 |
 |
| Anim Biosci > Volume 34(2); 2021 > Article |
|
Abstract
Objective
Stress-induced cytotoxicity caused by xenobiotics and endogenous metabolites induces the production of reactive oxygen species and often results in damage to cellular components such as DNA, proteins, and lipids. The cytochrome P450 (CYP) family of enzymes are most abundant in hepatocytes, where they play key roles in regulating cellular stress responses. We aimed to determine the effects of the antioxidant compound, methylsulfonylmethane (MSM), on oxidative stress response, and study the cytochrome P450 family 3 subfamily A (CYP3A) gene expression in fetal horse hepatocytes.
Methods
The expression of hepatocyte markers and CYP3A family genes (CYP3A89, CYP3A93, CYP3A94, CYP3A95, CYP3A96, and CYP3A97) were assessed in different organ tissues of the horse and fetal horse liver-derived cells (FHLCs) using quantitative reverse transcription polymerase chain reaction. To elucidate the antioxidant effects of MSM on FHLCs, cell viability, levels of oxidative markers, and gene expression of CYP3A were investigated in H2O2-induced oxidative stress in the presence and absence of MSM.
Results
FHLCs exhibited features of liver cells and simultaneously maintained the typical genetic characteristics of normal liver tissue; however, the expression profiles of some liver markers and CYP3A genes, except that of CYP3A93, were different. The expression of CYP3A93 specifically increased after the addition of H2O2 to the culture medium. MSM treatment reduced oxidative stress as well as the expression of CYP3A93 and heme oxygenase 1, an oxidative marker in FHLCs.
The liver plays a vital role in detoxification of xenobiotics such as carcinogens, environmental toxins, and drugs. The cytochrome P450 (CYP) family is a superfamily of heme-containing monooxygenases involved in the metabolism of approximately 80% of all drugs, xenobiotics, and endogenous metabolites in the liver [1]. CYP genes are classified into families and subfamilies according to their amino acid sequence identities. Enzymes that share more than 40% sequence identity belong to the same family, whereas those that share more than 55% sequence identity belong to the same subfamily [2]. CYP1, CYP2, and CYP3 are involved in the metabolism of numerous compounds and their expression is influenced by various factors such as xenobiotics, cytokines, and hormones, as well as genetics, disease, sex, and age [3]. CYP3A is expressed in many tissues, including the liver, intestines, gastric system, kidneys, lungs, adrenal glands, olfactory system, skin, prostate, and brain [4]. Functional studies have shown that each CYP3A isoform functions in a substrate-specific manner [5,6]. In the CYP3A subfamily, CYP3A4 is the most abundant isoform found in hepatocytes and plays a key role in the metabolism of drugs including steroids [7]. Several CYP3A genes have been annotated in the horse genome, for example, CYP3A89, CYP3A93, CYP3A94, CYP3A95, CYP3A96, and CYP3A97, which exhibit high amino acid similarities with the human CYP3A4 gene [8]. However, there is limited information on the involvement of horse CYP3As in substrate metabolism.
Reactive oxygen species (ROS), for example, hydrogen peroxide (H2O2), hydroxyl radicals, and superoxide anions that are generated by CYP metabolic processes, can induce and intensify oxidative stress in the liver [9]. The rate of H2O2 generation varies depending on the intrinsic properties of each CYP enzyme [10]. Excess ROS not only mediate cellular toxicity but also activate endogenous antioxidant mechanisms [11]. One of these endogenous mechanisms involves the translocation of NF-E2-related factor 2 (NRF2) from the cytosol to the nucleus during the transcription of antioxidant response-related genes and subsequent transcriptions of other genes such as NADPH:quinone oxidoreductase-1 (NQO1), NQO2, and heme oxygenase 1 (HO-1) [12,13].
