 |
 |
| Anim Biosci > Volume 30(7); 2017 > Article |
|
Abstract
Objective
The bursa of Fabricius (BF) is a central humoral immune organ belonging specifically to avians. Recent studies had suggested that miRNAs were active regulators involved in the immune processes. This study was to investigate the possible differences of the BF at miRNA level between two genetically disparate duck breeds.
Results
The results showed that there were 66 differentially expressed miRNAs and 28 novel miRNAs in bursa. A set of abundant miRNAs (i.e., let-7, miR-146a-5p, miR-21-5p, miR-17~92) which are involved in immunity and disease were detected and the predicted target genes of the novel miRNAs were associated with duck high anti-adversity ability. By gene ontology analysis and enriching KEGG pathway, the targets of differential expressed miRNAs were mainly involved in immunity and disease, supporting that there were differences in the BF immune functions between the two duck breeds. In addition, the metabolic pathway had the maximum enriched target genes and some enriched pathways that were related to cell cycle, protein synthesis, cell proliferation and apoptosis. It indicted that the difference of metabolism may be one of the reasons leading the immune difference between the BF of two duck breeds.
Compared with mammals, avains have a specific central humoral immune organ which is named the bursa of Fabricius (BF) and plays an important role in immune resistance [1,2]. In 1621, the bursa was originally described by Hieronymus Fabricius [3]. Subsequent researches have shown that BF is involved in the occupation, proliferation, differentiation and maturity of B lymphocytes which are one of the most significant immune cells that can produce immunoglobulin (Ig) when the body is stimulated by antigens [4,5].
MicroRNAs (miRNAs), as functional and small non-coding (~22nt) RNA, are essential regulators for gene expression [6]. miRNAs play a critical role in various aspects of the immune system. In one aspect, miRNAs exist in the immune organs among cells that are proliferation, differentiation and dying [7]. For instance, through establishing and comparing small RNA libraries derived from the bursa and spleen of embryonic chicks at days 15 and 20, Hicks et al [8] found that many identified differentially expressed miRNAs displayed dynamic expression patterns and some of them can regulate lymphocyte proliferation and apoptosis, such as miR-221 and miR-17-5p [8]. Treating chicken embryos lymphocytes with anti-miR-143 in vitro, Trakooljul et al [9] detected expression changes of downstream target genes and found that miRNA-143 may be involved in the regulation of lymphocyte proliferation and apoptosis [9]. Chen et al [10] found that the miRNA-181-a expressed selectively in B cells of thymus and can lead to B cell proliferation through adaptive special expression in hematopoietic stem cells of mice [10]. On the other aspect, miRNAs are involved in the regulation of the immune response. For example, 79 miRNAs that showed distinct expressions between marekâs disease virus infected-chicken splenic tumors and non-tumorous spleen tissues were identified by microarray [11]. Wang et al [12] demonstrated that gga-miRNA-21 can inhibit replication of infectious bursal virus through down-regulating of infectious bursal disease virus viral structural protein viral protein 1 at translation level [12]. The miRNA-155 participated in the reaction that B cells produce high affinity antibodies and memory cell [13,14]. Although some miRNAs have been identified in chicken BF, and they were subsequently revealed to be involved in BF immunity processes [8], there is still much to investigate regarding the function of conserved miRNAs and novel miRNAs in poultry BF.
Immune resistance to disease can be greatly influenced by the genetic background. It had been demonstrated that there were differentially expressed miRNAs between two genetically disparate chicken lines after inducing virus, and further studies demonstrated that different disease phenotypes led to the identification of host immunity genes regulated by miRNAs [15,16]. Therefore, the present study aimed to distinguish conservative and differently expressed miRNAs of BF resistance in ducks by enriching the differentially expressed miRNAs in BF between two genetically different duck breeds through next generation sequencing technology.
Jianchang duck (JCH) and Nonghua P-strain meat duck (NH-P) were two different duck breeds being reared in Waterfowl Breeding Experimental Farm at Sichuan Agricultural University. The two breeds of duck had the same nutritional standards and environmental conditions. Food and water were available ad libitum. Six randomly selected JCH and NH-P were euthanized at week 3, 4, and 5. Then bursa of Fabricius was separated from adhering tissues and all samples were frozen in liquid nitrogen and stored at â80°C. Tissue samples of six individuals in each duck breed were pooled and partial pooled samples of week 4 were sent to sequence. Total RNA of all samples was extracted using trizol (Invitrogen, Carlsbad, CA, USA), following the manufacturerâs instructions. All procedures of the entire experiment were approved by the Animal Ethics Committee of Sichuan Agricultural University.
RNA integrity of the two samples was confirmed using Agilent 2100. Two sRNA libraries (bursa of Fabricius in JCH and NH-P) were constructed using Small RNA Sample Pre Kit (Illumina, San Diego, CA, USA) following the manufacturerâs instructions. Making use of the special structure of sRNA that 5Ⲡhas the integrated phosphate group and 3Ⲡhas hydroxyl, the total sRNA was ligated to 3Ⲡand 5Ⲡadapters using T4 RNA ligase. Subsequently, reverse transcription reactions were performed using the reverse transcription (RT) primer, and polymerase chain reaction (PCR) were performed using the forward and reverse Illumina primers. The PCR products were purified by gel electrophoresis (PAGE) and the separated DNA fragments were the cDNA libraries. Then the library preparations were sequenced on Illumina Hiseq 2000 platform.
