INTRODUCTION
After strong artificial selection for over 150 years, morphological variation has been created in dog populations and more than 400 canine breeds are currently registered worldwide by the Fédération Cynologique Internationale (FCI), and other federations such as the American and British Kennel Clubs [
1]. With over 470 million dogs being kept as companion animals worldwide, they were ranked as the leading type of pet in 2018 [
2]. In Taiwan, according to survey statistics from the Council of Agriculture, the number of dogs being bred by the end of 2017 had reached 1.78 million. The overall sales revenue of pet-related industries has increased from 15.5 billion New Taiwan dollars (NT$) in the past 10 years to 26.6 billion NT$ (Statistical Bulletin, Ministry of Finance, Republic of China;
http://service.mof.gov.tw/File/Attach/86088/File_21588.pdf). The previous roles of companion animals, such as hunting, security, and assistance, have gradually shifted towards them being regarded as family members. With the improvement of the status of pets, owners are increasingly paying attention to their quality of life. Therefore, the demand for various goods and services aimed at companion animals is also increasing. In the early stage of the Taiwan dog-sales market, the characteristics prioritised included the ability to guard, search, attack, and hunt; more recently, these ideals have been superseded by the need to provide companionship and prestige, which have made a recognized pedigree a major factor for buyers. In Taiwan, some legal purebreed grounds used closed flock breeding to keep their dog broodlines, including those three breeds in this study. Especially the Beagles, which have been used to hunt hares in the British Isles for centuries, and which were brought to the United States in 1880 to breed in large numbers. The modern Beagles have been modified to become a pet dog, and were often used as experimental dogs [
3]. It has considerable medical research value, and this dog was also cultivated as laboratory animal for medical research in Taiwan.
So far, many dog breeds have been developed in order to meet appearance standards and maintain the purity of their bloodlines. Breeding companies usually adopt inbreeding methods, which can lead to the occurrence of many genetic diseases. Generally, in the natural state—unlike with experimental animals for which it can be essential to reduce individual differences for study purposes—inbreeding approaches have seldom been used, in order to avoid inbreeding depression [
4]. The Royal Society for the Prevention of Cruelty to Animals (RSPCA) pointed out that dogs are now subject to more than 300 genetic diseases. Not only do the animals have to bear great suffering, but their owners also experience mental pressures and financial losses [
5]. Therefore, in order to maintain high heterozygosity and stability of the genetic background of the entire population, it is necessary to have a reliable breeding system and genetic monitoring [
6]. However, in Taiwan, the genealogical and registration data requirements for many breeds of dog are incomplete or remain to be established, and most of the certificates of pedigree produced by breeding sites in the market lack the backing of publicly trusted authorities, so trading disputes arise from time to time. Although trading in companion animals is discouraged in many countries today, the market in Taiwan is booming. In order to prevent companion animals from being afflicted by genetic diseases, in addition to promoting care by and education of owners, an important factor is reducing inbreeding. The Kennel Club has established a breeding certification system for 68 breeds since 2008, with the total number now reaching 222 breeds. Breeders can inquire about the diseases to be screened for each dog and the procedures for obtaining certification. After the puppies are certified by the Kennel Club, the breeders are issued with a puppy sale wallet; this measure not only protects the profits of the owner, but also reduces disputes over the sale of companion animals [
7].
Regarding the research on dog microsatellite markers, a lot of information has been discovered in conjunction with the elucidation of DNA sequences [
8]. As early as the 1990s, there have been many studies on dog microsatellite markers [
9,
10]. The application of canine microsatellite markers in modern times has mainly focused on two fields: one is evaluating the genetic structure polymorphism of populations, and the other is proving a platform for individual identification or paternity [
11,
12]. Wictum et al [
13] searched for published dog genome sequences, and selected suitable microsatellite markers based on the stability and high polymorphism that were required for forensic applications. A total of 15 microsatellite markers and a marker related to gender comprise the multiplex system, DogFiler [
13]. This is the first dog-identification data system created based on the recommendations of the Scientific Working Group for DNA Analysis Methods (SWGDAM) in the United States. At present, DogFiler has been integrated into forensic casework, and is widely accepted by courts in the United States. Owing to the relatively long period of strong artificial selection, the differentiation between dog breeds has been large. The sequences on both sides of microsatellite markers may have different degrees of variation, making it impossible to perform polymerase chain reaction (PCR) amplification in similar breeds. Even if the amplification is successful, the number of alleles and polymorphisms may be far lower than in the original breeds [
14]. Therefore, when conducting research on the population genetics of specific breeds of dog, it is necessary to develop new microsatellite markers.