Many natural substances have been screened and studied for the development of antioxidant supplements. Methylsulfonylmethane (MSM) is a small molecule containing sulfur and a methyl group and is found in fruits, vegetables, grains, and milk [14]. MSM restores the activities of catalase, superoxide dismutase, glutathione reductase, and glutathione S-transferase, and can directly or indirectly reduce oxidative stress [15]. Therefore, MSM has historically been used in the treatment of inflammatory disorders such as arthritis, interstitial cystitis, acute allergic rhinitis, exercise-induced inflammation, autoimmune disease, and cancer [14].
Considering the limited number of studies on stress-induced metabolism in horse liver in vivo, alternative methods such as the use of in vitro systems are needed to study and confirm the effects of natural substances. In this study, we used fetal horse liver derived cells (FHLCs) in vitro and investigated the effects of MSM on the level of cellular oxidative stress induced by H2O2, to improve current understanding of hepatic oxidative stress responses.
The experimental design was approved by the Pusan National University Institutional Animal Care and Use Committee (Approval Number: PNU-2015-0864). Tissue samples were collected via biopsies of the cerebrum, spinal cord, lungs, heart, liver, kidneys, and cecum of slaughtered horses.
FHLCs were kindly provided by Professor Tae Sub Park from Seoul National University. The liver tissue of a 7-month-old fetal Jeju horse was used in this study. Cells were cultured in DulbeccoŌĆÖs modified EagleŌĆÖs medium (Gibco, Grand Island, NY, USA) supplemented with 10% fetal bovine serum (Invitrogen, Carlsbad, CA, USA) and 1% antibioticŌĆōantimycotic solution (Invitrogen, USA) at 37┬░C in a 5% CO2 incubator. After 3 to 4 days of incubation, at 80% confluence, cells were treated with various concentrations of H2O2 (Junsei, Tokyo, Japan) either in the presence or absence of MSM (Merck, Darmstadt, Germany). MSM was mixed with H2O2 in the medium and stored at 4┬░C for 1 h before treatment. Then, the cells were incubated for 4 h and washed twice with 1├Ś phosphate-buffered saline (PBS) prior to RNA isolation.
Total RNA was isolated using TRIzol reagent (Invitrogen, Karlsruhe, Germany), according to the manufacturerŌĆÖs instructions. One microgram of RNA from each sample was used for reverse transcription with the SuperScript III First-Strand Synthesis System (Invitrogen, Germany). PRIMER3 software was used to design the primer sets. Information on the primers used in this study is provided in Table 1.
NCBI (http://www.ncbi.nlm.nih.gov) and the Ensembl Genome Browser (www.ensembl.org) were used to retrieve gene sequence information. The primers for amplification of the genes (Table 1) were designed using PRIMER3 software. Quantitative reverse transcription-polymerase chain reaction (qRT-PCR) was performed using a thermal cycler (C1000 Thermal Cycler; Bio-Rad, Hercules, CA, USA) to measure the relevant expression of target genes in a 20 ╬╝L reaction volume composed of 2 ╬╝L diluted cDNA (20 ng/╬╝L), 14 ╬╝L SYBR Green Master Mix (Bio-Rad, USA), and 1 ╬╝L each of diluted 5 pmol/╬╝L forward and reverse primers. The qRT-PCR cycling conditions were as follows: initial denaturation at 94┬░C for 10 min, followed by 40 cycles of denaturation at 94┬░C for 30 s, annealing at 65┬░C for 30 s, and extension at 72┬░C for 30 s. All measurements were performed in triplicate, and the 2ŌłÆ╬ö╬öCt method was used to determine relative gene expression. Relative expression of the target genes was normalized to the expression of glyceraldehyde-3-phosphate dehydrogenase.
