 |
 |
| Anim Biosci > Volume 35(10); 2022 > Article |
|
Abstract
Objective
Methods
Results
Notes
CONFLICT OF INTEREST
We certify that there is no conflict of interest with any financial organization regarding the material discussed in the manuscript.
FUNDING
This research was funded by the National Key R&D Program of China (No. 2018YFD0502000), Key Laboratory of Molecular Animal Nutrition of Ministry of Education of China, the Science and Technology Major Project of Anhui Province (No. 201903b06020001), National Innovative Training Program for College Student (No. 201910364046), The National Natural Science Foundation of China (No. 32002199), Anhui province Natural Science Foundation of China (No. 2008085MC86), and National Natural Science Foundation of China (No. 32002199).
Figure 1
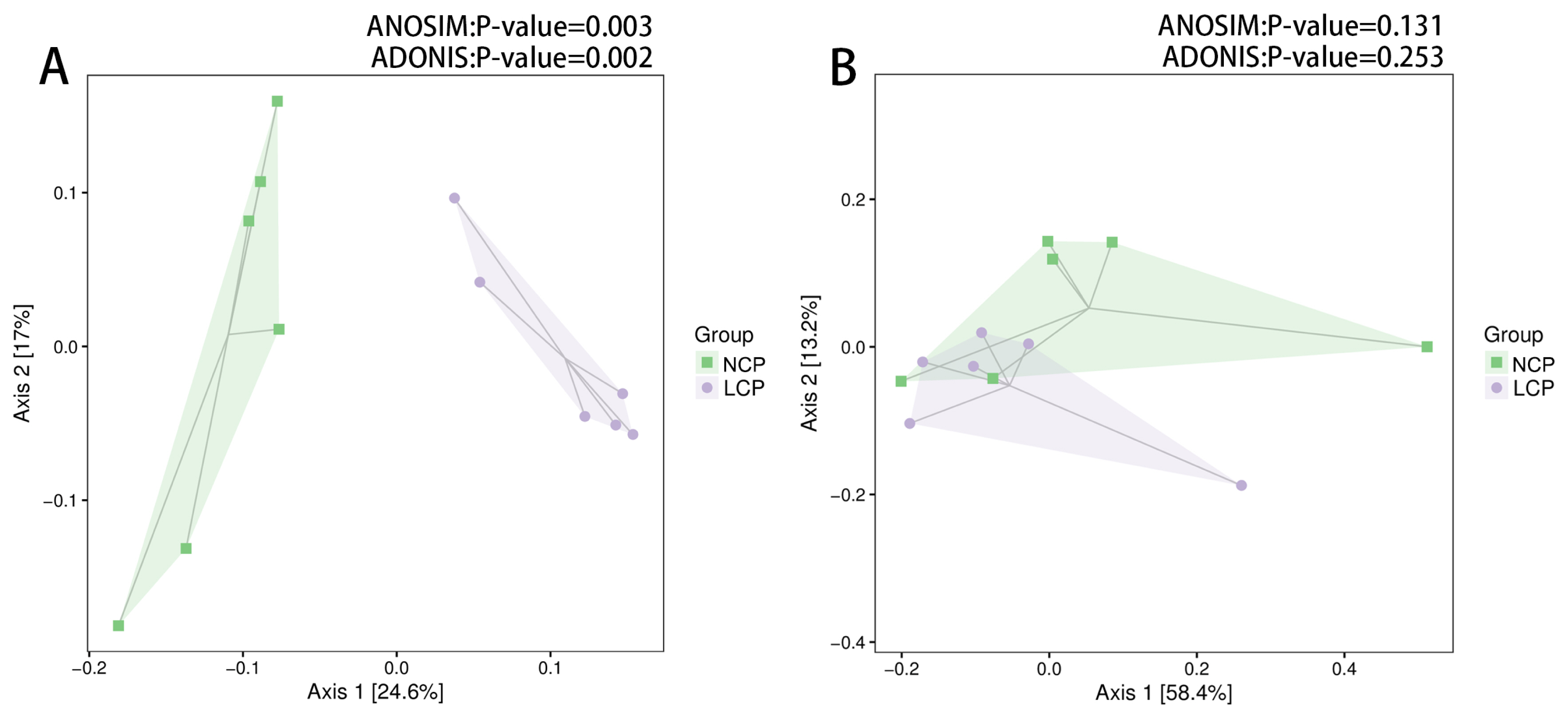
Figure 2
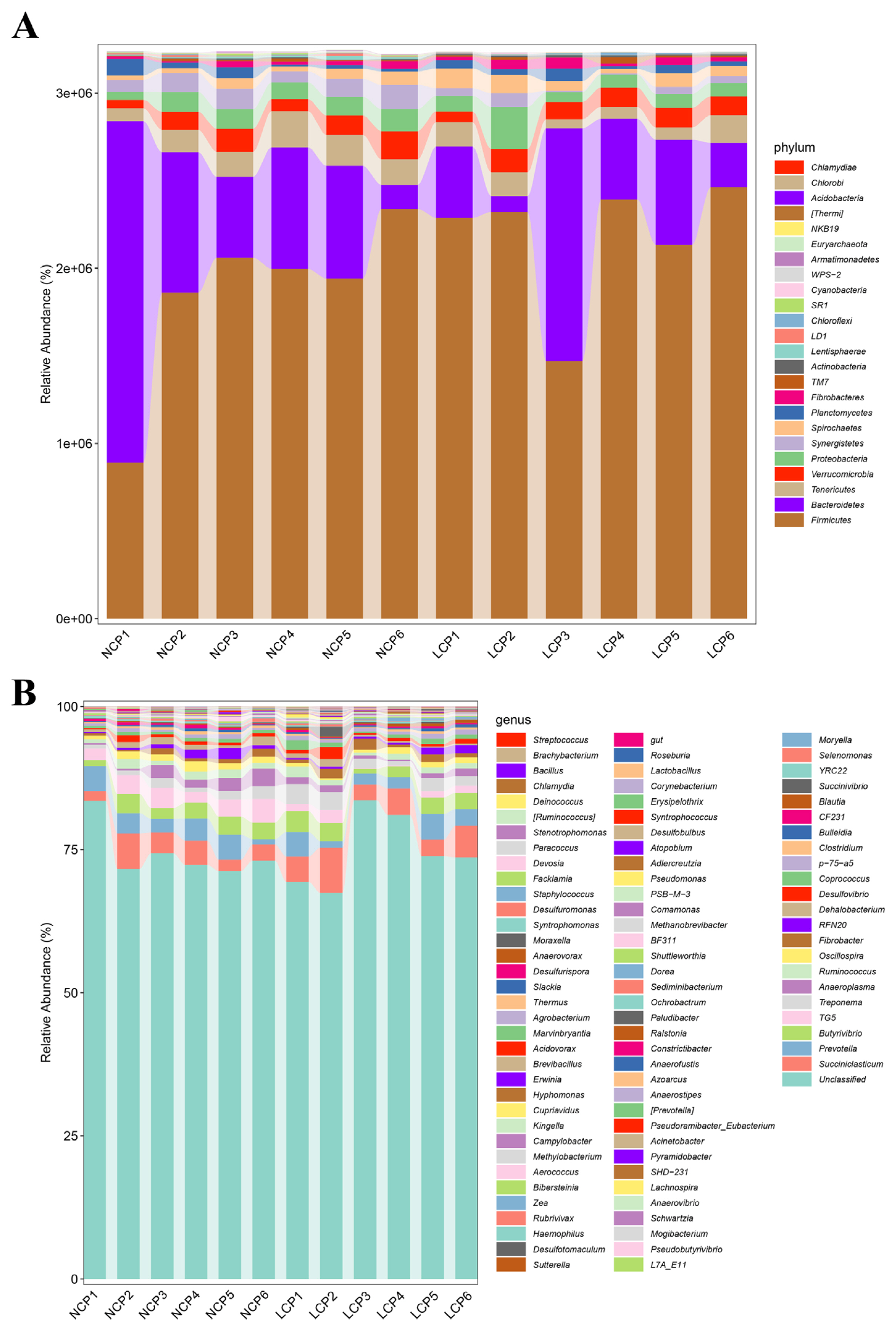
Figure 3
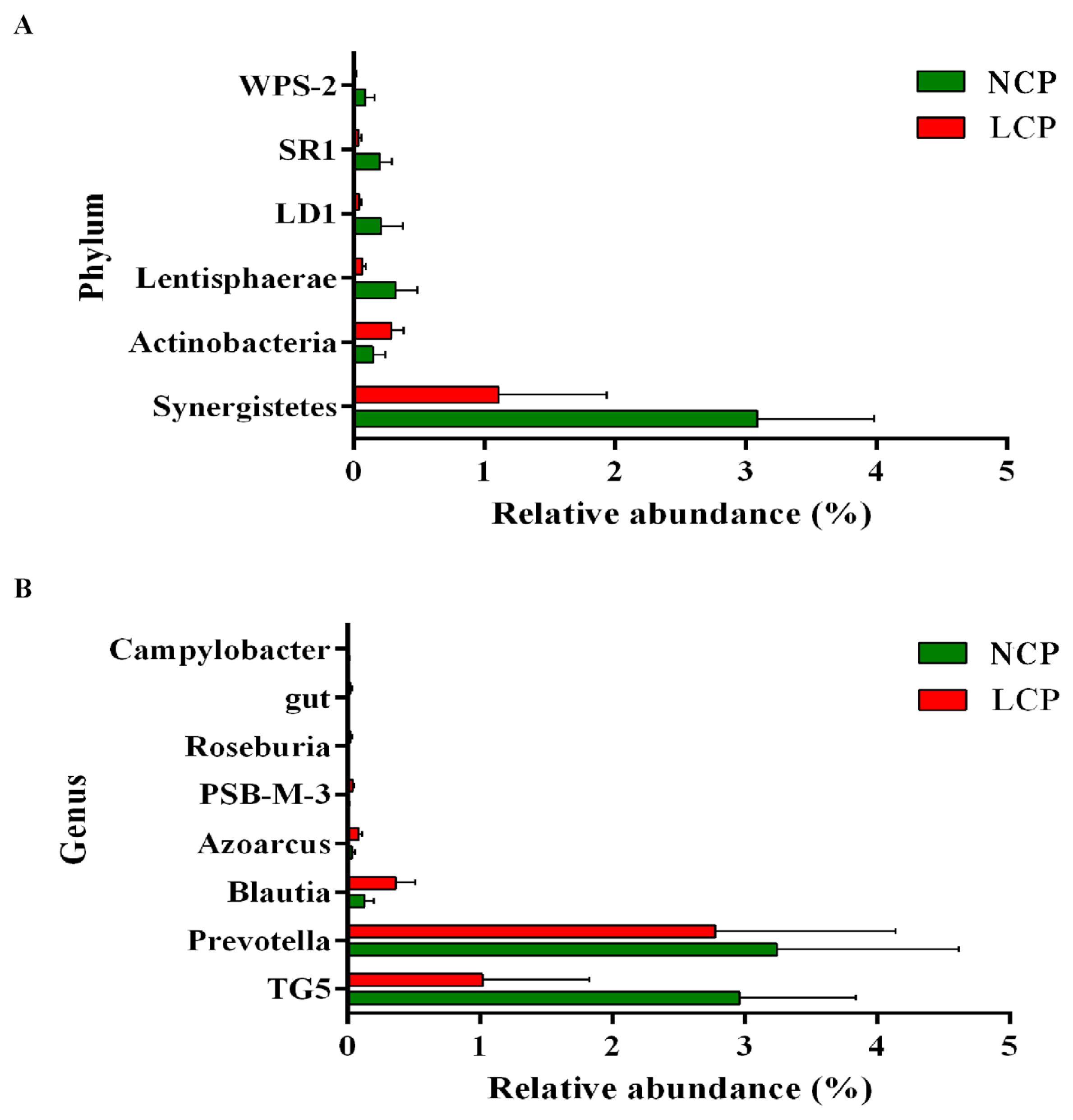
Figure 4

Figure 5
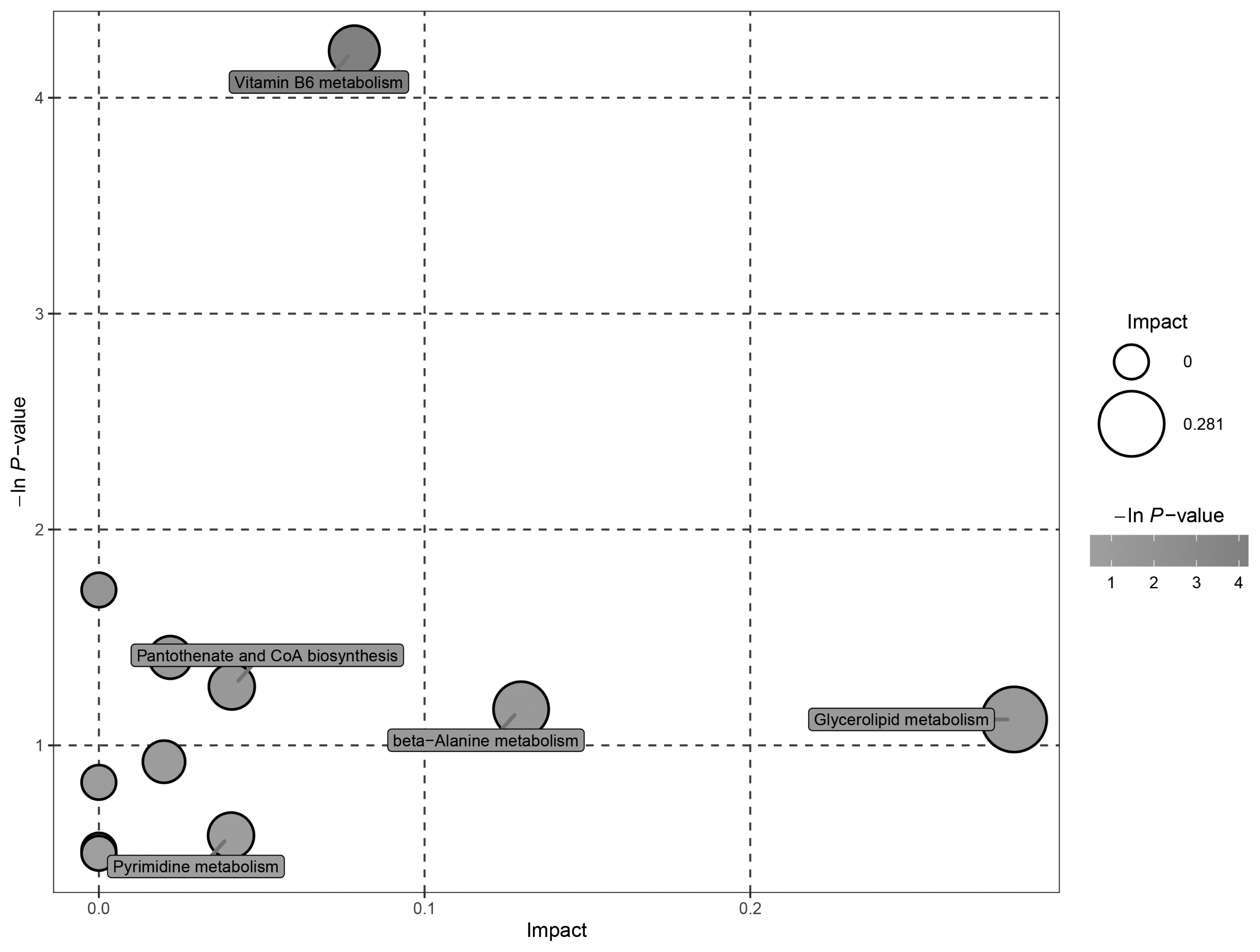
Table 1
| Item | Dietary treatments, % dry matter | |
|---|---|---|
|
|
||
| NCP | LCP | |
| Ingredient | ||
| Ground corn grain | 13.0 | 21.0 |
| Soybean meal2), 43.5% CP | 24.0 | 16.0 |
| Wheat bran | 7.5 | 7.5 |
| Sodium bicarbonate | 1.0 | 1.0 |
| Salt | 1.0 | 1.0 |
| Dicalcium phosphate | 0.5 | 0.5 |
| Calcium carbonate | 1.0 | 1.0 |
| Premix3) | 1.0 | 1.0 |
| Peanut vine | 28.0 | 28.0 |
| Chinese wild rye | 22.0 | 22.0 |
