 |
 |
| Anim Biosci > Volume 28(12); 2015 > Article |
|
Abstract
Since ancient days, domestic horses have been closely associated with human civilization. Today, horse racing is an important industry. Various genes involved in energy production and muscle contraction are differentially regulated during a race. Among them, creatine kinase (CK) is well known for its regulation of energy preservation in animal cells. CK is an iso-enzyme, encoded by different genes and expressed in skeletal muscle, heart, brain and leucocytes. We confirmed that the expression of CK-M significantly increased in the blood after a 30 minute exercise period, while no considerable change was observed in skeletal muscle. Analysis of various tissues showed an ubiquitous expression of the CK-M gene in the horse; CK-M mRNA expression was predominant in the skeletal muscle and the cardiac muscle compared to other tissues. An evolutionary study by synonymous and non-synonymous single nucleotide polymorphism ratio of CK-M gene revealed a positive selection that was conserved in the horse. More studies are warranted in order to develop the expression of CK-M gene as a biomarker in blood of thoroughbred horses.
Domestic horses (Equus caballus) have been associated with humans since the ancient days. At present, horses are mainly used in sports such as horseracing. Performance in a race is tightly linked with the skeletal muscle, nutrition and the training strategies. Fifty-five percent of body weight in thoroughbreds consists of skeletal muscles, which is widely used as an animal model in exercise physiology (Hinchcliff et al., 2008). The skeletal muscle in a horse is responsible for locomotion and energy production; with numerous genes involved in these processes (Schr├Čder et al., 2011). Various gene differential expression occurs in equine skeletal muscle in the pre and post exercise period (Eivers et al., 2010).
Creatine Kinase (CK) is present in various tissues, such as skeletal muscle, heart muscle, spleen, thyroid and blood as iso-enzymes, and is involved in energy metabolism through catalysis of creatine into the phosphocreatine mechanism. In the skeletal muscle, it is coded by CK-M gene, and knockout studies in mice revealed a significant increase in fatigue resistance as cellular adaptations (Echegaray and Rivera, 2001). Moreover, genetic polymorphisms are associated with the A/G variant in humans, and individuals carrying this genotype have considerably better running economy; larger VO2 and higher adaptations to training compared with G/G and A/A homozygotes (Zhou et al., 2006). CK-M attempts to maintain energy homeostasis by providing a steady supply of creatine phosphate, which is critical for sustaining the Ca2+- adenosine triphosphatase of the sarcoplasmic reticulum and other energy-dependent enzymes (Korge et al., 1993).
The main purpose of this study was to identify the gene expression level in blood and skeletal muscle of thoroughbred horses during the pre- and post- exercise period and compare nucleotide and amino acid sequence with other species. Quantitative real-time polymerase chain reaction (PCR) results revealed an increased expression of CK-M in the blood in response to exercise, but not in the skeletal muscle. In addition, the degree of expression of the CK-M gene was investigated in various tissues of horses, and when compared with other species, phylogenetic analysis of CK-M gene revealed a close relationship between the nucleotides and the amino acids, which shows a positive evolution in the horse.
Two stallions and one mare Throughbred horses, weighing from 500 to 700 kg, which were maintained in Ham-An Racing Horse Resort and Training Center and not genetically related, were used for blood samples. Three Jeju horses (3 mares), which were not genetically related, were used for tissue samples. Tissue samples of Jeju horses were provided by the National Institute of Subtropical Agriculture, Rural Development Administration, Republic of Korea. Sampling occurred from August 2014 to November 2014. Age of the horses were from 36 month to 40 months. The Pusan National University-Institutional Animal Care and Use Committee (PNU-2015-0864) approved this study.
