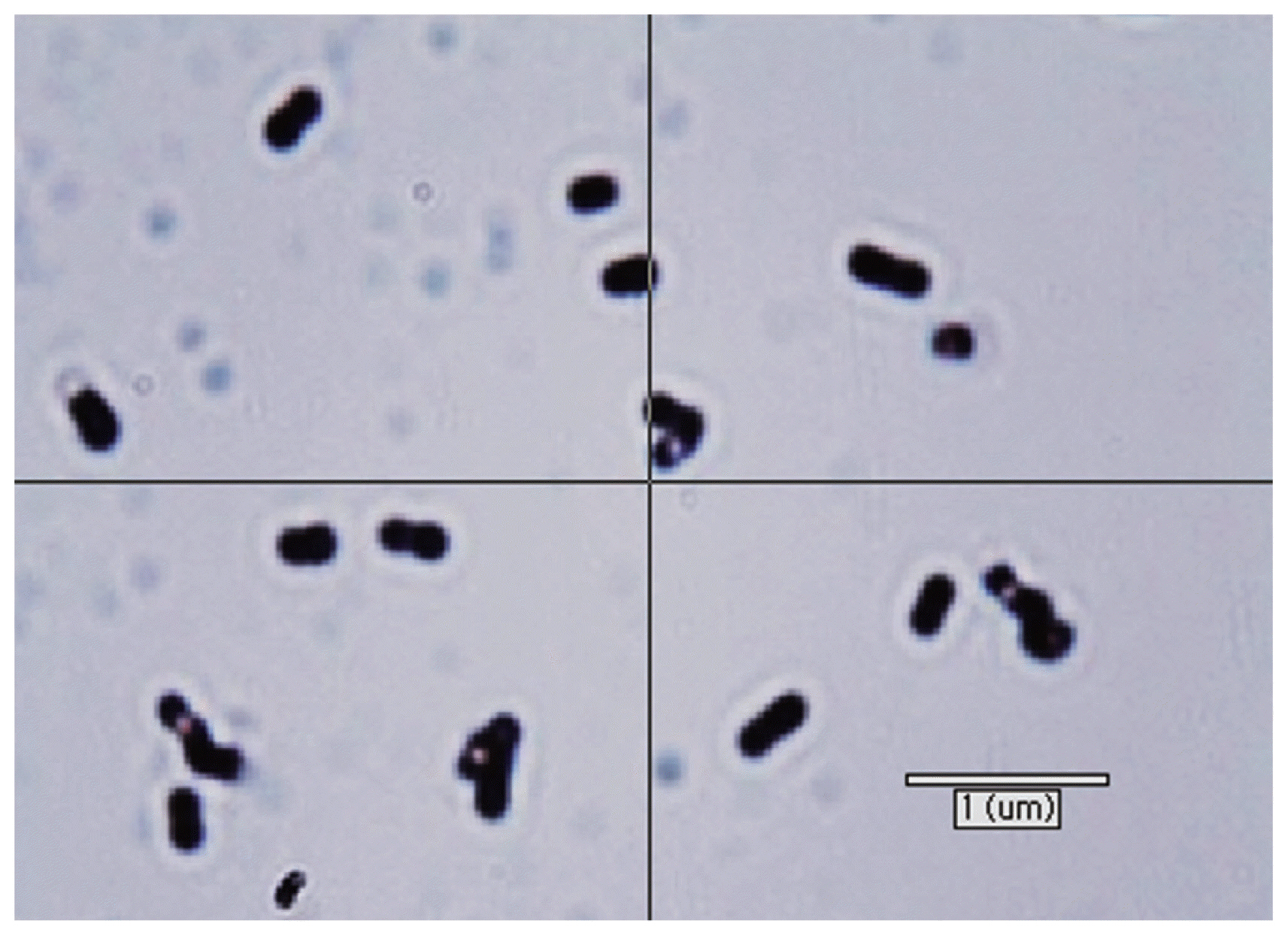
Table 2 shows the comparison between the phenotypic and growth characteristics of KOR-2 strain and M. bourgensis MS2 [25–27]. KOR-2 cells were irregular cocci, non-motile, 0.2 to 0.5 μm in diameter, and occurred singly or in pairs (Figure 1). The growth of KOR-2 strain was observed at a temperature range of 25°C to 45°C, with the fastest growth reported at 38°C. The pH range suitable for its growth was 4.0 to 9.0, and the optimum pH for growth was 6.8 to 7.0. KOR-2 could grow well in salinity up to 3.0% (w/v) and the optimum NaCl concentration for the strain was 1.0%. This range of salinity is typical for halotolerant organisms. The G+C content of genomic DNA of KOR-2 strain was 55.5 mol%. KOR-2 used H2/CO2 and sodium formate (50 mM) but was unable to metabolize sodium acetate, methanol, trimethylamine, 2-propanol, and isobutanol as substrates for growth and methane production. The isolate was phenotypically similar to the compared strain M. bourgensis. Maestrojuán et al [26] reported that the cells of Methanoculleus are irregular cocci, 0.5 to 2.0 μm in diameter, and gram-negative and occur singly or in pairs. Some species are motile. Members of the family Methanomicrobiaceae have been found in a wide variety of anaerobic environments where methane is produced, such as the rumen of ruminant animals, anaerobic marine sediments, wetlands, and oil wells [28]. The genus Methanoculleus contains nine species [29]. M. bourgensis (basinym: Methanogenium bourgense), a species including the former M. olentangyi comb. nov. isolated from a tannery by-product enrichment culture inoculated with sewage sludge [25] (basinym: Methanogenium olentangyi) [30], and M. oldenburgensis [31] were described as junior heterotypic synonyms. M. hydrogenitrophicus was obtained from a wetland soil. Methanoculleus strains, including M. bourgensis MS2, are obligate anaerobes that produce methane from H2/CO2 or formate [25,29]. Some species also produce methane from secondary alcohols and CO2. Acetate is generally required as a carbon source, and additional growth factors may be required. Two types of methanogens, Methanosarcina sp. and Methanosaeta sp., were known to be capable of metabolizing acetate [32]. KOR-2 has typical mesophilic temperature optima, whereas other Methanoculleus species have higher or lower optima. Furthermore, some species are moderately thermophilic [33]. The G+C content of the DNA varies between 55.5 and 62.9 mol%. The G+C content value for KOR-2 was within the range for the strains belonging to the genus Methanoculleus, as reported by Ollivier et al [25]. Based on the morphology and substrate utilization, KOR-2 strain may exhibit the basic characteristics of M. bourgensis.
