 |
 |
| Anim Biosci > Volume 33(9); 2020 > Article |
|
Abstract
Objective
Methods
Results
ACKNOWLEDGMENTS
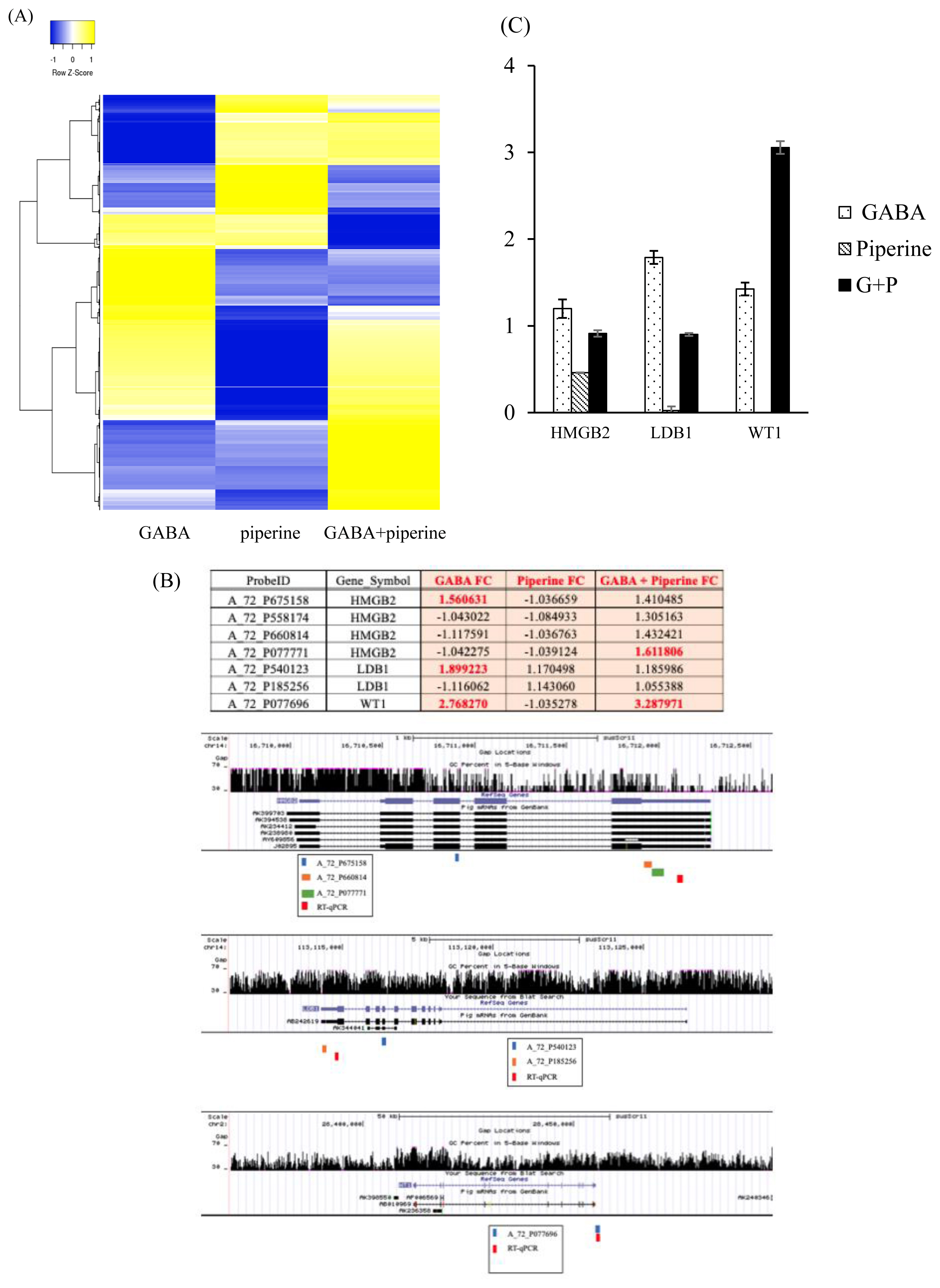
Figure┬Ā1
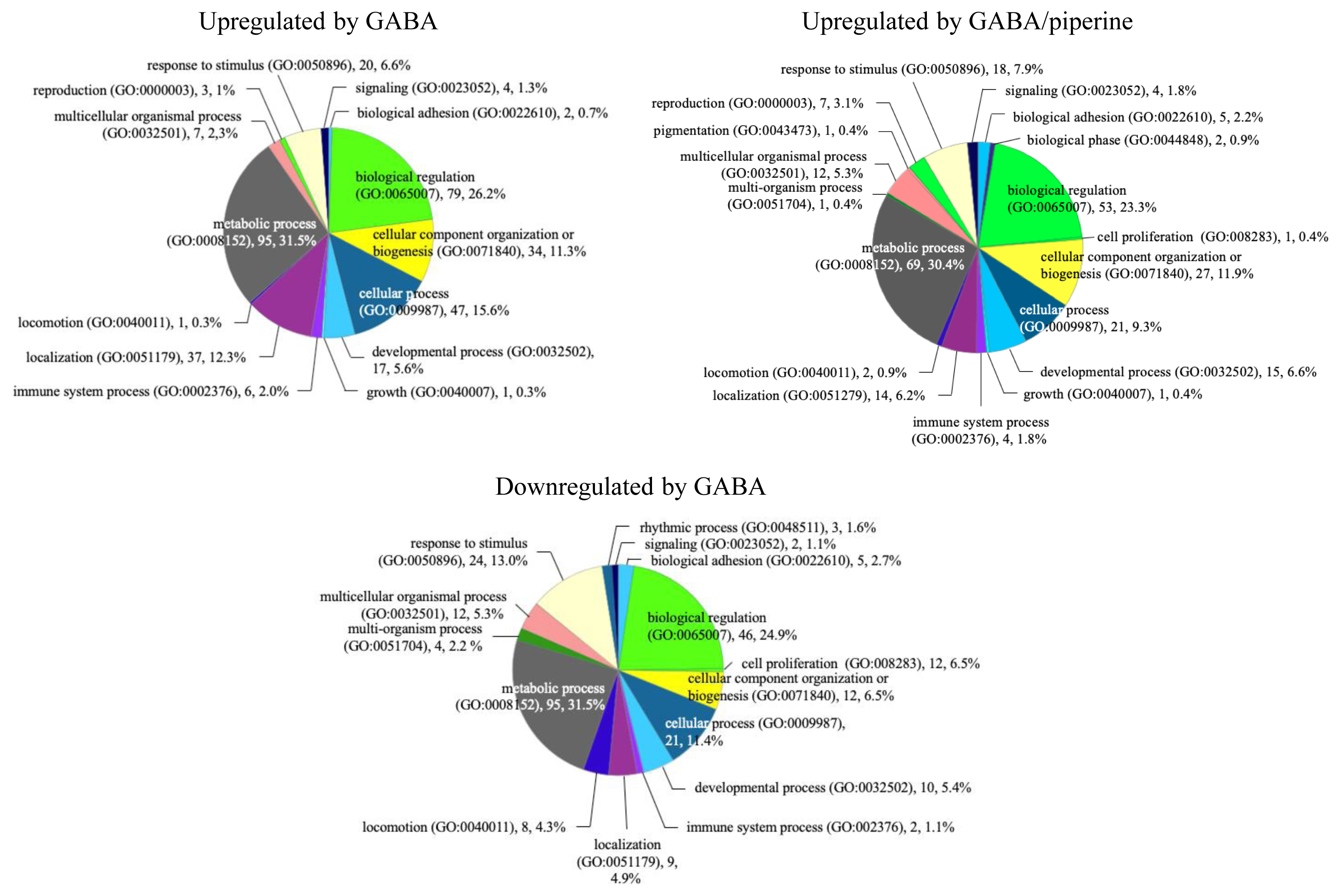
Figure┬Ā2
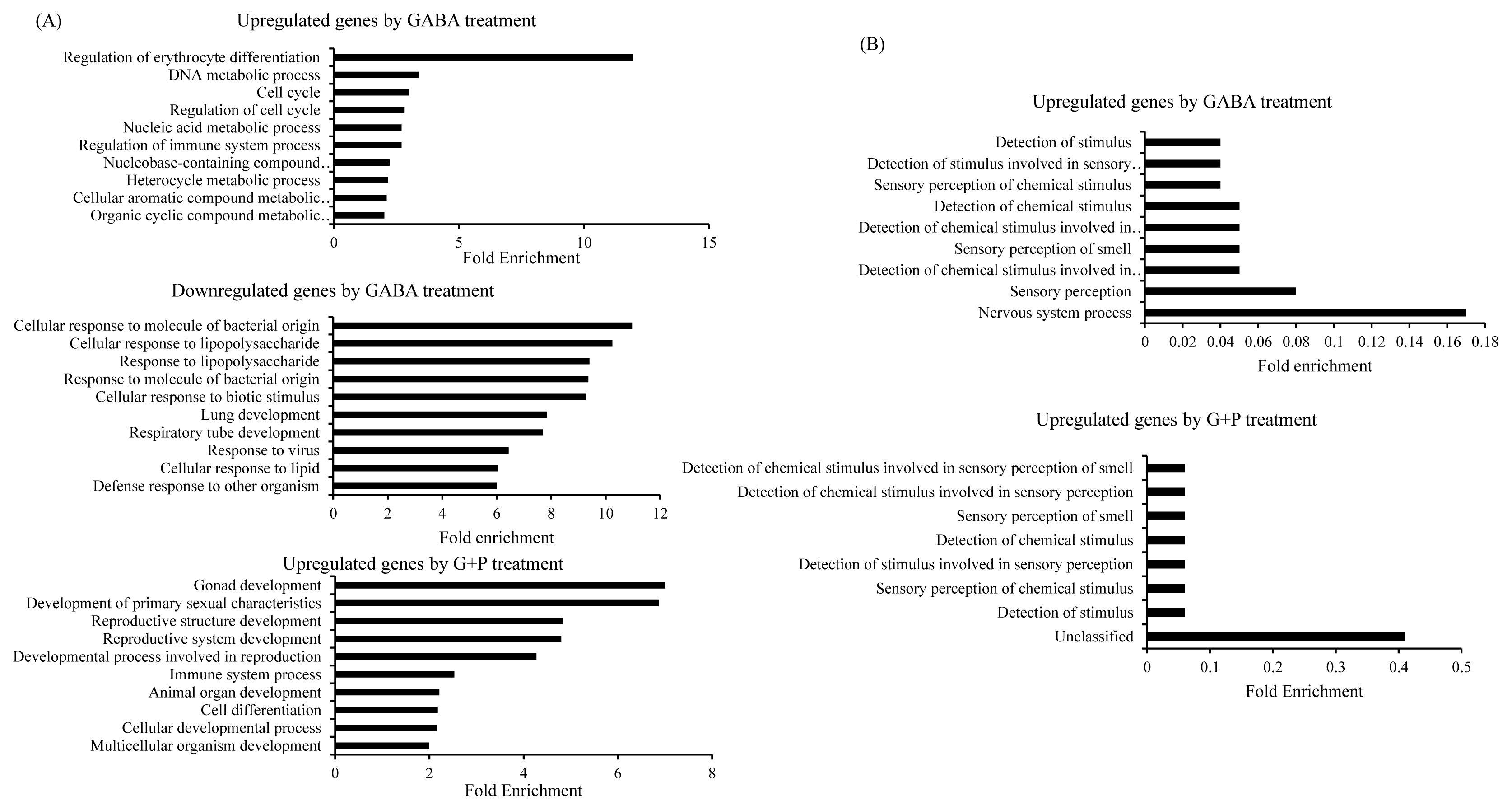
Figure┬Ā3
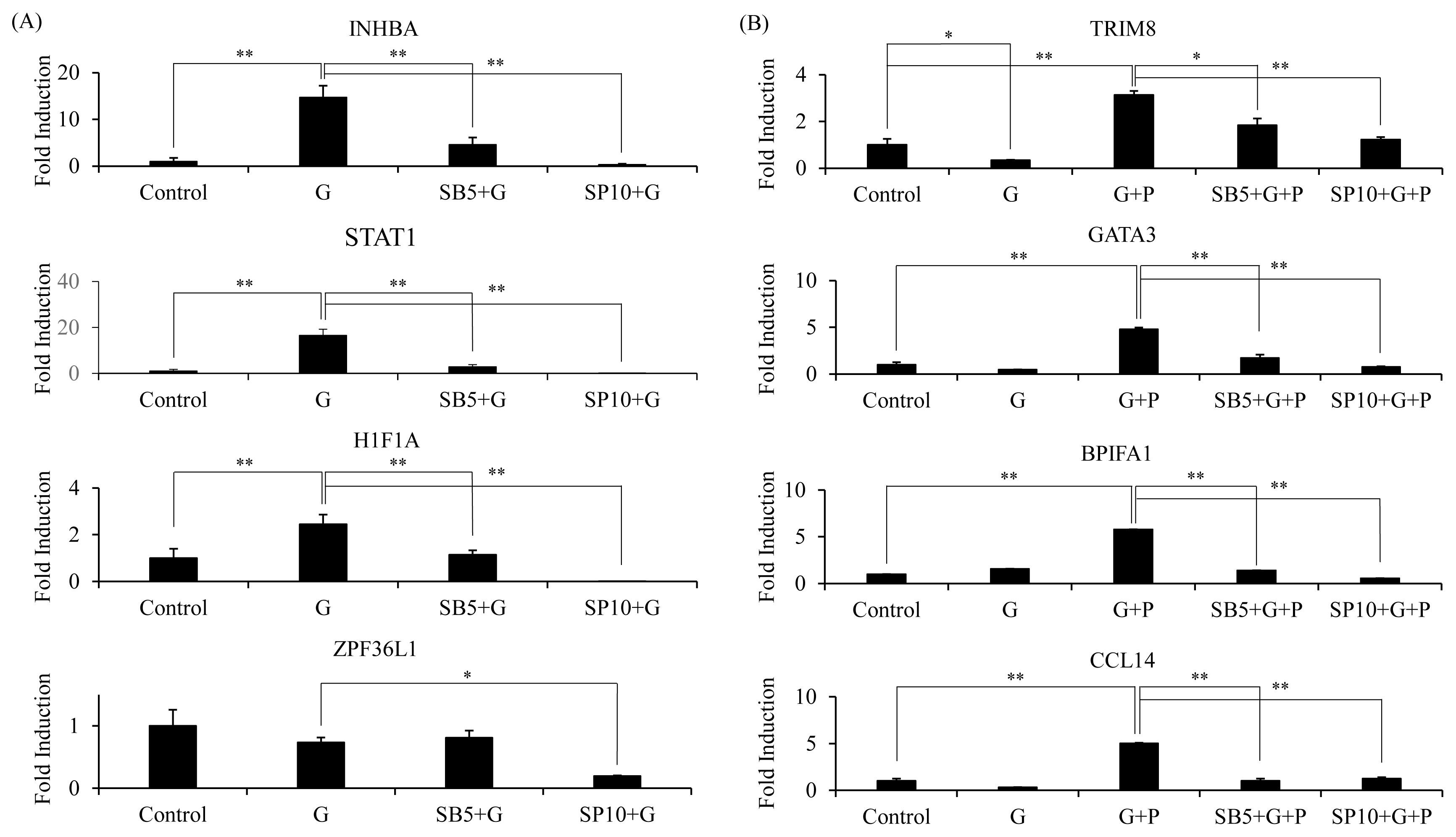
Figure┬Ā4
Table┬Ā1
Table┬Ā2
GenesŌĆÖ fold changes (FCs) compared to control are listed for GABA, PIP, and GABA/PIP (G+P) co-treatment.