To date, only a few domestic biotechnology companies in Taiwan use foreign dog microsatellite commercial kits, such as the StockMarks for dogs genotyping kit, for individual genetic analysis or genetic structure analysis at dog breeding sites. So, are they applicable to Taiwan? To our knowledge, there are no relevant published reports on existing dog breeds for reference and analysis. Therefore, the development of microsatellite markers suitable for the companion animal population in Taiwan is a crucial task to establish a molecular-detection platform for domestic dogs. Moreover, such a platform will make it possible to evaluate the inbreeding level of the dog population in Taiwan. In addition to assisting with the formulation of domestic dog breeding-management policies, this could also enhance Taiwan’s positive governance perspectives on animal protection and animal welfare.
DISCUSSION
In this study, 14 sets of novel microsatellite markers were developed and used to analyze the genetic variation of three dog breeds—SC, BI, and BE. The results showed that the Na of the 14 novel microsatellite loci was 6.3, and the Ne was 3.6 (
Table 3). The Na of locus SEL115 was 13, whereas the Ne was only 3.8. The reason might have been that the distribution of allele frequency was mainly concentrated on three alleles—243 bp (13.3%), 247 bp (10.6%), and 251 bp (47.4%) (
Supplementary Table S1)—such that in some cases the uneven distribution of allele frequencies caused a large gap between the Na and the Ne.
The H
E, H
O, and PIC are commonly used to assess the polymorphism of microsatellite loci in the analyzed population. The H
O refers to the observed heterozygosity of each locus, which represents the actual proportion of heterozygous individuals in the population. The H
E is the expected heterozygosity of each locus, which is the expected proportion of heterozygous individuals in the population that is calculated according to the Hardy–Weinberg Law. The PIC is the degree of polymorphism of each locus. Using the 14 novel microsatellite markers to analyze our dog populations, the average H
E was 0.662, which showed that the values of most of the microsatellite markers were within high expected heterozygosity (H
E>0.5). The average value of H
O was 0.567, which also fell within the range of high observed heterogeneity (0.7>H
O>0.5) [
26]. The average value of PIC was 0.612, which fell within the range of high polymorphic information content (PIC>0.5) [
27] (
Table 3). The results of this experiment were similar to the study of Radko et al [
28] using 18 sets of microsatellite markers to analyze the Polish Tatra shepherd dog. Their results showed that the average H
E was 0.643, the average H
O was 0.645, the average PIC was 0.598, and the values of the three variables were all greater than 0.5. Therefore, their study indicated that the tested Tatra shepherd dog population was highly polymorphic. In another study [
29], eight breeds of dog were surveyed by 21 microsatellite markers. PIC values over 0.5 were measured for 15 markers. The average value of the PIC was 0.555. Compared with these reports, the three variables in the current experiment were highly polymorphic, indicating that the 14 novel microsatellite loci should be able to effectively analyze the genetic structure and genetic variation of the three breeds of dogs analyzed in this experiment.
The H
E (0.412 and 0.293) and H
O (0.451 and 0.248) values of loci SEL093 and SEL094 were both less than 0.5, as were the respective PIC (0.326 and 0.249) values. The cause of this result, for which there were only two alleles in two loci, was supposed to be sampling error. However, some reports have suggested that the number of alleles for microsatellite markers should be three or more to reduce the standard deviation of distance calculation [
30]. The reason why the loci SEL093 and SEL094 were selected in this experiment was that the number of dog breeds analyzed was relatively small. If the number of breeds is increased, perhaps the allele number of these two microsatellite loci could be increased, and the three variables will be likely to increase as well. On this basis, the two microsatellite loci SEL093 and SEL094 were retained as potential canine microsatellite loci in this experiment.