KOR-2 was able to grow in the presence of ampicillin, penicillin G, kanamycin, spectromycin, and tetracycline in the media, while the cell growth was inhibited by chloramphenicol. Sensitivity to antibiotics was reported for a limited number of species of the family Methanomicrobiaceae, M. receptaculi [34], Methanofollis aquaemaris [35], Methanofollis formosanus [36], and Methanogenium frittonii (a later heterotypic synonym of M. thermophilus) [37] and members of the genus Methanoplanus [38]. Cells are sensitive to chloramphenicol and resistant to penicillin, ampicillin, kanamycin, vancomycin, and streptomycin [29]. Methanogenium frittonii, Methanofollis aquaemaris, and Methanofollis formosanus were sensitive to tetracycline, but Methanoplanus spp. were reported to be resistant [29]. M. receptaculi, but not Methanogenium frittonii, was inhibited by erythromycin [29]. Hilpert et al [39] found that archaea were insensitive to many antibiotics that inhibit eubacteria and eukaryotes, such as those inhibiting the synthesis or cross-linkage of the peptide subunit of murein or those suppressing RNA synthesis. Thus, KOR-2 was insensitive to antibiotics used in this study except for chloramphenicol. Chloramphenicol as a protein inhibitor interferes with the cell membrane function of methanogens. However, it is unknown if this insensitivity to chloramphenicol was associated with the impermeability of the cytoplasmic membrane or inactivation of the antibiotic by the cell, rather than the absence of a particular target for the antibiotic [39].
Polymerase chain reaction of 16S rRNA yielded an amplicon size of 1,350 bp. mcrA gene-based amplification was also used for identification purposes and yielded a product size of 790 bp. Comparative 16S rRNA gene sequence analysis showed that the strain was affiliated with the order Methanomicrobiales. The closest relatives of KOR-2 strain were M. bourgensis MS2 (98%, sequence similarity), M. palmolei (97%), M. receptaculi ZC-2 (96%), Methanofollis tationis (91%), and Methanoplanus limicola DSM 2279 (90%) (Figure 2). KOR-2 showed a small difference in physiological characteristics (Table 2) and 98% sequence similarity to the 16S rRNA gene of M. bourgensis MS2 (Figure 2). The RDP Release 11 (Update 3) reported that a total of 8623 sequences of archaeal 16S rRNA gene sequences were originated from the rumen of ruminants. About 90% of these sequences were assigned to methanogens. These sequences were classified to 10 known genera, with Methanobrevibacter being represented by 63.2% of all the sequences followed by Methanosphaera (9.8%), Methanomicrobium (7.7%), and Methanobacterium (1.2%) [3]. Among the gene sequences of the rumen, the 5 sequences of Methanoculleus were identified, in which 4 sequences were recovered from isolates including one gene from M. bourgensis KOR-2. It is noted that the genus of Methanoculleus is not probably major species in the rumen. The mcrA gene sequence also indicated that KOR-2 strain was a member of the order Methanomicrobiales. The closest relatives of KOR-2 strain based on the mcrA gene sequence were M. bourgensis MS2 (100%) and M. chikugoensis (94%) (Figure 3). All phylogenetic results were similar in the experiment. Luton et al [40] stated that the mcrA gene sequence may be alternatively used instead of the 16S rRNA-based sequence methods, demonstrating far greater diversity than that observed with 16S rRNA gene sequences in the methanogen population.