Briefly, the samples were collected from three thoroughbreds in the morning before and after subjecting them to physical exercise. The animals performed a combination of different horse gaits which included trotting and cantering through lunging and long-reining (circular bridge lunging) as their form of exercise. Generally, racehorses are subjected to exercise for 17 to 18 min per day. However horses in this study followed a combined 30-min exercise of trotting and cantering. Before and after the exercise, venous blood samples from the jugular vein were taken from the Thoroughbreds using a 15 mL syringe. For skeletal muscle biopsy collection, local anesthesia was given in the gluteus medius, and a biopsy collection syringe was used to get muscle samples before and after the exercise. The National Institute of Subtropical Agriculture provided various tissues from three Jeju horses. Skeletal muscle, kidney, thyroid, lung, appendix, spinal cord and heart tissues were kept in a liquid nitrogen tank until the RNA extraction.
Trizol reagent (Invitrogen) was used to extract total RNA from samples, according to the manufacturerŌĆÖs manual. In order to prevent contamination of genomic DNA, RNase-free DNase kit (Qiagen, Valencia, CA, USA) was used according to the manufacturerŌĆÖs operating manual. Total RNA quantification was performed using a NanoDrop ND-1000 Spectrophotometer (Thermo Scientific, Wilmington, DE, USA). cDNA was synthesized in a reaction with oligo-dT primers, M-MLV (moloney-murine leukemia virus) reverse transcriptase (Promega, Madison, WI, USA), RNase inhibitor (Promega, USA) and RNase-free ddH2O, which was incubated at 37┬░C for 4 h.
The horse CK-M transcript primers were designed using the PRIMER3 software (http://frodo.wi.mit.edu/primer3/). The following CK-M primers were used: forward: (5ŌĆ▓-GACCT CAGCA AGCAC AACAA -3ŌĆ▓) and reverse: (5ŌĆ▓-AGAGT GAAGC CCGAT GGAGT -3ŌĆ▓). For the normalization process, we used a house keeping gene, GAPDH, as described (Song et al., 2014). The expected product size of this gene was 95 base pairs (bp).
cDNA was analyzed by BioRad CFX-96 (BioRad, Hercules, CA, Country), and each reaction was executed in total volume of 25 ╬╝L: 12.5 ╬╝L SYBR green master mix, 0.5 ╬╝L (5 pmol) CK-M forward primer, 0.5 ╬╝L (5 pmol) reverse primer, 10.5 ╬╝L distilled water and 1 ╬╝L (50 ng/╬╝L) of the cDNA. PCR conditions were as follows: pre-denaturation step at 94┬░C for 7 min, 49 cycles at 94┬░C for 30 s and at 60┬░C for 30 s, 72┬░C for 30 s, followed by 72┬░C for 10 min as a final step. Dissociation was accomplished in a condition, in which temperature increased from 60┬░C to 95┬░C over 25 min (linear increase at 0.5┬░C). All samples were measured in triplicate to ensure reproducibility, and the Ct value was calculated using 2ŌłÆ╬ö╬öCt method (Livak et al., 2001).
The coding sequence region of CK-M gene was sequenced for comparison analysis with other species. The following CK-M primers were used: i) Forward: (5ŌĆ▓-ATATA AGGCC AGGGC TGGAG -3ŌĆ▓) and Reverse: (5ŌĆ▓-GTCGA ACAGG AAGTG GTCGT -3ŌĆ▓), ii) Forward: (5ŌĆ▓-CTGGA CCCTC ATTAC GTGCT -3ŌĆ▓) and Reverse: (5ŌĆ▓-GCTGG GGCTA CAGCA GTC -3ŌĆ▓). Also, the CK-M coding sequence and amino acid sequences in various species (human, chimpanzee, mouse, rat, cow, pig, dog and cat) were retrieved from National Center for Biotechnology Information (NCBI, www.ncbi.nlm.nih.gov/). The BioEdit (Ibis Biosciences, Carlsbad, CA, USA) software with CLUSTERW option was used to align the sequences in various species. MEGA5 (http://www.megasoftware.net) software was used to obtain the Ka/Ks ratio analysis data (Tamura et al., 2011). Substitutions of nucleotides were obtained using the Kimura 2-parameter model. We also showed synonymous substitutions per site (Ks) and non-synonymous substitutions per site (Ka) in CK-M genes for individual variation.
The CK-M gene sequences in various species were obtained from NCBI for comparative genome composition analysis. Table 1 represents the nucleotide similarity and the amino acid identity (AI) of the CK-M gene. When compared with the horse, the rat is the least similar while the pig is the most similar, with similarity being 0.016 and 0.059, respectively. The AI was calculated by dividing the number of different amino acids to the number of all amino acids by the pair wise distance.
When compared to other species, the AIs of the horse CK-M and those of the rat (0.039) were the lowest while the highest AIs were those of the cat (0.018), but the entire amino acids pair wise distance was shown to be close. In order to characterize horse CK-M coding protein, amino acid sequences of CK-M from various species were aligned and analyzed. The amino acid sequences are highly conserved between species except for the N-terminal of the sequence (Figure 1). Domain search results through SMART program (http://smart.embl-heidelberg.de) confirmed that the horse CK-M domain function is related SMART to enzyme adenosine triphosphate guanido phosphotransferases. Phylogenetic analysis of the equine CK-M gene with the Neighbor-Joining method showed that the pig and the rodent were, respectively, the closest and the furthest species (Figure 2).