Cell viability was assayed by measuring the amount of blue formazan generated from 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT; Sigma-Aldrich, St. Louis, MO, USA) by the activity of mitochondrial dehydrogenases. The cells were resuspended in the medium one day before H2O2 treatment at a density of 2├Ś105 cells per well in 24-well culture plates. The control treatment consisted of a culture plate in which the medium was replaced with fresh medium containing dimethyl sulfoxide (DMSO). Cells were incubated with various concentrations of H2O2 (Junsei, Japan) for 4 h in either the presence or absence of MSM. MTT (0.5 mg/mL) was added to each well and incubated for 4 h at 37┬░C. The formazan product was dissolved by adding 200 ╬╝L DMSO to each well, and the absorbance was measured at 570 nm using a microplate reader (Tecan US Inc., Durham, NC, USA). All measurements were performed in triplicate and repeated at least three times.
H2O2-stimulated and unstimulated FHLCs were incubated in the presence or absence of MSM (100 mM or 200 mM) at 37┬░C for 4 h. The medium was removed, and 10 ╬╝M dihydroethidium (DHE; Sigma-Aldrich, USA) was added to the cells and incubated for an additional 1 h at 37┬░C. Then, the solution was removed, and the FHLCs were detached using 0.05% trypsin-ethylenediaminetetraacetic acid. Cells were washed with 1├Ś PBS, resuspended in FACS buffer, and analyzed using a Muse Cell Analyzer (Merck, Germany).
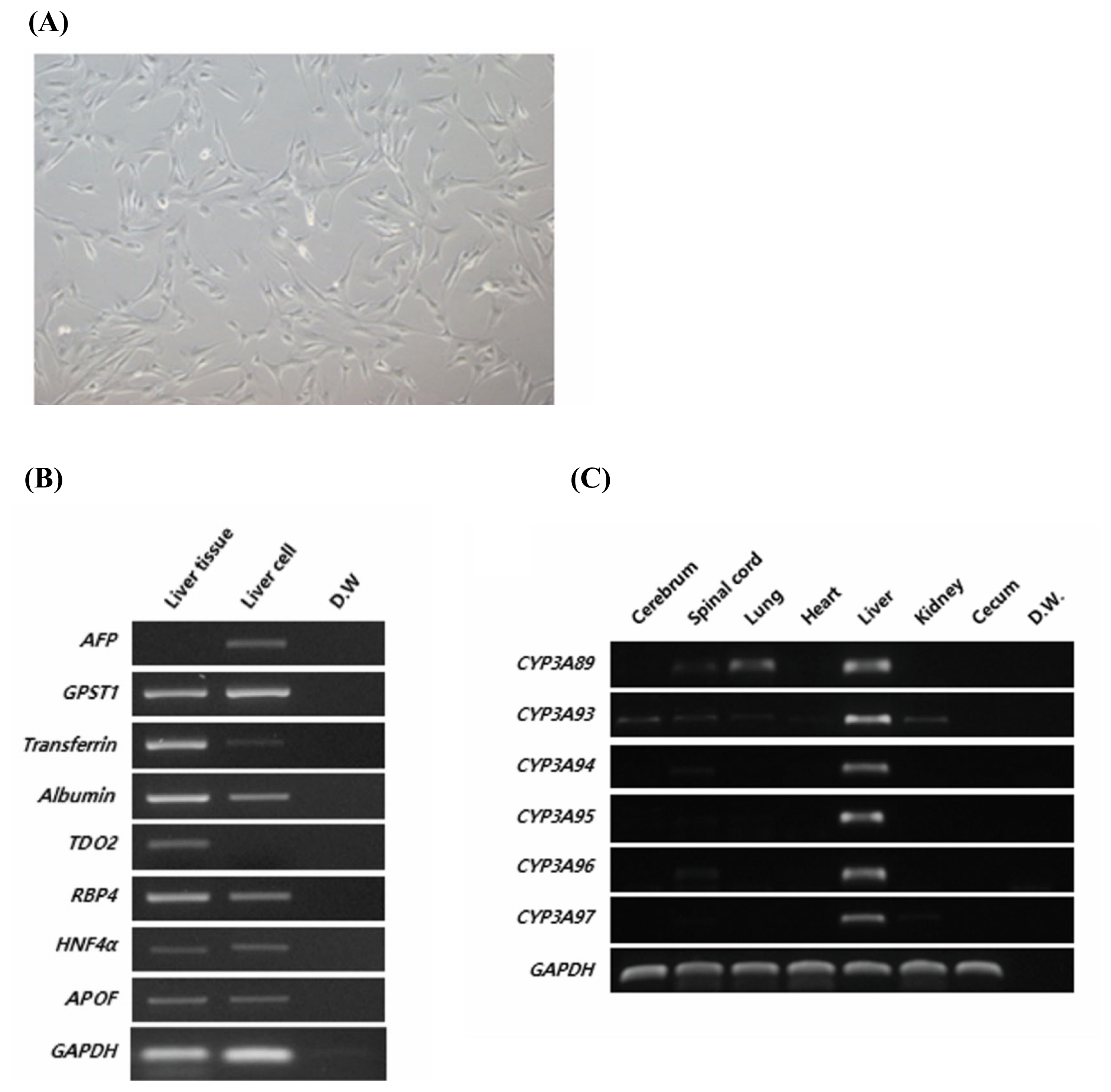
We used FHLCs derived from the liver of a 7-month-old fetal Jeju horse (Figure 1A) and studied the expression of liver-specific markers (Figure 1B) to verify the lineage of cells. Fetal liver markers such as ╬▒-fetoprotein and glutathione S-transferase P were highly expressed in FHLCs, whereas the expression of liver markers such as transferrin and albumin, typically observed in livers of adult horses, were reduced. Other markers such as retinol-binding protein 4 (RBP4), hepatocyte nuclear factor 4╬▒ (HNF4╬▒), and apolipoprotein F (APOF) were expressed at similar levels in both FHLCs and liver tissue in adult horses. It is noteworthy that tryptophan 2,3-dioxygenase was expressed in the adult liver but not in FHLCs.
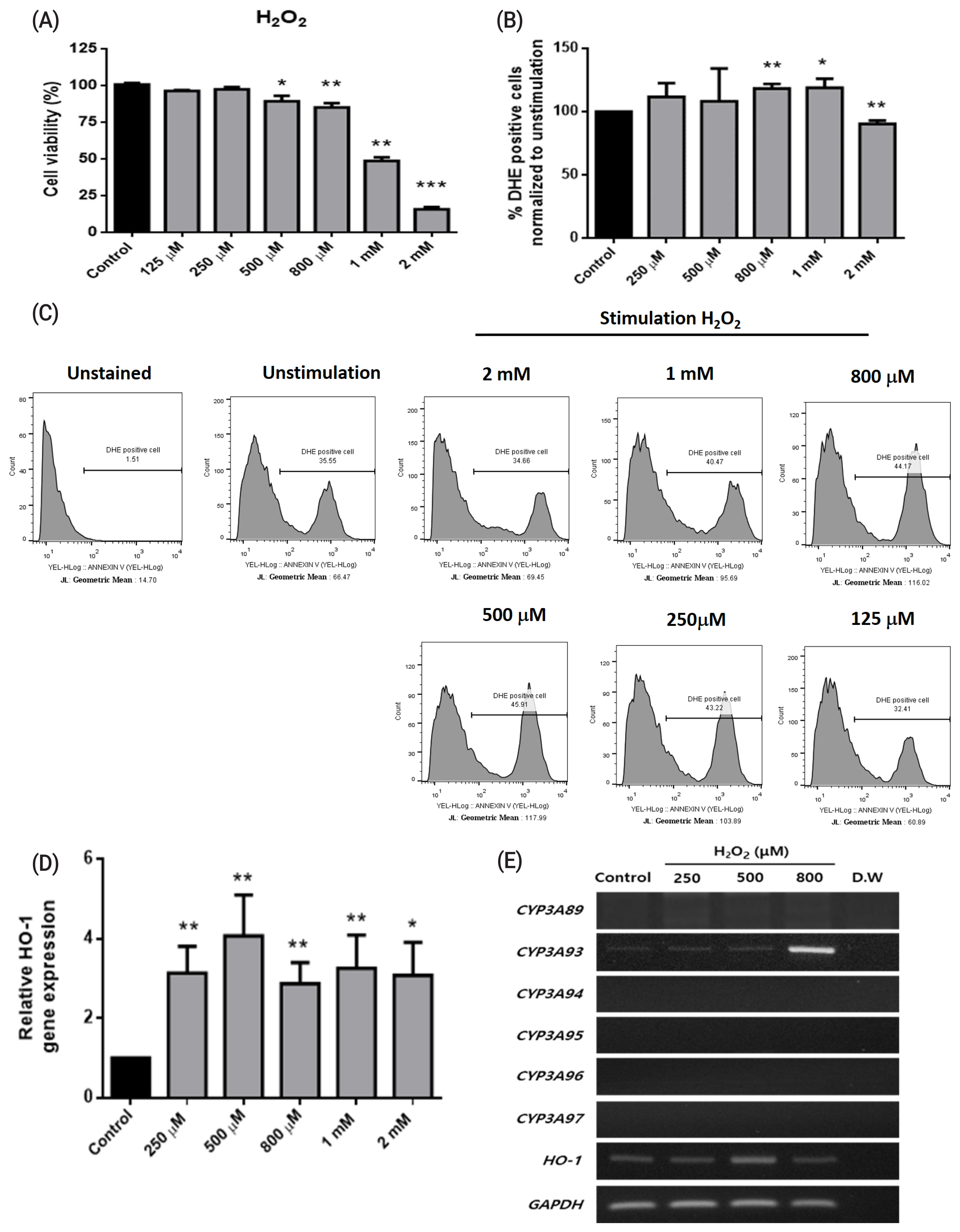
To determine the expression patterns of the CYP3A gene family in horse, we designed specific primer sets to detect six CYP3A genes, namely, CYP3A89, CYP3A93, CYP3A94, CYP3A95, CYP3A96, and CYP3A97, in horse liver tissues (Table 1). All CYP3A genes were strongly expressed in liver tissue (Figure 1C). CYP3A89 was weakly expressed in the spinal cord and lungs, and CYP3A93 was differentially expressed in the cerebrum, spinal cord, liver, and kidneys. These results demonstrated that the CYP3A gene family was specifically expressed in liver tissues. In contrast to the adult liver tissue in which all CYP3A genes were expressed, only the expression of CYP3A93 was detected in FHLCs (Figure 2E). In addition, the expression of the CYP3A gene family in FHLCs under oxidative stress was analyzed; the expression of CYP3A93 was upregulated, but the expression of other CYP3A genes was not detected (Figure 2E).
Schmitz et al [8] annotate seven CYP3A genes in the horse genome and report that CYP3A128P is considered a pseudogene, while CYP3A129 is expressed at undetectable levels in the liver. Previous studies have investigated the gene expression of CYP isozymes in horse liver tissues [13,16] and attempted functional characterization of these proteins [17, 18]. In this study, we observed that only CYP3A93 was expressed in FHLCs, whereas all CYP3A genes were expressed in the adult liver of the horse. One possible explanation for the differential expression CYP3A genes and liver-specific genes in horse, e.g., RBP4, HNF4╬▒, and APOF, may be attributed to the differentiation and maturation of hepatocytes. Zabulica et al [19] performed comparative experiments to provide a reliable guide for evaluating the differentiation of stem cells to hepatocyte-like cells using 60 genes expressed in the normal liver; they used a sample set of 17 fetal and 25 mature livers and revealed that the expression of CYP3A genes and other liver-marker genes are attributed to age [19]. Other studies have also shown that CYP3A mRNA expression follows a pattern very similar to that of CYP3A as revealed using immunostaining [20,21].