On the basis of morphology, physiological characteristics, and phylogenetic analyses described above, the strain was identified as a new strain of M. bourgensis and named as KOR-2.
ACKNOWLEDGMENTS
This work was supported by the Korea Institute of Planning and Evaluation for Technology in Food, Agriculture, Forestry and Fisheries (IPET) through the Agri-Bio industry Technology Development Program, funded by the Ministry for Agriculture, Food and Rural Affairs (MAFRA) (116086-3).
Figure 2
Phylogenetic dendrogram of 16S rRNA gene sequences showing the position of the Methanoculleus bourgensis strain KOR-2 relative to other species of the genus Methanoculleus as well as selected reference sequences of methanogens. Methanofollis tationis and Methanoregula formicica were used as outgroup references. The evolutionary distances were computed using the maximum composite likelihood method [24]. Bootstrap values are shown at nodes (percentages of 500 replicates). GenBank accession numbers are indicated. A bar represents 0.01 substitutions per nucleotide position.

Figure 3
Phylogenetic tree of deduced mcrA gene sequences indicating the relationship between the Methanoculleus bourgensis strain KOR-2 and members of the genus Methanoculleus. GenBank accession numbers are indicated. Bootstrap values are shown at nodes (percentages of 500 replicates). A bar represents 0.01 substitutions per nucleotide position.

Table 1
Composition of the concentrate mix
| Items | |
|---|---|
| Ingredients | g/kg of dry matter |
| Ground corn | 30.2 |
| Wheat | 21.0 |
| Soybean meal | 24.0 |
| Rice bran | 10.0 |
| Tapioca | 178.0 |
| Sesame oil meal | 14.0 |
| Palm kernel meal | 410.8 |
| DDGS | 220.0 |
| Molasses | 50.0 |
| Condensed molasses soluble | 10.0 |
| Salt | 3.0 |
| Limestone | 20.0 |
| CaCO3 | 7.0 |
| Minerals and vitamin mixture1) | 7.0 |
| Chemical composition | |
| Dry matter | 882.7 |
| Crude protein | 145.0 |
| Ether extract | 67.7 |
| Crude fiber | 126.8 |
| Undegradable protein | 73.6 |
| Ash | 74.9 |
| NFE | 471.0 |
| NFC | 198.3 |
| ADF | 230.8 |
| NDF | 396.9 |
| TDN | 710.2 |
Table 2
Comparative characteristics of KOR-2 strain isolated from the rumen of Holstein steers1)
| Characteristics | KOR-2 | MS2T |
|---|---|---|
| Cell morphology | Coccoid | Coccoid |
| Cell width (μm) | 0.2–0.5 | 1.0–2.0 |
| Temperature for growth (°C) | ||
| Range | 25–45 | 30–49 |
| Optimum | 38 | 37 |
| pH for growth | ||
| Range | 4.0–9.0 | 5.5–8.0 |
| Optimum | 6.8–7.0 | 6.7 |
| NaCl for growth range (%) | ||
| Range | 0.5–3.0 | 0–4.0 |
| Optimum | 1.0 | 1.0 |
| DNA G+C content (mol%)2) | 55.5 (Tm) | 59 (Bd) |
| Substrate utilization | ||
| H2/CO2 | + | + |
| Formate | + | + |
| Acetate | − | − |
| Methanol | − | − |
| Trimethylamine | − | − |
| 2-propanol | − | − |
| Isobutanol | − | − |
| Tolerance for antibiotics | ||
| Ampicillin | + | ND |
| Penicillin G | + | ND |
| Spectromycin | + | ND |
| Kanamycin | + | ND |
| Tetracycline | + | ND |
| Chloramphenicol | − | ND |
REFERENCES
1. Knapp JR, Laur GL, Vadas PA, Weiss WP, Tricarico JM. Invited review: Enteric methane in dairy cattle production: Quantifying the opportunities and impact of reducing emissions. J Dairy Sci 2014; 97:3231–61.