The non-synonymous (Ka) and synonymous (Ks) substitution sites in the CK-M genes of various animals were analyzed for an evolutionary study (Table 2). Ka ranged from 0 to 0.22 whereas Ks ranged from 0.023 to 0.456. The average of Ka was 0.014 and that of Ks was 0.306. The results of the Ka and Ks ratio revealed a positive selection in horse in contrast to other species. In other words, synonymous substitutions occurred faster than non-synonymous substitutions. These results are based on the supposition that the CK-M coding sequence regions of protein in the human genome are under positive selection during evolution (Bustamante et al., 2005). As a result, we confirmed that evolution of horse CK-M gene was very similar to that of other species.
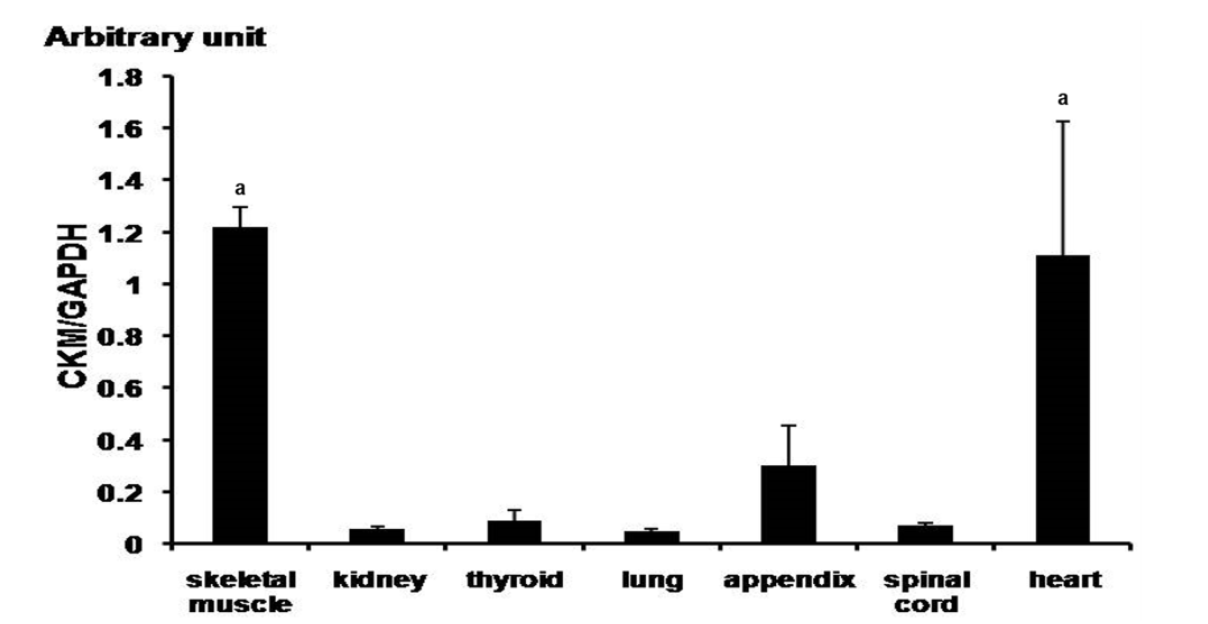
In order to confirm ubiquitous expression of CK-M in various tissues of horse, we analyzed skeletal muscle, kidney, thyroid, lung, appendix, and spinal cord and heart tissues. In particular, the CK-M was highly expressed in the skeletal muscle and the heart (Figure 3). Results of the digital gene expression analysis in equine skeletal muscle showed the highest distribution of CK-M transcript (McGivney et al., 2010). Generally, CK-M type iso-enzyme is abundant in skeletal muscle, while CK-MB type is found more in the cardiac muscle. There were very low levels of CK-M expression found in all tissues except for the skeletal muscle and the heart muscle when compared to the human gene (Tsung, 1976). We analyzed CK-M gene level in the skeletal muscle and the blood of thoroughbred horses. The results revealed a significant increase of CK-M expression in blood after 30-minute exercise, while in the skeletal muscle, no considerable change was observed (Figure 2). These results strongly correlate with the elevation of CK enzyme in blood of thoroughbred horses, at the completion of the exercise (Octura et al., 2014). The main function of the CK-M gene is to synthesize CK enzyme, which catalyzes energy formation concerned with energy metabolism in the cytoplasm (Wallimann et al., 1992). The CK-M expression is related to muscle contraction by occurrence of energy and it has an effect on muscle contraction proteins in muscular fiber. In a previous study, the immunoblotting results showed that the muscle specific CK has a higher ratio in the skeletal muscle than in the cardiac muscle in laboratory animals (Fredericks et al., 2001). The discordant results regarding the gene expression level in blood might be due to the physiological process during the exercise. However, more studies with large sample size are warranted to confirm these preliminary results.
This study characterized the equine CK-M gene at the molecular level, and compared its nucleotide and amino acid sequences with other species. Our analysis of various tissues of the horse showed a higher expression of CK-M gene in the skeletal muscle than in other tissues. This correlates with another previous study, which showed highly abundant CK-M transcripts in the skeletal muscle (McGivney et al., 2010). The sequence variations of CK-M and other genes in equine skeletal muscle (Gu et al., 2010) were also reported (Hill et al., 2010; 2012). Expression of CK-M increased in the blood after the exercise period, but not in the skeletal muscle. These results supported the notion that CK-M expression can be developed as a molecular marker for the racing ability of racehorses and the blood could be used as a specimen, rather than the invasive procedure of the skeletal muscle collection.
ACKNOWLEDGMENTS
This work was supported by a grant from the Next Generation Bio Green 21 Program (No. PJ01104401, PJ011173), Rural Development Administration, Republic of Korea.
Figure┬Ā1
Amino acid sequence comparison analysis of CK-M gene in nine species. (A) Amino acid alignment of various species indicates strong conservation. The domain is related to enzyme ATP guanido phosphotransferases. (B) Phylogenetic tree of CK-M in various species. The Phylogenetic tree analysis of horse confirmed that its closest and farthest clade are pigs and rodents respectively. CK-M, creatine kinase, muscle.