Through base calling, the original image data of sequencing was analyzed into the sequenced reads, they were called the raw reads. In order to obtain the clean reads, we filtered out the reads of low-quality, and the reads with the percentage of N more than 10%. In addition, reads were removed with 5Ⲡprimer contaminants or without 3Ⲡprimer insert fragments. After trimming the 3Ⲡadaptors, removing the reads including polyA, poly T, poly C, and poly G, the clean reads were mapped duck genome database (BGI duck 1.0 reference Annotation Release 101) and compared with the distribution of the known miRNAs. The novel miRNAs were predicted by the software miREvo and mirdeep2 [17,18]. With the statistics and normalization of the expression of miRNAs in two samples through transcript per million (TPM, normalization expression = readCountĂ1,000,000)/libsize) [19], the differential expression of miRNAs were compared by DGEseq in the two samples [20]. The 0.05 of p-value and log2 Fold-change (log2[JCH/NH-P]) were calculated to filter the significant differential expressions. GOseq Release 2.12 was used to analyze the gene ontology (GO) enrichment of differential expressed miRNAs. According to the relationship between miRNAs and target genes, we analyzed the distribution of target genes in GO through three categories (cellular component, biological process, molecular function). KEGG is the main database of pathway for understanding the high-level functions and utilities of biological system (http://www.genome.jp/kegg/). We used KOBAS v2.0 software to calculate the expression of the miRNA target genes in order to find which pathway was involved.
The total RNA were reversed transcribed into first-strand cDNA according to the manufacturerâs instructions of miRcute miRNA reverse transcription kit (TIANGEN, Beijing, China). Some significantly differentially expressed miRNAs were selected to verify by real-time PCR. The mature sequence of miRNAs and information of primers are listed in Supplementary file 3. U6 was used as the housekeeper genes. The miRcute miRNA RT-PCT kit (TIANGEN, China) was used to determine miRNA expression, which was carried out using an iCycler IQ5 Multicolor real-time PCR detection system (Bio-Rad, Hercules, CA, USA). The procedures included 30 s of pre-denaturation at 95°C, followed by 40 cycles of denaturation at 95°C for 5 s and 60°C for 30 s. Real-time PCR analysis of each sample was repeated in triplicate. The data were calculated by the normalized relative quantification method followed by 2âÎÎCt [21]. All data were calculated according to the normalization of log2 Fold-change (log2[JCH/NH-P]) to compare the predicted value and experimental value and verify the accuracy of the sequence.
All data were processed using the MS Excel program. A t-test was employed to analyze the significance of the differences between the groups at a level of p<0.05 using SPSS 13.0 software [22].
Two small RNAs (sRNA) libraries (JCH and NH-P) were constructed and sequenced. A total of 11.5 and 12.0 million raw reads were obtained in the two libraries, respectively (Table 1). The data of Q20 and Q30 were both more than 90% of the raw reads. The percentage of total G and C in all bases of JCH was 51.67% and it was50.63% in NH-P.
Referencing the duck genome (GCF_000355885.1), the total sRNA reads were 6215728 and 6463060 in the JCH and NH-P libraries (Supplementary file 2). In the two libraries, there were 4570475 (73.53%) and 4887684 (75.62%) reads mapped to the reference genome, respectively for JCH and NH-P groups. As seen in Figure 1, the length of sRNA was mainly distributed from 21nt to 27nt in JCH and NH-P libraries, and the most abundant length size were 21nt, 22nt, and 23nt. The number of 22nt miRNAs accounted for 18.88% in JCH group, and accounted for 21.03% in NH-P group.
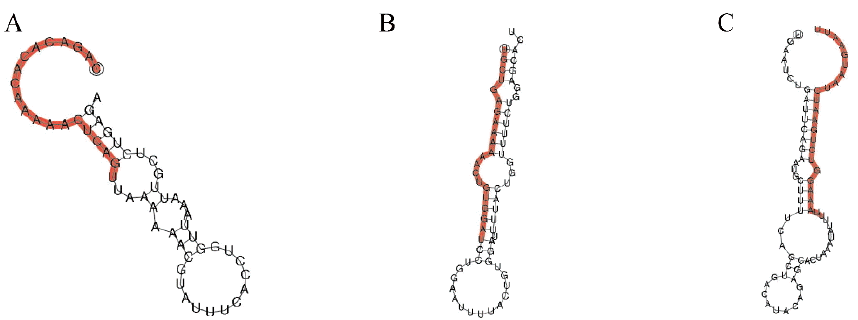
By comparing to the known miRNAs of miREVO and miRDeep2 software, conserved miRNAs were identified in JCH and NH-P group, which was 218 and 228, respectively (Supplementary file 3). According to the symbolic structure of the miRNA hairpin, we used miRDeep2 and miREVO software to discover the novel miRNAs. The results showed that the numbers of novel miRNAs were 26 and 27 in JCF and NH-P. Comparing these novel miRNAs, there are 25 novel miRNAs which are shared in two libraries (Supplementary file 4). Therefore, these novel miRNAs may be specific for duck species or had not identified yet (Supplementary file 5). The typical secondary structures of 28 novel miRNAs precursors were predicted by miRDeep2 software and shown in Figure 2.
The differential expression of miRNAs were enriched by using two standards that were p value<0.05 and fold-change log2 (JCF/NH-P)>0.05. The volcano plots (Figure 3A) of JCF vs NH-P concluded the distribution in different expressed miRNAs at the whole. There were 66 significant differentially expressed miRNAs enriched between the two libraries, including 23 significant up-regulated (p<0.05) and 43 significant down-regulated (p<0.05) (Figure 3B).
The top 21 miRNAs with the different expression levels in JCF and NH-P (Table 2) are listed. It was found that gga-miR-6651-5P, which is involved in regulating the process of resistance to the MADV [11], was the largest up-regulated in JCH compared to NH-P and its expression was only detected in JCH. We also noticed that gga-miR-122-5p was the largest down-regulated in JCH and it played an important role in the growth and development of liver, lipid metabolism [23]. The fold changes of 21 listed differential expressed miRNAs were all above 0.5 times.