In terms of the analysis of population genetic structure, we applied the
FIS,
FST, and
FIT statistics to evaluate the distribution of genetic variation within and between populations. The average
FIS value of the 14 new microsatellite markers was 0.002. This value was positive and low. The percentage of heterozygotes in the overall tested dog population was less than expected—that is, there was an inbreeding phenomenon—but the average value of
FIS was around 0.002, which indicated that the situation in the dog population was not serious. The mean
FST for all the loci was 0.212. This fell within the range of high differentiation (0.15<
FST<0.25), according to the Sewall Wright rules [
31], indicating that there was high differentiation among the three breeds of dogs in this study.
Kang et al [
32] investigated the genetic structure of local dogs in South Korea and establish an individual and paternity identification system through evaluating the polymorphisms of the populations with three variables: H
O, H
E, and PIC. Between nine and 11 microsatellite loci were used for genetic analysis of two local breeds from South Korea and three exotic dog breeds in their study. The sample selection criterion was at least one generation of unrelated dog individuals. The results showed that the average H
O for each breed ranged from 0.65 to 0.78; the H
E ranged from 0.71 to 0.85; and the PIC ranged from 0.66 to 0.82. In this study, the average value of H
E for the three varieties ranged from 0.480 (SCs) to 0.624 (BEs), and average H