Figure┬Ā2
Quantification of CK-M expression by q-PCR in various tissues of the horses (Jeju horses, Equus caballus). Expression value was calculated with 2ŌłÆ╬ö╬öCt method. The expression value was revealed to gene expression in horse (n = 3, p<0.01). The mean fold values are presented by mean┬▒standard error. The difference value of gene expression was verified by TukeyŌĆÖs t-test. The GAPDH gene was used as control. CK-M, creatine kinase-muscle; GAPDH, glyceraldehyde 3-phosphate dehydrogenase. a, p<0.05.
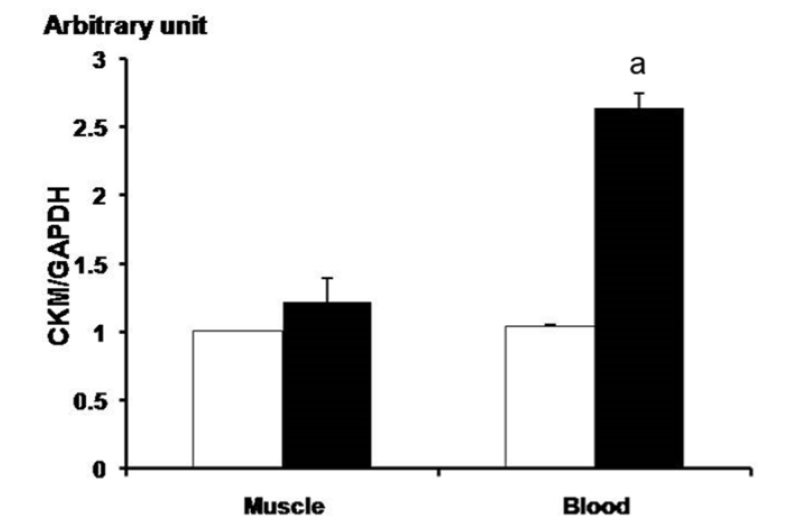
Figure┬Ā3
Analysis of CK-M gene expression before and after exercise using qPCR in the thoroughbred horsesŌĆÖ muscle and blood. The white and black bars means before condition and after, respectively. Before: before the exercise; after: 30 min after the exercise. Expression level was calculated by 2ŌłÆ╬ö╬öCt method. The expression value of the gene appeared significantly in leukocytes (n = 3). The mean fold values are presented by mean┬▒standard error. The GAPDH gene was used for normalization. The asterisk mark means p<0.05. CK-M, creatine kinase-muscle; GAPDH, glyceraldehydes 3-phosphate dehydrogenease. a, p<0.05.
Table┬Ā1
Nucleotide similarity (NS) and amino acid identity (AI) in the CK-M gene of various species1
1 The average AI and NS were 0.028 and 0.083, respectively. According to the results of comparative analysis of genetic informative, we found that horse nucleotide of CK-M was relatively in consensus to pigs (0.059) as amino acids were to cats (0.018), but generally, CK-M gene was well conserved with various species.
REFERENCES
Bustamante CD, Fledel-Alon A, Williamson S, Nielsen R, Hubisz MT, Glanowski S, Tanenbaum DM, White TJ, Sninsky JJ, Hernandez RD, Civello D, Adams MD, Cargill M, Clark AG. 2005. Natural selection on protein-coding genes in the human genome. Nature 437:1153ŌĆō1157.