Distribution of the candidate differential expressed miRNAs target-genes in the GO will clarify the main possible differences of BF impacted by genetic backgrounds. The results of GO analysis of miRNAs target genes is shown in Figure 4A. The significantly enriched GO terms were mainly distributed in biological process and molecular function. In addition, directed acyclic graph (DAG) was used to further display the detailed relationship of the enriched GO terms, and the deeper the color indicates the higher the level of enrichment in DAG dates. The target genes enriched in biological processes mainly concluded cellular response to stimulus, cellular process and cellular communication (Figure 4C). In cellular components (Figure 4D), the main differences were concentrated in the intrinsic to membrane and integral to membrane. In molecular function (Figure 4B), the main difference was concentrated in ion binding and receptor activity.
The scatter diagram of KEGG pathways enriched by differential expressed miRNAs target genes showed the top 20 enriched pathways, and the results revealed that the metabolic pathways (304 targets genes of enriched miRNAs) had the maximum enrich factor value and the lowest Q value (Figure 5A). Some pathways that ranked in the top 20 associated to immunity, such as pathways in cancer and bacterial invasion of epithelial cells. When considering the quantity of target genes enriched in each pathway, we found that PI3K-Akt, Epstein-Barr virus infection, human T-cell leukemia virus type I (HTLV-I) infection and mitogen-activated protein kinase (MAPK) signaling pathways were all ranked in the top 20 and all of them were related to immunity (Figure 5B).
Using the miRNAs target genes related to immunity to analyze KEGG enrichment, the top 18 enriched pathways are listed in Figure 5C. Some of the differentially expressed miRNAs targets genes were enriched in HTLV-I infection, Influenza A, bacterial invasion of epithelial cells, B cell receptor signaling pathway, T cell receptor signaling pathway, viral carcinogenesis, Apoptosis and MAPK signaling pathway. These enriched pathways indicated that there exists differences in immune resistance to disease both in innate and acquired immunity between two genetically disparate duck breeds.
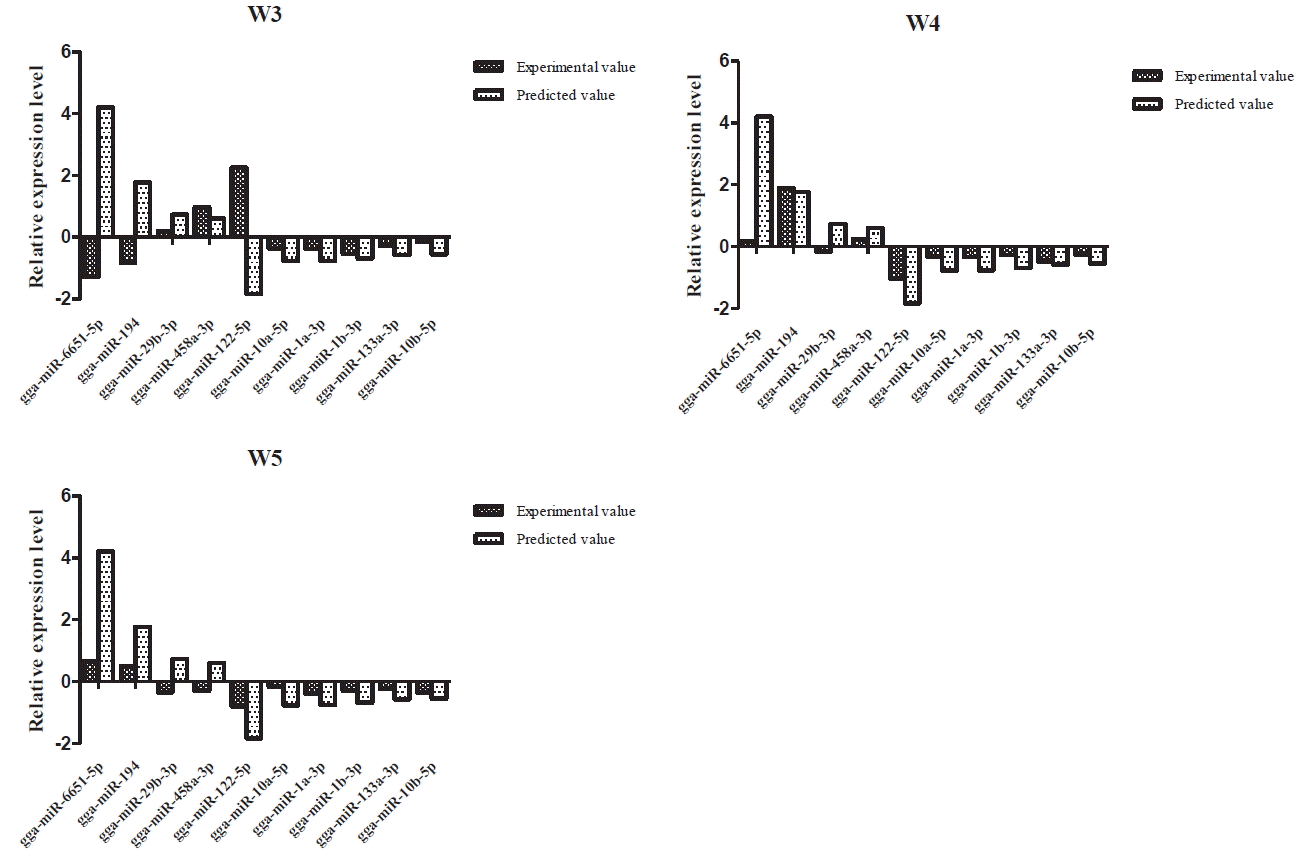
We chose 10 miRNAs which from the top 21 miRNAs with the highest expression levels in JCF and NH-P to verify the expression profiles. The selected miRNAs include the up-regulated miRNAs, such as gga-miR-6651-5p, gga-miR-194, gga-miR-29b-3p, gga-miR-458a-3p and the down-regulated miRNAs which were gga-miR-122-5p, gga-miR-10a-5p, gga-miR-1a-3p, gga-miR-1b-3p, gga-miR-133a-3p, gga-miR-10b-5p. We verified that the expression of 10 miRNAs in the BF of JCH and NH-P ducks at 3, 4, 5 weeks by real time PCR. Through calculating the results that the fold change log2(JCF/NH-P), the results showed that there were 9 miRNAs (90%) that had the same variation trend compared with the results as predicted at 4 week (Figure 6). By comparing the predicted dates of 4 weeks to the experimental dates of 3 and 5 weeks, there are 7 and 8 miRNAs with same variation trend, respectively.
MiRNAs, as small non-coding RNAs, take part in regulation of various biological processes at the post-transcriptional level [24]. By high-throughput sequencing, recent studies have illustrated that miRNAs play a role in different organs of ducks, including embryonic breast muscle, ovary, feather follicle and skin [25â27]. Previous studies have suggested that miRNAs are key regulators of the immune system [8]. Some researchers have discussed the functions of miRNAs after ducks were infected with disease [28, 29], however, fewer reports have displayed the impacts of genetic backgrounds on miRNAs function of ducksâ immune organs. In the present study, it was shown that some of the predicted miRNAs were closely related to proliferation, differentiation and death of immune organs or cells, e.g. miRNA-29b can work as a regulator of Th1 differentiation by regulating T-bet and IFN-Îł [30], the miR-17~92 cluster can regulate monocyte differentiation by targeting the transcription factor acute myeloid leukemia 1 (AML1) [31]. And, some of the predicted miRNAs were related to immune response, e.g. miRNA-146a has an important function in the innate immune system by controlling Toll-like receptors and cytokine signaling through a negative feedback loop involving the down-regulation of interleukin-1 receptor associated kinase (IRAK1) and tumor necrosis factor receptor associated factor 6 (TRAF6) (signaling components of the toll-like receptors [TLRs]) [32]. The let-7 is downregulated in many human tumors and overexpression of let-7 inhibits cancer growth [7]. All of these predicted miRNAs support the concept that the brusa of Fabricius is an essential immune organ and the BF of ducks play an important role in immunity and resistance to disease.
There were total 28 novel miRNAs identified in the present study. It was found that these novel miRNAs were involved in immune and resistance to disease through predicting their target genes. The tendency of novel miRNAs functions was same as with the conserved miRNAs. In the predicted target genes, RIG-1 (matched 3 novel miRNAs) works as a cytoplasmic protein that mainly recognizes viral dsRNA [33,34], NF-kB (matched 4 novel miRNAs) is a crucial transcription factor for immunosuppression and anti-inflammatory molecules [35], and IRF-7 (matched 3 novel miRNAs) can regulate type-I infection-dependent immune responses as a transcriptional regulation factor. In addition [36], all these genes (RIG-1, NF-kB, and IRF-7) are key components of RIG-1-like receptor signaling pathway. The duck is considered to have a higher anti-adversity ability than other birds, and as such are less susceptible to avian influenza [29,37]. These novel miRNAs, which might be specific to the duck and have a predicted correlation with avian influenza, indicte that these novel miRNAs may be associated with duck high anti-adversity ability. Further researches would be needed to investigate the detailed functions of these novel miRNAs in the resistance to disease.