To establish oxidative stress conditions, FHLCs were treated with different concentrations (125 ╬╝M, 250 ╬╝M, 500 ╬╝M, 800 ╬╝M, 1 mM, and 2 mM) of H2O2 for 4 h. Significant cytotoxicity was observed at concentrations more than 500 ╬╝M, and cell viability was significantly decreased after treatment with 1 mM and 2 mM H2O2 (p<0.05, Figure 2A). We performed DHE staining to measure the cellular state of oxidative stress; DHE used as a superoxide anion probe was oxidized to a fluorescent product (excitation, 480 nm; emission, 567 nm) [22]. DHE-positive cells, which were calculated using geometric means, were significantly increased at concentrations more than 800 ╬╝M and 1 mM H2O2, indicating that FHLCs treated with H2O2 were subjected to oxidative stress. We noted that the DHE positivity of FHLCs was significantly decreased at 2 mM H2O2, probably because of cell death (Figure 2B and Figure 2C). HO-1 is one of the downstream genes of CYP3A93, and HO-1 expression can be used as a marker of oxidative stress [13]. In FHLCs, HO-1 expression significantly increased after treatment with H2O2 at concentrations ranging from 250 ╬╝M to 2 mM (Figure 2D).
To determine the effects of oxidative stress on the expression of CYP3A genes, FHLCs were treated with different concentrations of H2O2 (250, 500, and 800 ╬╝M). We found that, among the six CYP3A genes, only CYP3A93 was upregulated under oxidative conditions and the highest expression was detected after treatment with 800 ╬╝M H2O2 (Figure 2E). For CYP3A93 and other CYP3A members, mRNA expression level and protein level were highly correlated in the horse liver and intestine [23]. It is reasonable to assume that oxidative stress upregulated the mRNA of CYP3A93 in FHLCs and in turn led to an increase in CYP3A93 proteins as well as CYP3A93 enzymatic activity. Nonetheless, the relationship between mRNA level and protein synthesis of horse CYP3A93 under oxidative stress in FHLCs needs to be studied further.
We tested whether the well-known antioxidant MSM could ameliorate oxidative stress in FHLCs and regulate CYP3A93 expression. We found that the MSM (12.5, 25, 50, 100, and 200 mM) did not change the viability of FHLCs (Figure 3A). Next, we examined the effects of MSM in FHLCs under oxidative stress by exposure to 800 ╬╝M H2O2. Treatment with MSM did not restore the viability of FHLCs that was reduced due to oxidative stress (Figure 3B). Although low concentrations of MSM did not ameliorate oxidative stress, treatment with 200 mM MSM clearly reduced the percentage of DHE-positive cells (Figure 3C and 3D), suggesting that MSM blocked ROS generation under oxidative stress. It is noteworthy that MSM had the capacity to suppress ROS generation; however, it did not improve cellular viability. This finding suggested that MSM affected other pathways to regulate cell death and suppress ROS generation. Further studies are required to determine these involved pathways.
Next, we investigated the effects of MSM on HO-1 and CYP3A93 expression after treatment with 800 ╬╝M H2O2. CYP3A93 expression decreased significantly after treatment with 100 and 200 mM MSM (Figure 4A, 4B). In addition, HO-1 showed an expression pattern similar to that of CYP3A93 (Figure 4A, 4C). These results indicated that CYP3A93 expression could be used as a marker to determine oxidative stress in FHLCs. Nonetheless, it was unclear how MSM regulated CYP3A93 expression and why reduced CYP3A93 expression was correlated with reduced ROS generation in FHLCs. A previous study reveals that MSM reduces cortisol-induced stress through the expression of p53-mediated succinate dehydrogenase complex flavoprotein subunit A/hypoxanthine phosphoribosyltransferase 1 in racehorse muscle [24]. In addition, MSM has been shown to increase the expression of antioxidant enzymes such as peroxiredoxin-1, thioredoxin-1, and HO-1, which are normally induced by the activation of Nrf2 [25]. MSM can inhibit the transcriptional activity of NF-╬║B during ROS production [14]. Further studies are required to determine whether these pathways regulate MSM-mediated CYP3A93 suppression and ROS reduction in FHLCs.
