 |
 |
| Anim Biosci > Volume 29(11); 2016 > Article |
|
Abstract
This study was conducted to estimate the genetic parameters of Ī²-hydroxybutyrate (BHBA) and acetone concentration in milk by Fourier transform infrared spectroscopy along with test-day milk production traits including fat %, protein % and milk yield based on monthly samples of milk obtained as part of a routine milk recording program in Korea. Additionally, the feasibility of using such data in the official dairy cattle breeding system for selection of cows with low susceptibility of ketosis was evaluated. A total of 57,190 monthly test-day records for parities 1, 2, and 3 of 7,895 cows with pedigree information were collected from April 2012 to August 2014 from herds enrolled in the Korea Animal Improvement Association. Multi-trait random regression models were separately applied to estimate genetic parameters of test-day records for each parity. The model included fixed herd test-day effects, calving age and season effects, and random regressions for additive genetic and permanent environmental effects. Abundance of variation of acetone may provide a more sensitive indication of ketosis than many zero observations in concentration of milk BHBA. Heritabilities of milk BHBA levels ranged from 0.04 to 0.17 with a mean of 0.09 for the interval between 4 and 305 days in milk during three lactations. The average heritabilities for milk acetone concentration were 0.29, 0.29, and 0.22 for parities 1, 2, and 3, respectively. There was no clear genetic association of the concentration of two ketone bodies with three test-day milk production traits, even if some correlations among breeding values of the test-day records in this study were observed. These results suggest that genetic selection for low susceptibility of ketosis in early lactation is possible. Further, it is desirable for the breeding scheme of dairy cattle to include the records of milk acetone rather than the records of milk BHBA.
Adipose tissues are mobilized for energy, and nonesterified fatty acids are converted to ketone bodies (acetone, acetoacetate, Ī²-hydroxybutyrate [BHBA]) when oxaloacetate becomes the limiting factor in the Krebās cycle (Wood et al., 2004). Acetoacetate can be detrimental to cows, leading to ketosis, which is a typical metabolic disorder that can appear with negative energy. High milking cows often do not ingest the necessary nutrients for milk production; thus, the highest incidence of ketosis is observed in dairy cows during the early lactation period (Baird, 1982; Sakha et al., 2006) and higher lactations (Vosman et al., 2015). Subclinical ketosis is characterized by increased concentrations of the ketone bodies in blood, milk, and urine without any clinical signs, while symptoms of clinical ketosis include anorexia, hypoglycemia and hyperketonemia (Baird, 1982; Duffield et al., 1997). However, both clinical and subclinical ketosis negatively affect milk production and reproduction (Enjalbert et al., 2001; Oetzel, 2012), and therefore lead to economic losses due to an increase in the forced culling of animals or higher veterinary costs.
Acetone and BHBA have been suggested as useful indicators of subclinical and clinical ketosis (Enjalbert et al., 2001; Nielsen et al., 2005). Concentrations of ketone bodies in the milk are closely correlated (r>0.8) with blood levels (Andersson, 1988). Further, milk Ī²-hydroxybutyric acid and milk acetone with information regarding the body condition score, milk fat and milk protein for energy balance can be used to establish ketosis in dairy cows (Koeck et al., 2014). Since April of 2012, Fourier transform infrared (FTIR) measurements for milk acetone and milk BHBA have been tentatively collected along with routinely evaluated milk measurements during dairy herd milk recording in Korea. Since ketosis is one of the most frequent diseases in dairy cattle, selective breeding for low incidence of hyperketonemia could be important to the dairy industry. Van der Drift et al. (2012) reported that there were moderate genetic relationships among concentrations of plasma BHBA, milk BHBA and milk acetone that may allow the use of milk BHBA and milk acetone in dairy cattle breeding. However, questions remain about whether the sensitivity of the records is sufficient to provide genetic information that can be used to select for low susceptibility to ketosis. Therefore, this study was conducted to investigate the heritabilities of milk BHBA and milk acetone and their genetic correlations, as well as the relationships with routinely evaluated test-day (TD) traits during first, second and third lactations.