Ducks from JCH and NH-P were disparate in genetic backgrounds, JCH is a local breed with strong physical features and only produced within in Sichuan Province in China. NH-P is a new cultivated breed, with high-strength selection process in order to acquire maximized economic traits. In our laboratory previous study, it had been demonstrated that the differences exist in immune organ index, organization structure and expression of immune related gene and protein in the blood between JCH and NH-P ducks. The data showed that there were 66 differential expressed miRNAs in the brusa of Fabricius between the two varieties of ducks. The accurate of sequencing data were confirmed by real-time PCR, and proved with a high repeatability (9 of 10). These differentially expressed miRNAs in BF may contribute to the divergent immunity between duck breeds. As in detail, the top 8 miRNAs with higher expression in JCH (Table 2) were almost all involved in immunity and disease by analyzing the function of differential expressed miRNAs. For example, the gga-miRNA-6651-5p was only detected expression in JCH, and it had been demonstrated that it was dramatically increased in the MADV-GA-infected nontumorous samples but not in the tumorous and mock-infected groups, suggesting that it may be involved in regulating the process of resistance to the MADV [11]. Moreover, study had showed that miR-215 functions as a tumor suppressor that regulates cell cycle progression [38]. The miR-147 was induced upon Toll-like receptor stimulation and functioned as a negative regulator of TLR-associated signaling events in murine macrophages and it was up-regulated when chickens were infected with Marekâs disease virus [39]. In addition, miRNA 194 might play an important role in various cancer invasion and progression [40]. Therefore, the most differentially expressed miRNAs with higher expression in JCH were almost all related to immunity and disease, which supports JCH as having a better ability of disease resistance than NH-P. Moreover, these enriched differential expressed miRNAs maybe provide new insight into molecular marker assisted selection of resistance to disease in poultry.
The GO terms, such as cellular response to stimulus, cellular communication, receptor activity, enriched the most target genes. The bursa of Fabricius, with many B cells, is one of essential immune organs [4]. The mature B cells can synthesis immunoglobulin and it had affirmed that there are IgM, IgG, and IgA in avian, which had a role in immune response to fight infection, cytotoxicity, cytolysis and neutralize virus infection [28]. In addition, the BF also provides microenvironment for the occurrence of humoral immunity [41,42]. Therefore, there was frequency recognition of external stimuli and signal transfer, and the BF miRNAs of two duck breeds were different in these aspects. KEGG pathway showed that there were many enriched pathways involved in immunity and disease, such as Pathway in cancers, Bacterial invasion of epithelial cells, HTLV-1 infection [43], Influenza A, T cell receptor interaction, and Ras signaling pathway [44]. Taken together, these results agreed widely with the results of differential expressed miRNAs function, suggesting that the BF functions of two duck breeds were differenct in immunity and miRNAs may play a crucial role in the process of immune regulation.
The KEGG analysis showed that the metabolic pathways had the maximum enriched factor value and the lowest Q value. As a part of metabolic, cell cycles were also enriched and ranked in top 20 enriched pathways, such as, MAPK signaling pathway [45], Focal adhesion [46], extracellular matrix-receptor interaction [47], and N-Glycan biosynthesis [48]. In addition, PI3K-Akt pathway had secondary enriched genes and it had demonstrated that it might be mediate the direct anti-apoptotic activity of chicken bursal B cell cultures [49]. Proteins, as main ingredients of cells, are synthesised in the process of cell proliferation and many pathways related to protein synthesis and modification were enriched, such as Protein processing in endoplasmic reticulum [50], Ubiquitin mediated proteolysis [51]. On the contrary, the results showed that differences existed in apoptosis and lysomsome. These results indicated that there were differences in cell proliferation and apoptosis. Previous research had demonstrated that only approximately 5% of daily B-cell production survives to form a mature postbursal B-cell population, which supports the observation that there were frequent B cell apoptosis and proliferation in the brusa [4]. Therefore, we inferred that cell cycle, apoptosis and proliferation were different between two duck breeds, which can influence the quantities of mature functional B cells. In addition, cycle of cellsâ renewal can influence immunity, the same as the speed of antibodiesâ production [52], efficiency of toxic production excretion [53], which both were included in metabolism. Therefore, the KEGG pathway results showed that the cell metabolism of the BF had obvious differences and this study inferred that one of the reasons leading to the immune differences in the BF was the metabolic difference.
In this study, the miRNA profiles revealed different miRNA regulation mechanism might exist in the brusa of Jianchang and Nonghua-P strain ducks. There were a total of 66 differential expressed miRNAs and 28 novel miRNAs. The predicted target genes of these novel miRNAs were associated with duck high anti-adversity ability. The targets of differential expression miRNAs were mainly involved in the process of immunity. In addition, metabolic pathways had the maximum enriched target genes and some enriched pathways were related to cell cycle, protein synthesis, cell proliferation and apoptosis. These data listed the main difference in the BF at miRNAs level between two genetically disparate duck breeds and lays a foundation for marker assisted selection of poultry in the future.
ACKNOWLEDGMENTS
This work was supported by the application foundation project of Sichuan Science and Technology Bureau (2015JY0110), the National Natural Science Foundation of China (No.31301964), Chinese Agriculture Research Service (No.CARS-43-6), the Major Project of Sichuan Education Department (13ZA0252).
Figure 1
The length distribution of small RNA in JCH and NH-P. JCH, Jianchang duck; NH-P, Nonghua P-strain meat duck.