2. Johnson KA, Johnson DE. Methane emissions from cattle. J Anim Sci 1995; 73:2483–92.
3. Patra A, Park T, Kim M, Yu Z. Rumen methanogens and mitigation of methane emission by anti-methanogenic compounds and substances. J Anim Sci Biotechnol 2017; 8:13
4. Rother M, Krzycki JA. Selenocysteine, pyrrolysine, and the unique energy metabolism of methanogenic archaea. Archaea 2010; 2010:453642
5. Valle ER, Henderson G, Janssen PH, et al. Considerations in the use of fluorescence in situ hybridization (FISH) and confocal laser scanning microscopy to characterize rumen methanogens and define their spatial distributions. Can J Microbiol 2015; 61:417–28.
6. Battumur U, Yoon YM, Kim CH. Isolation and characterization of a new Methanobacterium formicicum KOR-1 from an anaerobic digester using pig slurry. Asian-Australas J Anim Sci 2016; 29:586–93.
7. Battumur U, Yoon YM, Bae GS, Kim CH. Isolation and characterization of new Methanosarcina mazei strains KOR-3, -4, -5, and -6 from an anaerobic digester using pig slurry. Asian-Austalas J Anim Sci 2017; 30:1198–205.
8. Ziemer CJ, Sharp R, Stern MD, et al. Comparison of microbial populations in model and natural rumens using 16S ribosomal RNA-targeted probes. Environ Microbiol 2000; 2:632–43.
9. Garrity GM, Lilburn TG, Cole JR, et al. Taxonomic outline of the Bacteria and Archaea. Part 1. The Archaea, phyla Crenarchaeota and Euryarchaeota [Internet]. Release 7.7. Lansing, MI, USA: Michigan State University; 2007. [cited 2007 Mar 6]. Available from: www.taxonomicoutline.org
10. DeLong EF, Pace NR. Environmental diversity of bacteria and archaea. Syst Biol 2001; 50:470–8.
11. Janssen PH, Kirs M. Structure of the archaeal community of the rumen. Appl Environ Microbiol 2008; 74:3619–25.
12. Lee J-H, Kumar S, Lee G-H, et al. Methanobrevibacter boviskoreani sp. nov., isolated from the rumen of Korean native cattle. Intl J Syst Evol Microbiol 2013; 63:4196–201.
13. Official methods of analysis. 16th edAssociation of Official Analytical Chemists. Washington, DC, USA: AOAC International; 1995.
14. Van Soest PJ, Robertson JB, Lewis BA. Methods for dietary fiber, neutral detergent fiber, and nonstarch polysaccharides in relation to animal nutrition. J Dairy Sci 1991; 74:3583–97.
15. Hungate RE. The anaerobic mesophilic cellulolytic bacteria. Bacteriol Rev 1950; 14:1–49.
16. Balch WE, Fox GE, Magrum LJ, Woese CR, Wolfe RS. Methanogens: reevaluation of a unique biological group. Microbiol Rev 1979; 43:260–96.
17. Sowers KR, Schreier HJ. Media for methanogens. Sowers KR, Schreier HJ, editorsArchaea - a laboratory manual methanogens. New York, USA: Cold Spring Harbor Laboratory Press; 1995. p. 459–89.
18. Sly LI, Blackall LL, Kraat PC, Tian-Shen T, Sangkhobol V. The use of second derivative plots for the determination of mol% guanine plus cytosine of DNA by the thermal denaturation method. J Microbiol Methods 1986; 5:139–56.
19. Großkopf R, Janssen PH, Liesack W. Diversity and structure of the methanogenic community in anoxic rice paddy soil microcosms as examined by cultivation and direct 16S rRNA gene sequence retrieval. Appl Environ Microbiol 1998; 64:960–9.
20. Shlimon AG, Friedrich MW, Niemann H, Ramsing NB, Finster K. Methanobacterium aarhusense sp. nov., a novel methanogen isolated from a marine sediment (Aarhus Bay, Denmark). Int J Syst Evol Micrbiol 2004; 54:759–63.
21. Luton PE, Wayne JM, Sharp RJ, Riley PW. The mcrA gene as an alternative to 16S rRNA in the phylogenetic analysis of methanogen populations in landfill. Microbiology 2002; 148:3521–30.
22. Hales BA, Edwards C, Ritchie DA, et al. Isolation and identification of methanogen-specific DNA from blanket bog feat by PCR amplification and sequence analysis. Appl Environ Microbiol 1996; 62:668–75.