In this study, we investigated the expression of horse CYP3A genes in liver tissues of horse and FHLCs and found that CYP3A93 was expressed in FHLCs. CYP3A93 expression increased under oxidative conditions; however, the expression of other CYP3A genes did not increase. In FHLCs, MSM reduced ROS generation without affecting cell viability and suppressed the expression of CYP3A93 and HO-1; these findings suggested that MSM can be used to decrease oxidative stress-induced hepatotoxicity in horses. Further investigation is required to clarify the mechanisms underlying these effects in FHLCs.
ACKNOWLEDGMENTS
This study was supported by grants from the Individual Basic Science & Engineering Research Program (2017R1D1A1B0 3036432), National Research Foundation of Korea (NRF), Next-Generation BioGreen 21 Program (Project No. PJ01325 701), Rural Development Administration, Republic of Korea, and Research Base Construction Fund Support Program funded by Jeonbuk National University in 2020.
Figure┬Ā1
Characterization of horse liver-derived cells and validation of CYP3A family genes in different tissues of horse. (A) Morphology of horse liver-derived cells. Scale bar: 50 ╬╝m. (B) Liver marker expression analyzed using RT-PCR in horse liver tissue and liver-derived cells. GAPDH was used as the reference gene. (C) Expression of CYP3A genes in different horse tissues, namely, cerebrum, spinal cord, lungs, heart, liver, kidneys, and cecum, was analyzed using RT-PCR. GAPDH was used as the reference gene. The data are presented as one of three independent experiments. CYP3A, cytochrome P450 family 3 subfamily a; RT-PCR, reverse transcription-polymerase chain reaction; AFP, ╬▒-fetoprotein; GSTP1, glutathione S-transferase P; TDO2, tryptophan 2,3-dioxygenase; RBP4, retinol-binding protein 4; HNF4╬▒, hepatocyte nuclear factor 4╬▒; APOF, apolipoprotein F; GAPDH, glyceraldehyde 3-phosphate dehydrogenase; CYP, cytochrome P450.
Figure┬Ā2
Effects of oxidative stress on horse liver-derived cells. (A) Cell viabilities measured using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. (B) Effects of H2O2 treatment on the generation of reactive oxygen species analyzed using dihydroethidium (DHE) staining. (C) Histogram analysis of DHE-positive cells and geometric expression. (D) HO-1 gene expression analyzed using real-time polymerase chain reaction (PCR). Black and gray bars represent data from control and H2O2 treatment, respectively. The data are presented as means┬▒standard deviations of three independent experiments (* p<0.05, ** p<0.01, and *** p<0.001; unpaired StudentŌĆÖs t-test). (E) RT-PCR of CYP3A family gene expression after H2O2 treatment at 250, 500, and 800 ╬╝M for 4 h. Data are presented as one of three independent experiments. CYP, cytochrome P450; HO-1, heme oxygenase-1; GAPDH, glyceraldehyde 3-phosphate dehydrogenase.
Figure┬Ā3
Effects of methylsulfonylmethane (MSM) on cell viability and reactive oxygen species generation in horse liver-derived cells under oxidative stress. (A) Percentage cell viability with MSM treatment at different concentrations (12.5, 25, 50, 100, and 200 mM). (B) Percentage cell viability with 100 or 200 mM MSM under oxidative stress conditions (800 ╬╝M H2O2). Reactive oxygen species levels were determined using dihydroethidium (DHE) staining and analyzed using flow cytometry. (C) Black and white histograms represent DHE-negative and DHE-positive cells, respectively. (D) Black, grey, and white bars represent the percentage of DHE-positive cells in the presence or absence of MSM treatment (100 or 200 mM) under oxidative stress (800 ╬╝M H2O2). Data are presented as means┬▒standard deviations of three independent experiments (* p<0.05, ** p<0.01, and *** p<0.001; unpaired StudentŌĆÖs t-test). Cell viabilities were measured using 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay.