Figure 3
Differential expression of miRNAs between JCF and NH-PF. The differentially expressed miRNAs were filtered by |log2(FoldChange)|>0.05 and q-value<0.05. (A) The volcano plots of differential expressed miNRA. Each point in the figure represents a miRNA. Square points represent significant up-expressed miRNAs, triangular points represent significant down-expressed miRNAs, dot represent miRNAs which were no significant difference. (B) The statistic of differential expressed miRNA in JCH and NH-P. JCH, Jianchang duck; NH-P, Nonghua P-strain meat duck.
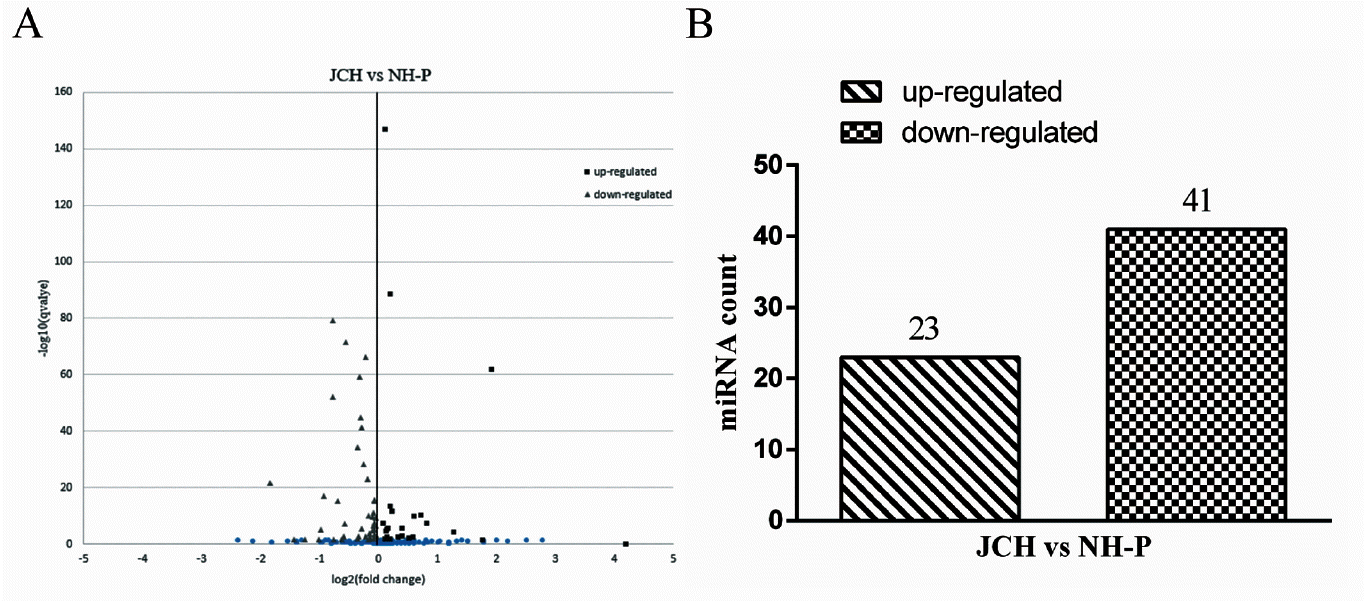
Figure 4
GO enrichment analysis of target genes. (A) The bar graph of the most enriched GO terms of target genes, y-axis means the percentages of differentially genes in term, x-axis means biological process (bp), cellular component (cc) and molecular function (mf); (B), (C), (D). Directed acyclic graph (DAG) in molecular function, biological process and cellular component, respectively. The deeper of the color means the higher level of enrichment in DAG dates.