O ranged from 0.485 (SCs) to 0.587 (BEs). The average PICs ranged between 0.407 (SCs) and 0.567 (BEs) (
Table 4). Compared with the abovementioned studies, the three variables in our experiment showed slightly lower values, which may have been caused by differences in the individual dogs included in this experiment: some of the animals were blood-relatives and were full-sibs or half-sibs, so their genetic backgrounds were similar, leading to slightly lower polymorphisms. Future testing of individual Taiwanese dogs with different origins or more distant blood relationships could improve the applicability of these new microsatellite markers.
Among the different varieties of
FIS, only the average value for BEs (0.045) was positive. The results showed that although BEs in this experiment had high genetic variation, the positive value of
FIS indicated that the proportion of heterozygous individuals was still too small to achieve the Hardy–Weinberg balance; Iindeed, it deviated significantly from the expected value (p<0.01). This result may reflect the fact that the fathers of the BE population in this experiment comprised a small number of male dogs, which was not reflective of the situation of mating by chance. The BIs and SCs did not deviate from the Hardy–Weinberg equilibrium, and the
FIS values of these two breeds were negative. It can thus be inferred that these two breeds had no inbreeding issues, the genetic backgrounds of the parents were different, and the number of male and female animals was equal [
33]. Therefore, it was supposed that the deviation of the entire dog population from the Hardy–Weinberg balance was attributable to the deviation of the BE population.
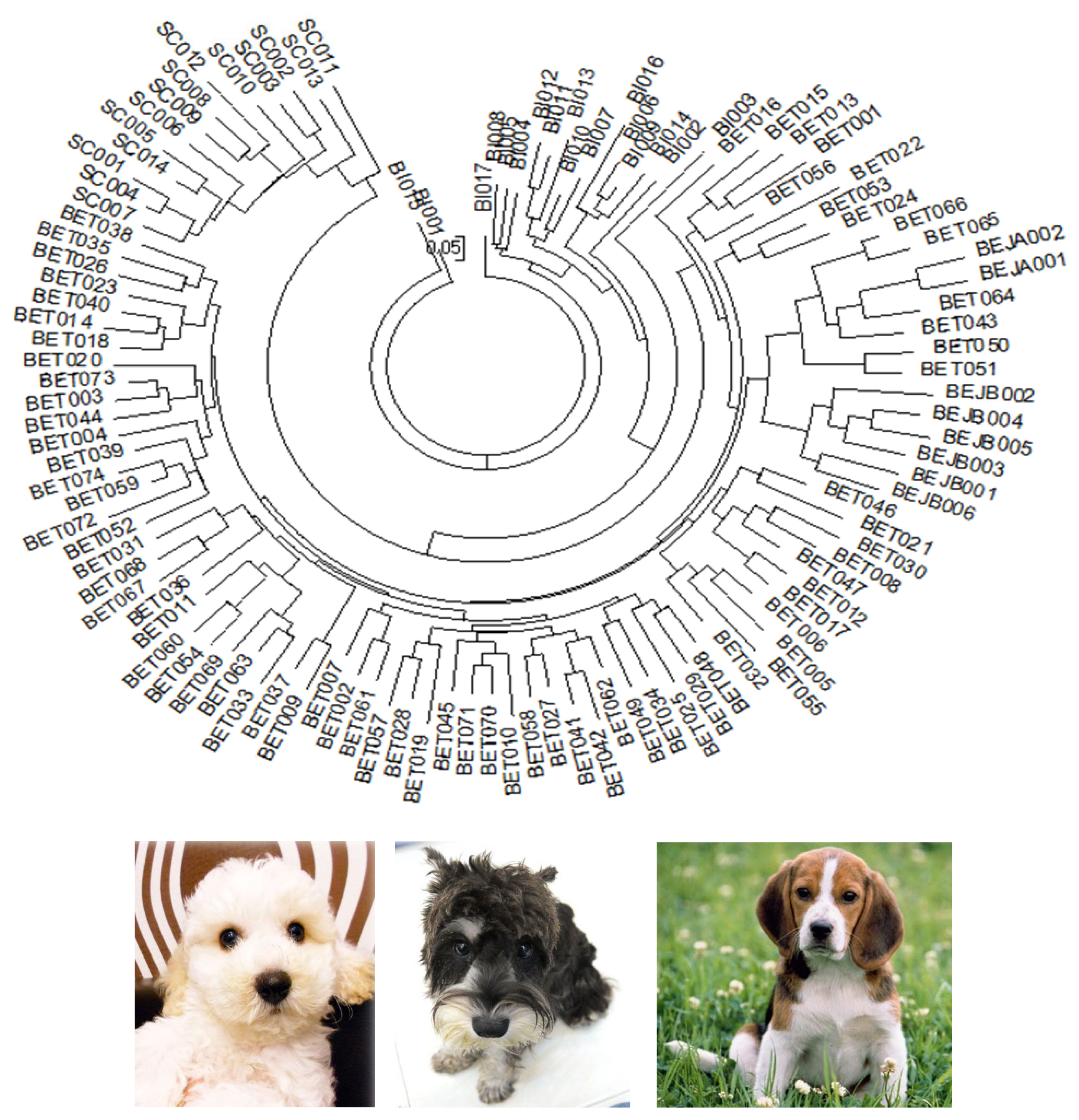
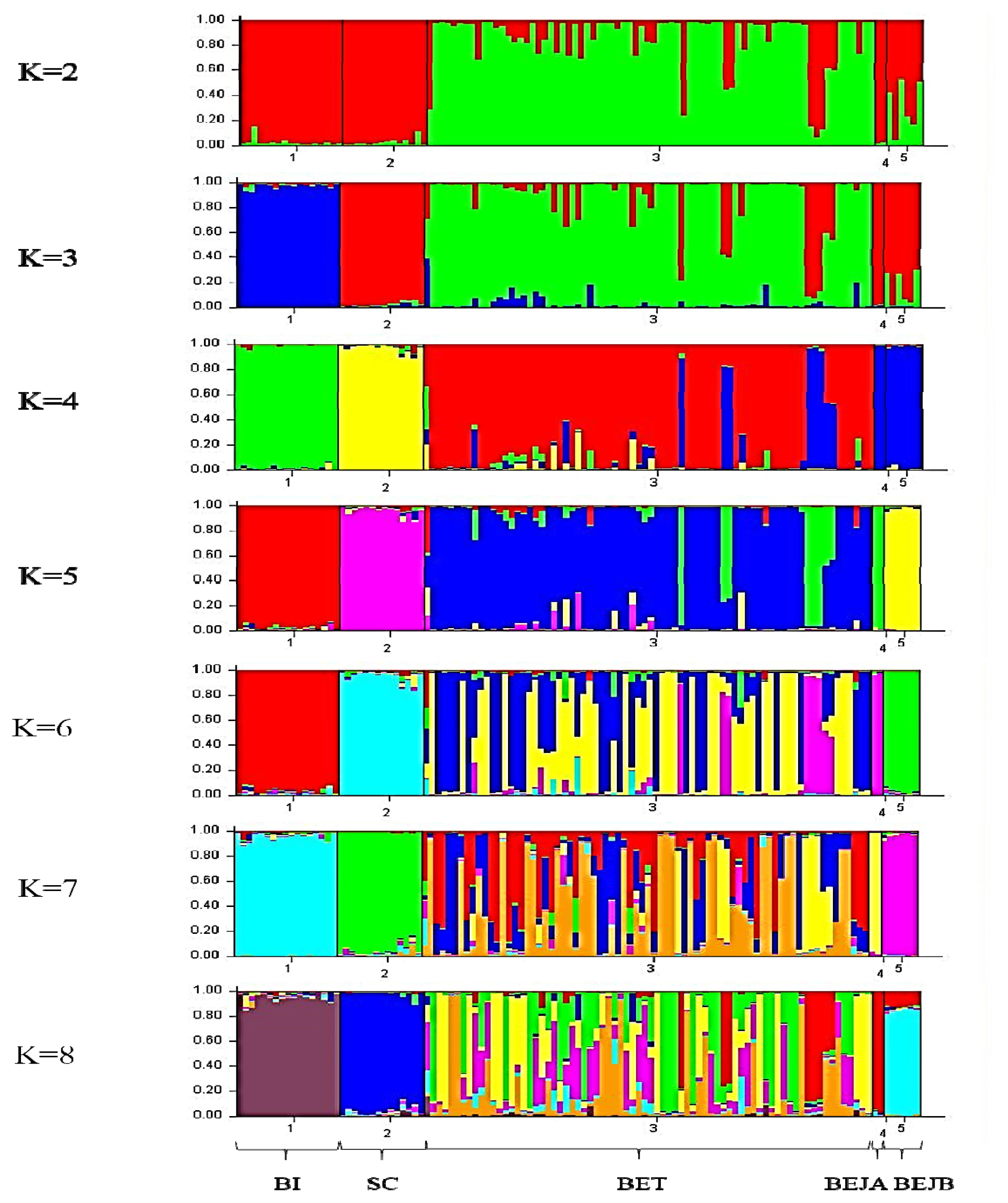
The individual phylogenetic tree of the dogs (
Figure 1) was constructed by the NJ method, and the cluster analysis diagram drawn by the STRUCTURE software (
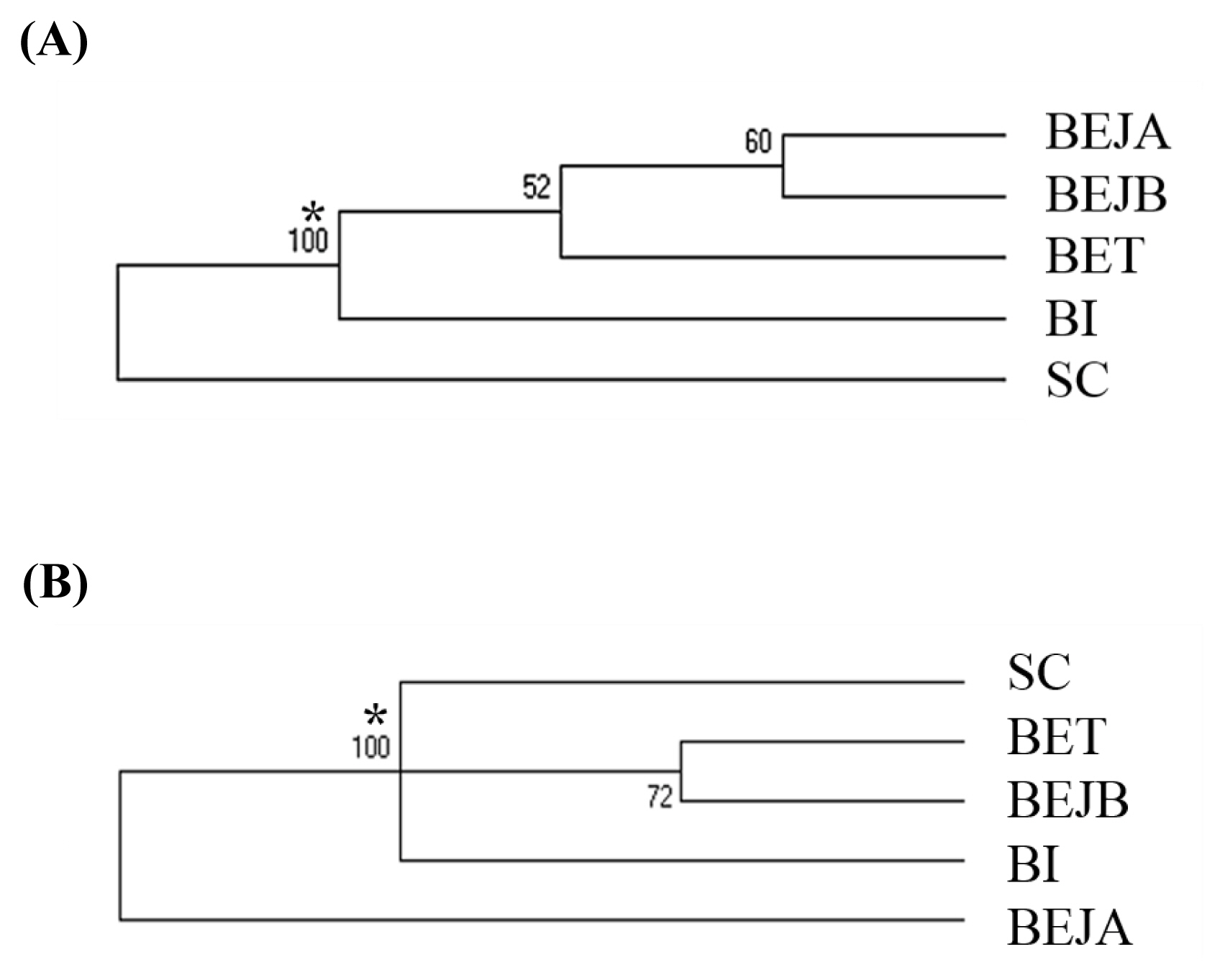
Figure 4), and the dogs in this experiment were divided into five groups: SC, BI, BET, BEJA, and BEJB. In the phylogenetic tree of dog populations drawn by the NJ method (
Figure 2A), the distinctive main clusters were consistent with the result of the individual phylogenetic tree: both could clearly distinguish the BI, SC, and BE groups, and the BET and BEJ groups were closely identified. The different breeds of dogs could be clearly differentiated by the microsatellite markers used in this experiment. The differentiation of dog breed in tree would be caused by the unique alleles. For example, the alleles 185 bp and 201 bp of the SEL035 locus were only found in the BI and SC populations, respectively (
Supplementary Table S1). In the phylogenetic tree drawn by the NJ method, the bootstrap value of the BET and the BEJ group was 52%, which means that only 52% of the analysis results separated them. When the bootstrap value between the two populations was not greater than 70% in neighbor joining tree, it showed that the clustering of that two populations were not obvious [
34]. In the NJ phylogenetic tree, the bootstrap value between the SC and other dog populations was 100%, and the result was in the highly reliable range (bootstrap value>70%). That showed that the genetic distance between the SC and other dog populations was relatively long. However, the UPGMA phylogenetic tree (
Figure 2B) shows that the BEJA group was far away from other dog groups, and the bootstrap value was 100%. But the UPGMA method based on constant-rate assumption, and the distance between samples on the same branch is the same, so this method is only suitable for the case where all samples have the same evolution distance [
35], so recently it was less used to construct a phylogenetic tree, and the NJ method was a more commonly used method for drawing a phylogenetic tree.
The PCoA draws a 3D stereogram based on the genetic distance between the populations (
Figure 3), and distinguishes the genetic distance among the populations by the variance of the three principal coordinate axes. It can be found that the two BEJ groups were relatively close, which was consistent with the close geographical relationship between the two, and the relative distance between the two groups and the BET group was closer than the relative distance between the two groups and the SC and the BI. The results of the NJ phylogenetic tree can also be confirmed in the 3D map of the PCoA. The SC and the BEJA were located on the farthest sides of the 3D map, so there was a maximum genetic distance between the two populations.