The FTIR measurements for milk acetone and milk BHBA levels along with routine monthly TD records were collected from April 2012 to August 2014 by the Korea Animal Improvement Association (KAIA). TD milk sampling and analysis were performed according to the Korean milk-recording procedures (Cho et al., 2013). Milk samples were taken in the evening and morning. TD milk samples were analyzed by FTIR spectroscopy using a CombiFos FT+ system (Foss Analytical A/S, HillerĆød, Denmark) with previously developed calibration equations for milk BHBA and milk acetone from the manufacturer. The TD milk records included milk yield and fat and protein percentages. The morning records of registered cows with pedigree information provided by KAIA and of test days from cows that were 4 to 305 days in milk (DIM) at sampling were included in the analysis. The ages at calving ranged from 18 to 48 months, 32 to 57 months and 46 to 71 months for first, second and third lactations, respectively. Outliers over Ā±3 standard deviation (SD) for milk production measurements were considered to be incorrect recordings and excluded from the analysis. The parent information was available for all records in the data set. The basic and descriptive statistics are provided in Table 1. The TD records (25,185, 18,836, and 13,169 records, respectively) and the numbers of cows (7,896, 6,095, and 3,902, respectively) and sires (308, 295, and 278, respectively) for the first, second and third lactations were used, after editing. In addition, even though the records are monthly collected for every cow in the milk recording system, the average numbers of TD records per lactation were 3.2, 3.3, and 3.4 for first, second and third lactations, respectively. Cows with third parity records had to have their records in first and second parity.
The prevalence of hyperketonemia varies across parity, stage of lactation and season in cows (Duffield et al., 1997; van der Drift et al., 2012). Preliminary analyses (general linear model procedure; SAS Institute Inc. Cary, NC, USA) of concentrations of the ketone bodies indicated that age classes, seasons at calving, herds and test days should be corrected in the models. The herd-test day subclasses take fixed regressions on DIM (Jamrozik et al., 1997) into account in the random regression model (RRM). The RRMs provide genetic and permanent environmental (co)variance throughout lactation. There are different functions that can be used to fit the yield trajectory over a period. The functions most often used for dairy cattle are Legendre orthogonal polynomials. Legendre polynomials are flexible, enabling the fitting of curves independent from the trait of interest. Jamrozik and Schaeffer (2002) found that the TD milk yield with Legendre polynomials outperformed the TD milk yield with a lactation curve function. Kirkpatrick and Heckman (1989) suggested the use of covariance functions defined by orthogonal Legendre polynomials to describe the (co)variance structure of longitudinal observations. For RRMs based on Legendre polynomials, Pool et al. (2000) suggested that at least a three coefficient polynomial was needed to model the (co)variance structure of the random components of the longitudinal observations. The order of random regressions often does not exceed three for computational ease, although regressions with higher orders model curves more accurately (Van Der Werf et al., 1998). Even though Van der Werf et al. (1998) and Pool et al. (2000) suggested a lower degree for animal genetic components than for permanent environmental components, an order 3 of fit was equally applied to provide equal opportunity for variation of both components over a period.
The genetic ability to produce milk varies among parities. Animals with an identical production potential for the first lactation can have a genetically different production level than matured animals. Moreover, the TD records obtained in first, second, and third lactation are treated as independent traits in many countries. The TD records of milk BHBA, acetone and other traits were separately analyzed for each parity. Genetic parameters, including genetic (co)variance components, were estimated by the restricted maximum likelihood procedure based on a random regression TD animal model using the Wombat program (Meyer, 2015). The linear model for analysis was as follows:
where, yijkl:t is the TD record at t DIM for one of the analyzed traits (milk BHBA, milk acetone, milk yield, fat, or protein %) of cow l in a lactation (parity 1 to 3) kth season within jth age of calving at ith herd TD, and t is any DIM between 4 and 305; HTDi = ith herd TD effect, which takes the fixed effect of the lactation curve for each trait at t DIM into account; AgeSeasonjk is the effect of season k of age j at calving [age was categorized into 4 groups and calving seasons were defined as summer (May to October) and winter (November to April)]; aln:t is the additive random regression coefficient of the lth animal on t DIM; pln:t is the permanent environmental random regression coefficient of the lth animal on t DIM; zln:t is the covariate associated with standardized DIM; and eijkl:t is a random residual effect. As zln:t is the Legendre polynomial that was recommended by Kirkpatrick et al. (1990), z1 ā z3 were:
where, d is a standardized DIM with a range of ā1 to +1 that is derived as:
where, tmin is the smallest DIM and tmax is the largest DIM represented in the data.