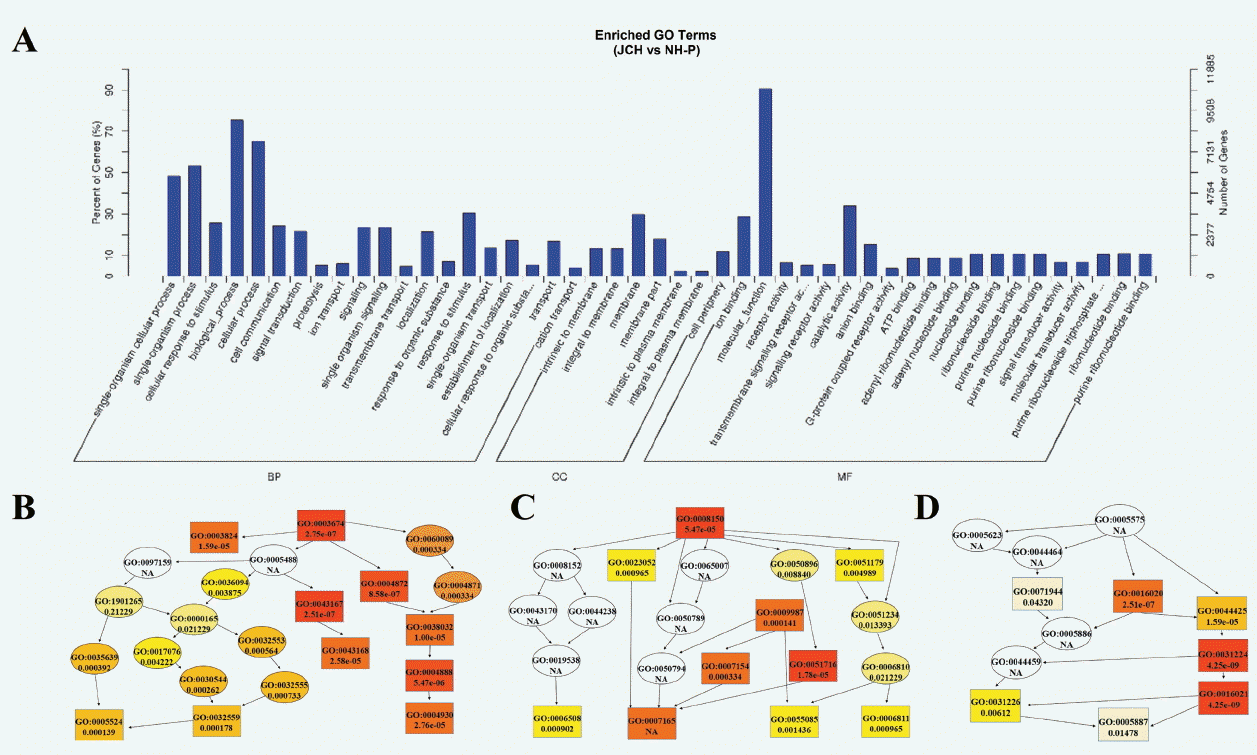
Figure 5
KEGG analysis of target genes. (A) KEGG scatter plot analysis of target genes for differentially expressed miRNAs. The samller of the qvalue indicated the more significant of the enrichment; the greater of enrichment factor indicated the more number of target genes. (B) The top 20 pathways of the most miRNAs target genes enriched. (C) The bar graph of the enriched pathways of target genes related to immunity.
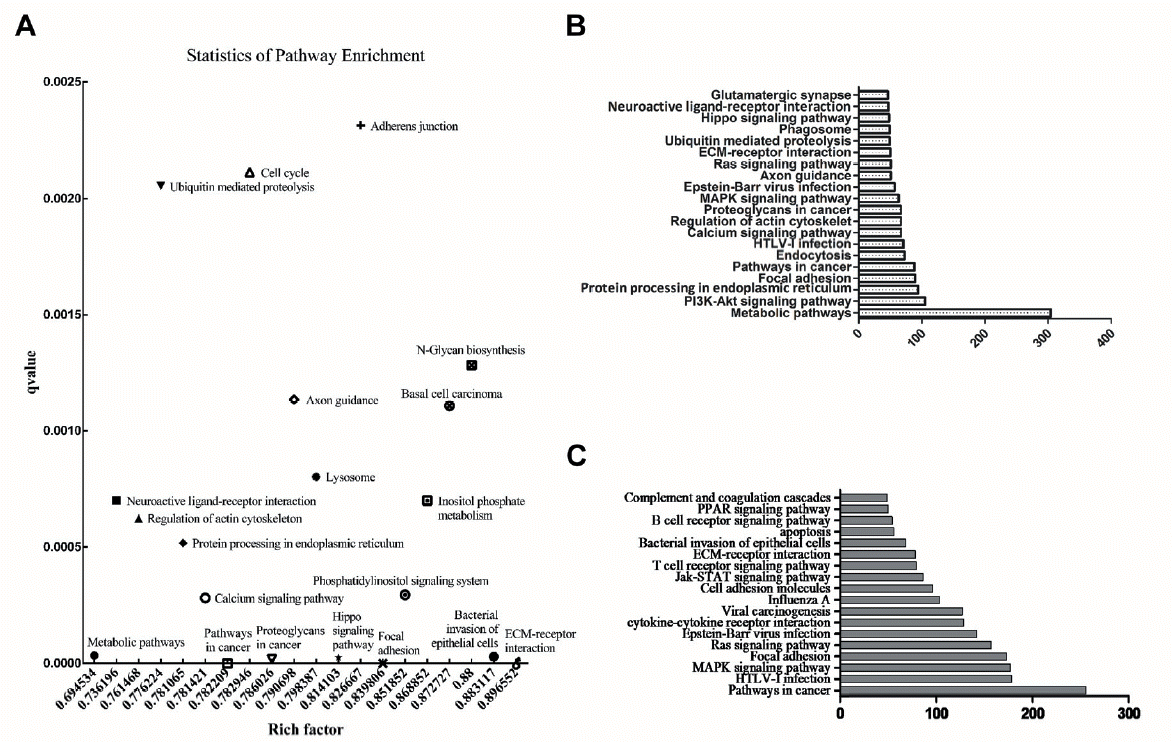
Table 1
Quality analysis of the two small RNA libraries
Table 2
The top 21 miRNAs with the different expression levels in JCH and NH-P
REFERENCES
1. Korte J, FrĂśhlich T, Kohn M, et al. 2D DIGE analysis of the bursa of Fabricius reveals characteristic proteome profiles for different stages of chicken B-cell development. Proteomics 2013; 13:119â33.