Table┬Ā1
Primer sets used in this study
REFERENCES
1. Guengerich FP. Cytochrome P450s and other enzymes in drug metabolism and toxicity. AAPS J 2006; 8:E101ŌĆō11.
https://doi.org/10.1208/aapsj080112




2. Nelson DR. Cytochrome P450 nomenclature, 2004. Methods Mol Biol 2006; 320:1ŌĆō10.
https://doi.org/10.1385/1-59259-998-2:1


3. Zanger UM, Schwab M. Cytochrome P450 enzymes in drug metabolism: regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacol Ther 2013; 138:103ŌĆō41.
https://doi.org/10.1016/j.pharmthera.2012.12.007


4. Ding X, Kaminsky LS. Human extrahepatic cytochromes P450: function in xenobiotic metabolism and tissue-selective chemical toxicity in the respiratory and gastrointestinal tracts. Annu Rev Pharmacol Toxicol 2003; 43:149ŌĆō73.
https://doi.org/10.1146/annurev.pharmtox.43.100901.140251


5. Schmitz A, Zielinski J, Dick B, Mevissen M.
In vitro metabolism of testosterone in the horse liver and involvement of equine CYPs 3A89, 3A94 and 3A95. J Vet Pharmacol Ther 2014; 37:338ŌĆō47.
https://doi.org/10.1111/jvp.12106