In matrix notation, the statistical model with multi-traits was:
where, y is the matrix of observations of the traits; X and Z are the known incidence matrices for fixed and random effects; b is the vector of fixed effects; a is the vector of additive genetic regression coefficients for each animal; p is the vector of permanent environmental regression coefficients for each animal and e is the vector of residual effect. The (co)variance matrices of random effect factors in a, p, and e were assumed to be:
where, G is the (co)variance matrix of the additive genetic random regression coefficients of order 3; A is the additive genetic relationship matrix between animals; P is the (co)variance matrix of the permanent environmental random regression coefficients of order 3; I is the identity matrix; and R is the residual (co)variance.
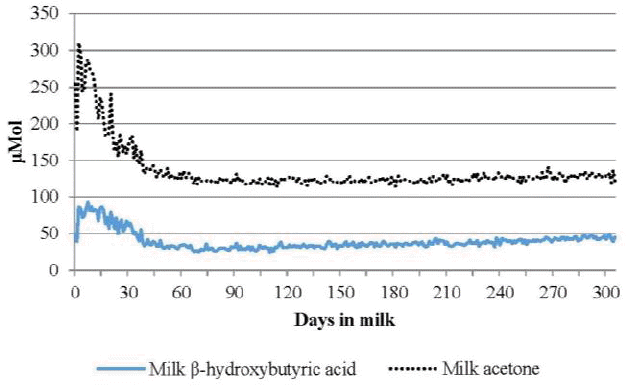
Herds are usually monitored for hyperketonemia by regular assessment of the percentage of fresh cows having ketone body concentrations above a defined threshold value (prevalence of hyperketonemia). Prevalence of hyperketonemia does not reflect biological variations in ketone body concentrations or the severity of hyperketonemia in cows. Consequently, attention was paid to concentrations rather than presence or absence of hyperketonemia in cows to investigate genetic variations in ketosis susceptibility. The means of milk BHBA concentrations in this study were high between 1 and 30 DIM and low around 70 DIM, after which they gently increased with DIM (Figure 1), which is consistent with reports by Baird (1982) and Sakha et al. (2006). The means of milk BHBA increased along with SDs through parities (Table 1). The mean concentrations of milk BHBA between 4 and 305 DIM for parities 1, 2, and 3 were 38.0, 39.3, and 43.7 Ī¼mol/L, respectively, which were consistent with the low occurrence of ketosis in the first lactation reported by Baird (1982). Comparatively, Koeck et al. (2014) found averages of 106 to 64 Ī¼mol/L from 5 to 100 DIM.
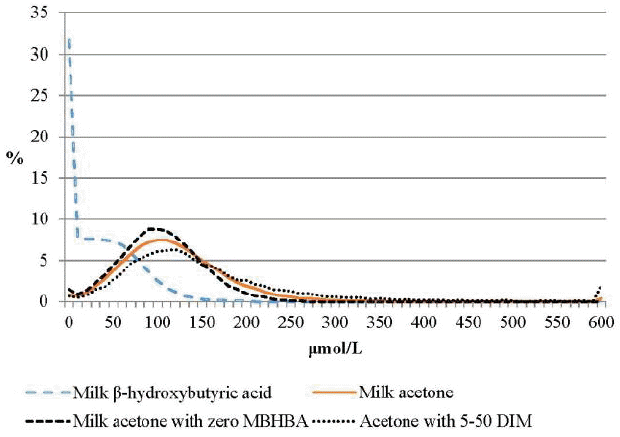
The distributions of milk BHBA and acetone concentrations are shown in Figure 2. Zero observations in milk BHBA concentration account for over 30% from the data set, consequently they were not normally distributed, which is considered a healthy signal for ketosis. Conversely, the distribution of milk acetone records showed a normal distribution, and further distribution in milk acetone of cows with no milk BHBA was similar (Figure 2). Log transformation for normal distribution in acetone concentration was not necessary as acetone data were transformed by Wood et al. (2004).