2. Glick B, Chang TS, Jaap RG. The bursa of fabricius and antibody production. Poult Sci 1956; 35:224â5.

3. Yin T-B, Liu X-Y. Poultry immunology. Beijing, China: China Agriculture Science and Technique Press; 1999.
4. Mustonen L, Alinikula J, Lassila O, Nera KP. Bursa of Fabricius Encyclopedia of Life Science. Hoboken, NJ: Wiley; 2010.
5. Ratcliffe MJ. Antibodies, immunoglobulin genes and the bursa of Fabricius in chicken B cell development. Dev Com Immunol 2006; 30:101â18.

6. Bartel DP, Chen C-Z. Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nat Rev Genet 2004; 5:396â400.


7. Schickel R, Boyerinas B, Park S, Peter M. MicroRNAs: key players in the immune system, differentiation, tumorigenesis and cell death. Oncogene 2008; 27:5959â74.


8. Hicks JA, Tembhurne PA, Liu H-C. Identification of microRNA in the developing chick immune organs. Immunogenetics 2009; 61:231â40.


9. Trakooljul N, Hicks J, Liu HC. Identification of target genes and pathways associated with chicken microRNA miR-143. Anim Genet 2010; 41:357â64.


10. Chen C-Z, Li L, Lodish HF, Bartel DP. MicroRNAs modulate hematopoietic lineage differentiation. Science 2004; 303:83â6.


11. Li Z-J, Zhang Y-P, Li Y, et al. Distinct expression pattern of miRNAs in Marekâs disease virus infected-chicken splenic tumors and non-tumorous spleen tissues. Res Vet Sci 2014; 97:156â61.


12. Wang Y-S, Ouyang W, Pan Q-X, et al. Overexpression of microRNA gga-miR-21 in chicken fibroblasts suppresses replication of infectious bursal disease virus through inhibiting VP1 translation. Antiviral Res 2013; 100:196â201.


13. Rodriguez A, Vigorito E, Clare S, et al. Requirement of bic/microRNA- 155 for normal immune function. Science 2007; 316:608â11.



14. Dahlberg JE, Lund E. Micromanagement during the innate immune response. Sci Signal 2007; 2007:pe25-pe

15. Dinh H, Hong YH, Lillehoj HS. Modulation of microRNAs in two genetically disparate chicken lines showing different necrotic enteritis disease susceptibility. Vet Immunol Immunopathol 2014; 159:74â82.


16. Tian F, Luo J, Zhang H, Chang S, Song J. MiRNA expression signatures induced by Marekâs disease virus infection in chickens. Genomics 2012; 99:152â9.


17. Wen M, Shen Y, Shi S, Tang T. miREvo: an integrative microRNA evolutionary analysis platform for next-generation sequencing experiments. BMC Bioinformatics 2012; 13:140



18. Friedländer MR, Mackowiak SD, Li N, Chen W, Rajewsky N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res 2012; 40:37â52.