23. Simankova MV, Kotsyurbenko OR, Lueders T, et al. Isolation and characterization of new strains of methanogens from cold terrestrial habitats. Syst Appl Microbiol 2003; 26:312–8.
24. Tamura K, Dudley J, Nei M, Kumar S. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 2007; 24:1596–9.
25. Ollivier BM, Mah RA, Garcia JL, Boone DR. Isolation and characterization of Methanogenium bourgense sp. nov. Int J Syst Bacteriol 1986; 36:297–301.
26. Maestrojuán GM, Boone DR, Xun L, Mah RA, Zhang L. Transfer of Methanogenium bourgens, Methanogenium marisnigri, Methanogenium olentangyi, and Methanogenium thermophilicum to the genus Methanoculleus gen. nov., emendation of Methanoculleus marisnigri and Methanogenium, and description of new strains of Methanoculleus bourgense and Methanoculleus marisnigri. Int J Syst Bacteriol 1990; 40:117–22.
27. Asakawa S, Nagaoka K. Methanoculleus bourgensis, Methanoculleus olentangyi and Methanoculleus oldenburgensis are subjective synonyms. Int J Syst Evol Microbiol 2003; 53:1551–2.
28. Boone DR, Whitman WB, Koga Y. Family I. Methanomicrobiaceae Barker 1956, 15, AL emend. Balch and Wolfe in Balch, Fox, Magrum, Woese and Wolfe 1979, 286. Boone DR, Castenholz RW, Garrity GM, editorsBergey’s manual of systematic bacteriology, vol 1. 2nd ed. The Archaea and the deeply branching and phototrophic Bacteria. New York, USA: Springer; 2001. p. 247
29. Oren A. The family Methanomicrobiaceae. Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F, editorsThe Prokaryotes: other major lineages of bacteria and the archaea. Berlin, Germany: Springer; 2014. p. 231–46.
30. Corder RE, Hook LA, Larkin JM, Frea JI. Isolation and characterization of two new methane-producing cocci: Methanogenium olentangyi, sp. nov., and Methanococcus deltae, sp. nov. Arch Microbiol 1983; 134:28–32.
31. Blotevogel K-H, Gahl-Janßen R, Jannsen S, et al. Isolation and characterization of a novel mesophilic, fresh-water methanogen from river sediment Methanoculleus oldenburgensis sp. nov. Arch Microbiol 1991; 157:54–9.
32. Ferry JG. Methanogenesis: ecology, physiology, biochemistry and genetics. New York, NY, USA: Chapman & Hall; 1993.
33. Chong SC, Boone DR. Genus II. Methanoculleus Maestrojuán, Boone, Xun, Mah and Zhang 1990, 121VP. Boone DR, Castenholz RW, Garrity GM, editorsBergey’s manual of systematic bacteriology, vol 1, 2nd ed. The Archaea and the deeply branching and phototrophic Bacteria. New York, USA: Springer; 2001. p. 251–2.
34. Cheng L, Qiu TL, Li X, et al. Isolation and characterization of Methanoculleus receptaculi sp. nov. from Shengli oil field, China. FEMS Microbiol Lett 2008; 285:65–71.
35. Lai MC, Chen SC. Methanofollis aquaemaris sp. nov., a methanogen isolated from an aquaculture fish pond. Int J Syst Evol Microbiol 2001; 51:1873–80.
36. Wu SY, Chen SC, Lai MC. Methanofollis formosanus sp. nov., isolated from a fish pond. Int J Syst Evol Microbiol 2005; 55:837–42.
37. Harris JE, Pinn PA, Davis RP. Isolation and characterization of a novel thermophilic, freshwater methanogen. Appl Environ Microbiol 1984; 48:1123–8.
38. Huber H, Huber G, Stetter KO. Genus VI. Methanoplanus Wildgruber, Thomm and Stetter 1984, 270VP (Effective publication: Wildgruber, Thomm, König, Ober, Ricchiuto and Stetter 1982, 36). Boone DR, Castenholz RW, Garrity GM, editorsBergey’s manual of systematic bacteriology, vol 1. 2nd ed. The Archaea and the deeply branching and phototrophic Bacteria. New York, USA: Springer; 2001. p. 259–61.
39. Hilpert R, Winter J, Hammes W, Kandler O. The sensitivity of archaebacteria to antibiotics. Zbl Bakt Mik Hyg I C 1981; 2:11–20.
40. Luton PE, Wayne JM, Sharp RJ, Riley PW. The mcrA gene as an alternative to 16S rRNA in the phylogenetic analysis of methanogen populations in landfill. Microbiology 2002; 148:3521–30.






 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Download Citation
Download Citation Print
Print