Subclinical and clinical ketosis were defined based on the BHBA concentration by Enjalbert et al. (2001) and McArt et al. (2012). The proportion of cows testing positive for hyperketonemia based on a milk BHBA ā„200 Ī¼mol/L as described by Koeck et al. (2014) was 6.6%. Additionally, the incidence rate for subclinical ketosis (10ā¤subclinical ketosis 200 Ī¼mol/L) was 67.4% based on an estimation from 31.7% milk BHBA ā¤10 Ī¼mol/L in early lactations (between 1 and 30 DIM) in this study. The prevalence of hyperketonemia was 0.9% between 1 and 305 DIM (Figure 2), which falls in the range of 0.5% to 5.1% incidence rates found for clinical ketosis in previous studies (van Dorp et al., 1999; Koeck et al., 2012). The incidence rate (67.4%) for subclinical ketosis was considerably higher than the range of 12% to 43% for subclinical ketosis observed by Duffield et al. (1997) and McArt et al. (2012). Those rates at 305 DIM were 0.7%, 0.9%, and 1.3% for hyperketonemia and 66.9%, 67.0%, and 68.6% for subclinical ketosis, respectively, for parity 1, 2, and 3. No clear reason was suggested for the high prevalence of subclinical ketosis in Korea. This could be explained by the short herd life (2.7 lactations on average) of dairy cows in Korea. Koeck et al. (2014) stated that lower milk BHBA was associated with a longer herd life, better conformation, and better feet and legs. There was a trend of increasing prevalence at later parities, which was consistent with the results reported by Vosman et al. (2015). The mean concentrations of milk acetone for parities 1, 2, and 3 were 131.9, 120.2, and 123.7 Ī¼mol/L, respectively, which were similar to the 140 Ī¼mol/L in lactations 1 and 2 reported by Wood et al. (2004). The mean acetone concentration was the highest between 1 and 30 DIM and lowest around 100 DIM, then gently increased with DIM (Figure 1).
The overall correlation of milk acetone with milk BHBA was 0.47 (p<0.001), which was lower than that of 0.68 found by Enjalbert et al. (2001). The regression equations for prediction (R2 = 0.2205) were as follows:
Andersson (1984) and Wood et al. (2004) suggested that concentration of acetone in milk could be used to estimate the incidence of clinical, or subclinical ketosis. The correlation between concentrations of acetone and acetoacetate was also quite high (r = 0.88; Tveit et al., 1992). Enjalbert et al. (2001) reported that milk acetone was very close to blood acetone, with a mean milk acetone/blood acetone ratio of 1.04. Andersson (1984) also observed a close relationship between these factors (r = 0.96). Moreover, Andersson (1988) reported that acetone had the highest relative concentration in milk (Table 1) and the least diurnal variation, making it the best of the three ketone bodies for ketosis testing. Tveit et al. (1992) found that milk acetone has a correlation of 0.88 with acetoacetate, which can lead to ketosis.
We focused on genetic variations in hyperketonemia and did not distinguish between cows with or without clinical signs of ketosis during data collection. However, milk BHBA showed a genetic correlation (0.48) with clinical ketosis (Koeck et al., 2014); therefore, genetic variations in ketone bodies could be used for selective breeding program for low incidence of ketosis. This study assessed genetic variation and heritability estimates for concentrations of milk BHBA and milk acetone in lactations 1, 2, and 3 to evaluate cows susceptible to hyperketonemia and the possibility of using TD milk ketone bodies for genetic improvement. Estimates of heritability and genetic and permanent environmental variance for milk BHBA and acetone are given in Table 2 and the traces of heritabilities and genetic variance components for milk BHBA and acetone from 4 to 305 DIM are shown in Figure 3 and 4.
Traces of heritabilities of milk BHBA for parities 1 and 2 were high at the beginning and end of the lactations, while the traces for parity 3 were high between 130 and 200 DIM (Figure 3). The highest trace of heritabilities with a mean of 0.11 and maximum of 0.17 was observed in parity 2 (Table 2 and Figure 3). The heritability for parity 3 was lowest at early lactation, being 0.04 at 4 DIM. In general, the peaks on the heritability traces were attained at mid lactation for parity 3 and at late lactation for parities 1 and 3. The genetic variance components were considerably higher until 30 DIM than during mid and late lactation for all parities. For parity 3, the additive genetic variances increased at mid lactation. Heritabilities were not as high as genetic variances during the same period, since phenotypic variances were also large during early lactation. Milk BHBA had similar and high traces of additive genetic variances at early lactation over parities, which might lead to high heritability with a proper statistical model.
In general, the heritability of acetone was higher than that of milk BHBA. The traces of heritabilities of milk acetone for all parities were low at the beginning of lactations, but increased steadily from mid-lactation. On the other hand, the trace for parity 3 was lower than those of other parities, except between 120 and 180 DIM (Figure 4). For parities 1 and 2, the traces of heritabilities were similar, with mean values of 0.29 and similar ranges of heritabilies (Table 2 and Figure 4). Parity 3 had the lowest heritability trace, which was 0.03 at 4 DIM. The heritability traces for parities had similar shapes as those reported by Strabel and Misztal (1999), who found a strong positive correlation of heritability (0.60 to 0.80) between first and second lactations upon random regression analysis. Similar to milk BHBA, the genetic variance components of parities 1 and 2 were considerably high until 15 DIM relative to mid lactation for all parities. For parity 3, the additive genetic variances were lower at the beginning and end of lactation than during mid lactation (140 to 220 DIM). Phenotypic variances were also high during early lactation, which was similar to the observations for milk BHBA, consequently resulting in low heritability during early lactation. Notably, heritability and additive genetic variance at mid lactation were very close over parities, indicating the consistency of heritability over parities in milk acetone.