In the individual analysis, the combined probability of identification (P
(ID)) of the 14 new microsatellite loci for the entire dog population was 1.7×10
−12 (
Table 6), and according to the survey of the Council of Agriculture up until the end of 2019, the total number of dogs in Taiwan was about 1.54×10
6 [
36]. That meant when the current number of Taiwanese dog populations is analyzed by the 14 new microsatellite markers used in this experiment, the probability of appearing exactly the same genotype is very low. Although the dog population in this experiment contained only three breeds, however, when examining P
(ID) of a single dog breed, the credibility is still high. There are no statistics on the numbers of dogs of different breeds currently in Taiwan; however, it can be ascertained that the number of dogs in any single breed cannot exceed the total number of dogs. Therefore, the P
(ID) of the three dog populations respectively in this study should cover the total number of dogs in Taiwan in 2019. It was confirmed that the probability of the same genotype being identified in two individuals with 14 the sets of novel microsatellite markers was very low.
The combined probability of identity among sibs (P
(ID)sib) was 1.6×10
−5 (
Table 7) in this study. According to Waits et al [
24], their markers were sufficient to identify close relatives of the natural population when the P
(ID)sib was between 10
−3 and 10
−4. The P
(ID)sib values of the three breeds of dogs in this experiment generally met this recommended standard. This confirmed that the 14 novel microsatellite markers were applicable in the individual identification of close relatives of BEs, BIs, and SCs in Taiwan.
In the paternity tests of dogs, the PE of the 14 new microsatellite markers in this experiment was 99.98%, which meant that when the genotypes of the mother and offspring were known, individuals that were not the biological father could be ruled out. According to the pedigree data for the BE population collected in this experiment, there was one of the cases where the mother–child genotypes were known, the individual registered as the biological father in the pedigree was compared with the genotypes of 14 novel microsatellite markers and which was not found to provid any of the alleles to the offspring at six of the loci. Therefore, it could be inferred that the male was not the biological father, and suggested that the registration of the dog’s pedigree data was inaccurate. This supports the suggestion that many dog-breeding facilities in Taiwan are still not rigorous enough for pedigree registration. Therefore, using paternity facilities with high PE to improve the registration of pedigree in Taiwan is important. The microsatellite markers developed in this study may be suitable for this purpose to avoid the potential damage caused by inbreeding.
At present study, many countries used microsatellite markers to analyze dog populations. For developing a platform of paternity and individual identification, the American Kennel Club (AKC) analyzed 108 dog breeds using 17 microsatellite markers. The results show that the average H
O was 0.60, and the average value of PIC was 0.56. PE was more than 99% in all breeds, and the combined P
(ID) was 3.2×10
−8. The American Kennel Association considered that 17 microsatellite markers were sufficient for ordinarily paternity identification [
37]. Eichmann et al. established Austrian dog DNA profiling for investigation of dog-related accidents and crimes, using 15 sets of highly polymorphic microsatellite markers to analyze 45 dog breeds [
38]. The results revealed that the average H
O was 0.74, the average PIC was 0.82, and the combined P
(ID)sib was 8.5×10
−8, showing that these 15 microsatellite markers are sufficient for individual identification of dogs in Austria. Kang et al [
32] used 9 to 11 sets of microsatellite markers to detect the genetic structure of local dogs in South Korea and established an individual and paternity identification system to perform genetic analysis of two local breeds of South Korea and three foreign dog breeds. The results showed that the average H
O for each breed was between 0.65 and 0.78; the average PIC was between 0.66 and 0.82; and the average PE was more than 99% in all breeds. The average H
O value (0.57) of the 14 new microsatellite markers in this study near the range of the research results of the previous countries (0.60 to 0.74). The average value of PIC (0.61) was also within the range of the results of the aforementioned countries (0.56 to 0.82). According to the researches before, the 14 new microsatellite markers developed in this study were highly polymorphic and suitable to analyze the three breeds of dogs in Taiwan.
In conclusion, using 14 novel microsatellite markers to analyze the beagle, bichon, and schnauzer populations in Taiwan, the results showed that their average expected heterozygosity, observation heterozygosity, and polymorphism information content were all at high levels. Therefore, these new microsatellite markers have high applicability to the analyzed populations. These results indicate that the new microsatellite markers have good resolution when applied to the detection of differences among dog breeds. It was confirmed that the opportunity of identifying the exact same genotype among the analysis of the 14 new microsatellite markers was very low. In addition, the power exclusion was enough high to be a good tool for paternity testing.










 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Supplement
Supplement Print
Print