6. Nakayama SMM, Ikenaka Y, Hayami A, et al. Characterization of equine cytochrome P450: role of CYP3A in the metabolism of diazepam. J Vet Pharmacol Ther 2016; 39:478ŌĆō87.
https://doi.org/10.1111/jvp.12303


7. Liu YT, Hao HP, Liu CX, Wang GJ, Xie HG. Drugs as CYP3A probes, inducers, and inhibitors. Drug Metab Rev 2007; 39:699ŌĆō721.
https://doi.org/10.1080/03602530701690374


8. Schmitz A, Demmel S, Peters LM, Leeb T, Mevissen M, Haase B. Comparative human-horse sequence analysis of the CYP3A subfamily gene cluster. Anim Genet 2010; 41:Suppl 272ŌĆō9.
https://doi.org/10.1111/j.1365-2052.2010.02111.x


9. Deavall DG, Martin EA, Horner JM, Roberts R. Drug-induced oxidative stress and toxicity. J Toxicol 2012; 2012:645460. https://doi.org/10.1155/2012/645460


10. Mishin V, Heck DE, Laskin DL, Laskin JD. Human recombinant cytochrome P450 enzymes display distinct hydrogen peroxide generating activities during substrate independent NADPH oxidase reactions. Toxicol Sci 2014; 141:344ŌĆō52.
https://doi.org/10.1093/toxsci/kfu133




11. Zangar RC, Davydov DR, Verma S. Mechanisms that regulate production of reactive oxygen species by cytochrome P450. Toxicol Appl Pharmacol 2004; 199:316ŌĆō31.
https://doi.org/10.1016/j.taap.2004.01.018


12. Kensler TW, Wakabayashi N. Nrf2: friend or foe for chemoprevention? Carcinogenesis 2010; 31:90ŌĆō9.
https://doi.org/10.1093/carcin/bgp231