The low heritabilties for both milk BHBA and acetone at 30 DIM for parity 3 are shown in Table 2. Since residual variance was fixed through lactation in the RRM, the sources of variation in heritability were variances in permanent environmental and additive genetic effects. The additive genetic variance of parity 3 was lowest among parities at 30 DIM, and further permanent environmental variance was highest, being 4 to 6 times greater than the permanent environmental variance of parities 1 and 2 in milk BHBA and 2 or 3 times greater than the permanent environmental variance of parities 1 and 2 in acetone. The average heritabilites for milk BHBA were 0.08, 0.11, and 0.09, respectively, for parities 1, 2, and 3, respectively. For milk BHBA, heritabilities ranged from 0.04 to 0.17 with a mean of 0.09 for the interval between 4 and 305 DIM in the three lactations. Koeck et al. (2014) found that the heritability estimates for milk BHBA at different stages of early lactation were between 0.14 and 0.29 and van der Drift et al. (2012) reported 0.16 during early lactation. The heritability estimates of this study were considerably lower than those of previous studies. Specifically, low phenotypic means and standard deviations (39.64 Ī¼mol/L and 56.99) were observed for the interval between 1 and 305 DIM (Table 1), while values of 80 Ī¼mol/L and 80 were reported for early lactation by van der Drift et al. (2012). The zero concentrations of milk BHBA were reduced to 19% from 31.5% when the interval between 1 and 30 was adopted instead of the interval between 1 and 305. Large variances in permanent environmental effects were observed during early lactation (Table 2). Some of the above statements could be reasons for the low heritabilities of milk BHBA, such as many zero concentrations of milk BHBA (Figure 2) and less phenotypic variations.
Wood et al. (2004) reported that heritability of milk acetone was less than 1% following log transformation of records. Additionally, the heritability for milk acetone was estimated to be 0.10 by van der Drift et al. (2012). As shown in Table 2, the heritabilities of acetone over lactations ranged from 0.06 to 0.29. The average heritabilites for milk acetone were 0.29, 0.29, and 0.22 for parities 1, 2, and 3, respectively. Unlike the results reported by van der Drift et al. (2012), the heritability traces of milk BHBA were lower than those of milk acetone in general (Figures 3 and 4; Table 2). Heritabilities of concentrations of ketone bodies in milk have been reported to vary from near zero to as high as 0.30 (Tveit et al., 1992).
Each parity was treated as a separate trait in RRMs, which may have resulted in large swings of heritabilities through lactation (Strabel and Misztal, 1999). Also it may be attributed to different distributions of records along the ranges of lactations at parities 1, 2, and 3. The criteria as an indicator for sub-clinical or clinical ketosis with milk acetone were suggested in the previous studies (Gustafsson and Emanuelson, 1996; van der Drift et al., 2012). Milk acetone with variation shown in Figure 2 may be more sensitive as an indicator of ketosis than milk BHBA with less variation. The higher heritability traces through lactations were obtained in milk acetone than milk BHBA, which is contrary to the results reported by van der Drift et al. (2012). Furthermore, the high heritability of milk acetone relative to milk BHBA in early lactation in this study was more useful as a trait for selection because of the high susceptibility to ketosis during early lactation (Baird, 1982; Sakha et al., 2006). These results suggest that genetic selection for low susceptibility of ketosis during early lactation is possible, which is concurrent with reports by van der Drift et al. (2012). Also the report that the occurrence ratio of ketosis in dairy cows is low in the first lactation (Baird, 1982) should be taken into account.