GABA, gamma-aminobutyric acid; PIP, piperine; ZFP36L1, SPF36 ring finger protein like 1; STAT1, signal transducer and activator of transcription 1; HIF1A, hypoxia-inducible factor 1-alpha; HMGB2, high mobility group protein B2; LDB1, LIM domain-binding protein 1; INHBA, inhibin subunit beta A; DEFB1, defensin beta 1; TRIM8, tripartite motif containing 8; ADAM15, ADAM metallopeptidase domain 15; TEC, Tec protein ty-rosine kinase; CAV1, caveolin 1; PLCL2, phospholipase C like 2; CD3E, CD3e molecule; GATA3, GATA binding protein 3; CCNB2, cyclin B2; ARID4A, AT-rich interaction domain 4A; C4A, complement C4A (Rodgers Blood Group); TMEM173, transmembrane protein 173; BPIFA1, BPI fold containing family A member 1; TREM2, triggering receptor expressed on myeloid cells 2; ENPP2, ectonucleotide pyrophosphatase/phosphodiesterase 2; ITM2A, integral membrane protein 2A; RAB29, ras-related protein Rab-7L; OAS2, 2ŌĆ▓-5ŌĆ▓-oligoadenylate synthetase 2; IRAK1, interleukin-1 receptor-associated kinase 1; THRA, thyroid hormone receptor alpha; TOP2A, DNA topoisomerase II alpha; SLA-DOA, major histocompatibility complex, classII Do alpha; KMT2E, myeloid/lymphoid or mixed-lineage leukemia 5; REST, RE1-silencing transcription factor; GCNT1, glucosami-nyl (N-acetyl) transferase 1; CCL14, C-C motif chemokine ligand 14.
REFERENCES
- TOOLS
-
METRICS

- Related articles
-
Effects of exercise on myokine gene expression in horse skeletal muscles2019 March;32(3)






 PDF Links
PDF Links PubReader
PubReader Full text via DOI
Full text via DOI Download Citation
Download Citation Print
Print