13. Loboda A, Damulewicz M, Pyza E, Jozkowicz A, Dulak J. Role of Nrf2/HO-1 system in development, oxidative stress response and diseases: an evolutionarily conserved mechanism. Cell Mol Life Sci 2016; 73:3221ŌĆō47.
https://doi.org/10.1007/s00018-016-2223-0




14. Butawan M, Benjamin RL, Bloomer RJ. Methylsulfonylmethane: applications and safety of a novel dietary supplement. Nutrients 2017; 9:290
https://doi.org/10.3390/nu9030290



15. Mohammadi S, Najafi M, Hamzeiy H, et al. Protective effects of methylsulfonylmethane on hemodynamics and oxidative stress in monocrotaline-induced pulmonary hypertensive rats. Adv Pharmacol Pharm Sci 2012; 2012:507278. https://doi.org/10.1155/2012/507278


16. Tyden E, Lofgren M, Hakhverdyan M, Tjalve H, Larsson P. The genes of all seven CYP3A isoenzymes identified in the equine genome are expressed in the airways of horses. J Vet Pharmacol Ther 2013; 36:370ŌĆō5.
https://doi.org/10.1111/jvp.12012


17. DiMaio Knych HK, DeStefano Shields C, Buckpitt AR, Stanley SD. Equine cytochrome P450 2C92: cDNA cloning, expression and initial characterization. Arch Biochem Biophys 2009; 485:49ŌĆō55.
https://doi.org/10.1016/j.abb.2009.02.009


18. DiMaio Knych HK, McKemie DS, Stanley SD. Molecular cloning, expression, and initial characterization of members of the CYP3A family in horses. Drug Metab Dispos 2010; 38:1820ŌĆō7.
https://doi.org/10.1124/dmd.110.032953


19. Zabulica M, Srinivasan RC, Vosough M, et al. Guide to the assessment of mature liver gene expression in stem cell-derived hepatocytes. Stem Cells Dev 2019; 28:907ŌĆō19.
https://doi.org/10.1089/scd.2019.0064



20. Lee SJ, Hedstrom OR, Fischer K, et al. Immunohistochemical localization and differential expression of cytochrome P450 3A27 in the gastrointestinal tract of rainbow trout. Toxicol Appl Pharmacol 2001; 177:94ŌĆō102.
https://doi.org/10.1006/taap.2001.9289


21. Cotreau MM, von Moltke LL, Beinfeld MC, Greenblatt DJ. Methodologies to study the induction of rat hepatic and intestinal cytochrome P450 3A at the mRNA, protein, and catalytic activity level. J Pharmacol Toxicol Methods 2000; 43:41ŌĆō54.
https://doi.org/10.1016/S1056-8719(00)00086-1


22. Zhao H, Kalivendi S, Zhang H, et al. Superoxide reacts with hydroethidine but forms a fluorescent product that is distinctly different from ethidium: potential implications in intracellular fluorescence detection of superoxide. Free Radic Biol Med 2003; 34:1359ŌĆō68.
https://doi.org/10.1016/S0891-5849(03)00142-4


23. Tyden E, Lofgren M, Pegolo S, Capolongo F, Tjalve H, Larsson P. Differential gene expression of CYP3A isoforms in equine liver and intestines. J Vet Pharmacol Ther 2012; 35:588ŌĆō95.
https://doi.org/10.1111/j.1365-2885.2012.01379.x


24. Sp N, Kang DY, Kim DH, et al. Methylsulfonylmethane inhibits cortisol-induced stress through p53-mediated SDHA/HPRT1 expression in racehorse skeletal muscle cells: a primary step against exercise stress. Exp Ther Med 2020; 19:214ŌĆō22.
https://doi.org/10.3892/etm.2019.8196


25. Withee ED, Tippens KM, Dehen R, Tibbitts D, Hanes D, Zwickey H. Effects of methylsulfonylmethane (MSM) on exercise-induced oxidative stress, muscle damage, and pain following a half-marathon: a double-blind, randomized, placebo-controlled trial. J Int Soc Sports Nutr 2017; 14:24
https://doi.org/10.1186/s12970-017-0181-z




- TOOLS
-
METRICS

- Related articles
-
Effects of exercise on myokine gene expression in horse skeletal muscles2019 March;32(3)






 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Download Citation
Download Citation Print
Print