19. Zhou L, Chen J, Li Z, et al. Integrated profiling of microRNAs and mRNAs: microRNAs located on Xq27. 3 associate with clear cell renal cell carcinoma. PLoS ONE 2010; 5:e15224



20. Wang L, Feng Z, Wang X, Wang X, Zhang X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 2010; 26:136â8.


21. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2âÎÎCT method. Methods 2001; 25:402â8.


22. Green SB, Salkind NJ. Using SPSS for Windows and Macintosh: Analyzing and understanding data. Upper Saddle River, NJ: Prentice Hall Press; 2010.
23. Li Y, Wang X, Yu J, et al. MiR-122 targets the vanin 1 gene to regulate its expression in chickens. Poult Sci 2016; 95:1145â50.


25. Yu D-B, Jiang B-C, Jing G, et al. Identification of novel and differentially expressed microRNAs in the ovaries of laying and non-laying ducks. J Integr Agric 2013; 12:136â46.

26. Zhang L, Nie Q, Su Y, et al. MicroRNA profile analysis on duck feather follicle and skin with high-throughput sequencing technology. Gene 2013; 519:77â81.


27. Gu L, Xu T, Huang W, et al. Identification and profiling of microRNAs in the embryonic breast muscle of pekin duck. PloS one 2014; 9:e86150



28. Cong L-X, Su J-Z, Xing S-Y, Zhao Z-H, Zhang J-Y. Detection of the expression and comparative study of miRNA-17a~* in H5N1 infected and uninfected SPF ducks. Heilongjiang Anim Sci Vet Med 2012; 21:005
29. Li Z, Zhang J, Su J, et al. MicroRNAs in the immune organs of chickens and ducks indicate divergence of immunity against H5N1 avian influenza. FEBS Lett 2015; 589:419â25.


30. Smith KM, Guerau-de-Arellano M, Costinean S, et al. miR-29ab1 deficiency identifies a negative feedback loop controlling Th1 bias that is dysregulated in multiple sclerosis. J Immunol 2012; 189:1567â76.



31. Baltimore D, Boldin MP, OâConnell RM, Rao DS, Taganov KD. MicroRNAs: new regulators of immune cell development and function. Nat Immunol 2008; 9:839â45.


32. Taganov KD, Boldin MP, Chang K-J, Baltimore D. NF-ÎşB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Nat Acad Sci 2006; 103:12481â6.



33. Lynn DJ, Winsor GL, Chan C, et al. InnateDB: facilitating systems-level analyses of the mammalian innate immune response. Mol Syst Biol 2008; 4



34. Yoneyama M, Kikuchi M, Natsukawa T, et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat Immunol 2004; 5:730â7.


35. Baeuerle PA, Baichwal VR. NF-kB as a frequent target for immunosuppressive and anti-inflammatory molecules. Adv Immunol 1997; 65:111â38.


36. Honda K, Yanai H, Negishi H, et al. IRF-7 is the master regulator of type-I interferon-dependent immune responses. Nature 2005; 434:772â7.


37. Huang Y, Li Y, Burt DW, et al. The duck genome and transcriptome provide insight into an avian influenza virus reservoir species. Nat Genet 2013; 45:776â83.



38. Georges SA, Biery MC, Kim S-Y, et al. Coordinated regulation of cell cycle transcripts by p53-Inducible microRNAs, miR-192 and miR-215. Cancer Res 2008; 68:10105â12.


39. Liu G, Friggeri A, Yang Y, et al. miR-147, a microRNA that is induced upon Toll-like receptor stimulation, regulates murine macrophage inflammatory responses. Proc Nat Acad Sci 2009; 106:15819â24.



40. Dong P, Kaneuchi M, Watari H, et al. MicroRNA-194 inhibits epithelial to mesenchymal transition of endometrial cancer cells by targeting oncogene BMI-1. Mol Cancer 2011; 10:1



41. Kumar R, Singh GK, Chauhan RS. Development of bursa of fabricius in relations to humoral immunity in chicken embryo. Indian J Anim Sci 2004; 74:838â40.
42. Singh S, Singh I, Singh G, Gangwar C, Kumar P. Postnatal development of bursa of Fabricius in relation to humoral Immunity in Keets. J Immunol Immunopathol 2011; 13:42â5.
44. Vojtek AB, Der CJ. Increasing complexity of the Ras signaling pathway. J Biol Chem 1998; 273:19925â8.


45. Zhang W, Liu HT. MAPK signal pathways in the regulation of cell proliferation in mammalian cells. Cell Res 2002; 12:9â18.


46. Zhao J-H, Reiske H, Guan J-L. Regulation of the cell cycle by focal adhesion kinase. J Cell Biol 1998; 143:1997â2008.



47. Lukashev ME, Werb Z. ECM signalling: orchestrating cell behaviour and misbehaviour. Trends Cell Biol 1998; 8:437â41.


48. Lau KS, Partridge EA, Grigorian A, et al. Complex N-glycan number and degree of branching cooperate to regulate cell proliferation and differentiation. Cell 2007; 129:123â34.


49. Luna-Acosta JL, Alba-Betancourt C, MartĂnez-Moreno CG, et al. Direct antiapoptotic effects of growth hormone are mediated by PI3K/Akt pathway in the chicken bursa of Fabricius. Gen Comp Endocrinol 2015; 224:148â59.


50. Lawson WE, Crossno PF, Polosukhin VV, et al. Endoplasmic reticulum stress in alveolar epithelial cells is prominent in IPF: association with altered surfactant protein processing and herpesvirus infection. Am J Physiol Lung Cell Mol Physiol 2008; 294:L1119âL26.


51. Ciechanover A, Orian A, Schwartz AL. Ubiquitin-mediated proteolysis: biological regulation via destruction. Bioessays 2000; 22:442â51.







 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Supplement
Supplement Print
Print