Echegaray M, Rivera MA. 2001. Role of creatine kinase isoenzymes on muscular and cardiorespiratory endurance. Sports Med 31:919ŌĆō934.


Eivers SS, McGivney BA, Fonseca RG, MacHugh DE, Menson K, Park SD, Rivero JL, Taylor CT, Katz LM, Hill EW. 2010. Alterations in oxidative gene expression in equine skeletal muscle following exercise and training. Physiol Genomics 40:83ŌĆō93.


Fredericks S, Merton GK, Lerena MJ, Heining P, Carter ND, Holt DW. 2001. Cardiac troponins and creatine kinase content of striated muscle in common laboratory animals. Clin Chim Acta 304:65ŌĆō74.


Gu J, MacHugh DE, McGivney BA, Park SDE, Katz LM, Hill EW. 2010. Association of sequence variants in CKM (creatine kinase, muscle) and COX4I2 (cytochrome c oxidase, subunit 4, isoform 2) genes with racing performance in Thoroughbred horses. Equine Vet J 42:569ŌĆō575.

Hill EW, Fonseca RG, McGivney BA, Gu J, MacHugh DE, Katz LM. 2012. MSTN genotype (g. 66493737C/T) association with speed indices in Thoroughbred racehorses. J Appl Physiol 112:86ŌĆō90.


Hill EW, Gu J, McGivney BA, MacHugh DE. 2010. Targets of selection in the Thoroughbred genome contain exercise-relevant gene SNPs associated with elite racecourse performance. Anim Genetics 41:56ŌĆō63.

Hinchcliff KW, Geor RJ. 2008. The Horse as an Athlete: A Physiological Overview. Equine Exercise Physiology: The Science of Exercise in the Athletic Horse. Saunders/Elsevier; Edinburgh, UK: p. 463

Korge P, Byrd SK, Campbell KB. 1993. Functional coupling between sarcoplasmic-reticulum-bound creatine kinase and Ca2+-ATPase. Eur J Biochem 213:973ŌĆō980.


Livak KJ, Schmittgen TD. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2ŌłÆ╬ö╬öCT method. Methods 25:402ŌĆō408.


McGivney BA, McGettigan PA, Browne JA, Evans AC, Fonseca RG, Loftus BJ, Lohan A, Machugh DE, Murphy BA, Katz LM, Hill EW. 2010. Characterization of the equine skeletal muscle transcriptome identifies novel functional responses to exercise training. BMC Genomics 11:398



Octura JER, Lee KJ, Cho HW, Vega RS, Choi JY, Park JW, Shin TS, Cho SK, Kim BW, Cho BW. 2014. Elevation of blood creatine kinase and selected blood parameters after exercise in thoroughbred racehorses (Equus caballus L). Quest J 2:7ŌĆō13.
Schr├Čder W, Klostermann A, Distl O. 2011. Candidate genes for physical performance in the horse. The Veterinary Journal 190:39ŌĆō48.


Song KD, Cho HW, Lee HK, Cho BW. 2014. Molecular Characterization and Expression Analysis of Equine Vascular Endothelial Growth Factor Alpha (VEGF╬▒) Gene in Horse (Equus caballus). Asian Australas J Anim Sci 27:743ŌĆō748.




Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. 2011. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731ŌĆō2739.



Tsung SH. 1976. Creatine kinase isoenzyme patterns in human tissue obtained at surgery. Clin Chem 22:173ŌĆō175.







 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print