| Chemical composition | ||
| Organic matter | 86.3 | 88.3 |
| Crude protein | 14.8 | 12.0 |
| Neutral detergent fiber | 46.4 | 44.8 |
| Acid detergent fiber | 31.8 | 31.2 |
| Ether extract | 3.74 | 3.27 |
| Digestible energy4) (MJ/kg) | 13.9 | 13.8 |
1) Cited from Zhu et al [4].
2) Soybean meal contained: 89.5% dry matter, 43.5% crude protein, 28.2% neutral detergent fiber and 10.5% acid detergent fiber.
3) Formulated to provide (per kg of dry matter): 600,000 IU of vitamin A, 80,000 IU of vitamin D3, 5,000 IU of vitamin E, 8,000 mg of Zn, 60 mg of Se, 200 mg of I, 9,400 mg of Fe, 72 mg of Co, 10,400 mg of Mn, and 1,600 mg of Cu; Peanut vine contained: 91.3% dry matter, 7.17% crude protein, 57.2% neutral detergent fiber, and 50.5% acid detergent fiber on dry matter basis; Chinese wild rye contained: 89.6% dry matter, 6.7% crude protein, 67.5% neutral detergent fiber, and 38.2% acid detergent fiber on dry matter basis;
Table 2
| Items | Treatment | SEM | p-value | |
|---|---|---|---|---|
|
|
||||
| NCP | LCP | |||
| pH | 6.64 | 6.51 | 0.038 | 0.49 |
| Ammonia-nitrogen (mg/dL) | 10.5a | 8.94b | 0.431 | 0.03 |
| Total volatile fatty acid (mM) | 74.3a | 65.1b | 2.51 | 0.03 |
| Acetate (mM) | 52.6a | 45.8b | 1.71 | 0.04 |
| Propionate (mM) | 13.1a | 11.2b | 0.46 | 0.02 |
| Butyrate (mM) | 7.78 | 7.28 | 0.37 | 0.59 |
| Valerate (mM) | 0.81 | 0.83 | 0.029 | 0.67 |
| Acetate:propionate | 3.73 | 3.75 | 0.121 | 0.68 |
Table 3
Table 4
| Pathway | Total | Hits1) | p-value2) | Impact3) | Hits conpounds |
|---|---|---|---|---|---|
| Vitamin B6 metabolism | 9 | 1 | 0.015 | 0.000 | 4-Pyridoxic acid |
| Arginine and proline metabolism | 44 | 1 | 0.245 | 0.022 | Argininosuccinic acid |
| Glycerolipid metabolism | 18 | 1 | 0.326 | 0.281 | Glycerol |
| Alanine, aspartate and glutamate metabolism | 23 | 1 | 0.397 | 0.020 | Argininosuccinic acid |
| Galactose metabolism | 26 | 1 | 0.436 | 0.000 | Glycerol |
| Tryptophan metabolism | 41 | 1 | 0.596 | 0.000 | Indoleacetic acid |
REFERENCES
- TOOLS






 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Supplement1
Supplement1 Print
Print