Van der Drift et al. (2012) found that concentrations of milk BHBA and milk acetone were correlated with concentrations of plasma BHBA (0.52) for the interval between 5 and 60 DIM. These findings may indicate that both traits are equally useful for selection. The genetic relationships between milk BHBA and milk acetone and between ketone bodies and milk production traits were investigated. The breeding values of individuals for each DIM were estimated based on regression coefficients for individual breeding values. The covariance and correlations of breeding values for two traits from 4 to 305 DIM were estimated rather than the covariance functions through lactation (Meyer and Hill, 1997). The trajectory of correlations between breeding values of the traits through lactation were drawn and then shown in Figure 5, 6, 7, and 8. The statics of the correlations were shown in Table 3.
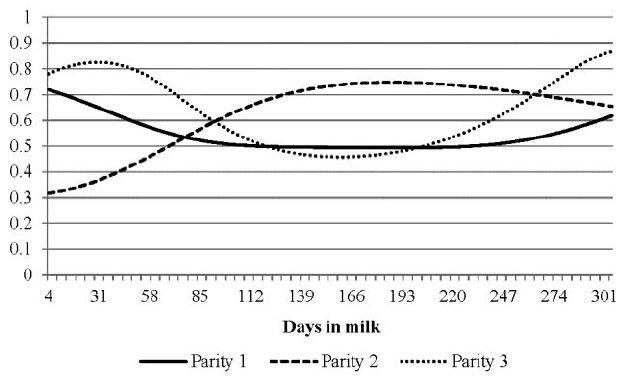
The trace of correlations between milk BHBA and acetone for each parity is shown in Figure 5. Both traits were ketone bodies and considered indicator traits for ketosis (Wood et al., 2004; van der Drift et al., 2012). The correlations were high at both ends of lactations in parities 1 and 3 and the shapes of the traces were similar. The correlations were high in mid lactation for parity 2. The average correlation for primiparous cows was lower than the overall average (Table 3). The averages and SDs of parities 2 and 3 were similar, but the shapes of their traces were dramatically different (Figure 5). During early lactation, the sizes of correlations were in the order of parities 3, 1, and 2. The overall average of correlations was 0.60, ranging from 0.32 to 0.87. Van der Drift et al. (2012) found that genetic correlations between the concentrations of milk BHBA and milk acetone was 0.90 for the interval between 5 and 60 DIM. The correlations were highest (over 0.8) from week 4 to 6 of lactation 3, in which ketosis most frequently occurs (Baird, 1982; Sakha et al., 2006; Vosman et al., 2015). In general, both traits as indicators of ketosis showed strong positive correlations of breeding values through lactations of three parities. However, milk BHBA and acetone were genetically less related in early lactation of second parity and showed increased genetic correlation afterward. Even if the exact cause may not be conjectured at this stage, we should pursue to find it with biological means or statistically with large data set later. The decision for genetic evaluation of ketosis resistance includes choice of traits to be evaluated as indicatory traits that would be milk BHBA or acetone, or both. The genetic relation of milk BHBA and acetone shown in Figure 5 should be taken into account before decision.
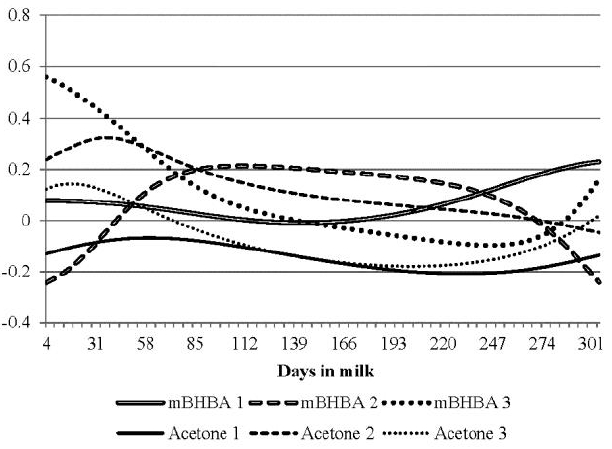
The breeding value correlations of two ketone bodies with breeding values of TD protein % were estimated to investigate genetic relationships and are shown in Figure 6. The correlations for milk BHBA with protein % did not show any pattern through lactations of parities 1, 2, and 3. The average of the overall correlations was 0.08 with a range of ā0.24 to 0.56. The average was 0.06 for primiparous cows with a range of ā0.01 to 0.23, in which the trace of correlations was lower than those for parities 2 and 3. The correlations of milk BHBA with protein % for parity 2 were generally low (no greater deviation of 0.2 from 0). In general, the milk BHBA relationship with protein % was insignificant for all parities, but a significant relationship during early lactation for parities 2 and 3. High occurrence of ketosis takes place commonly at early lactation of parities 2 and 3. Therefore it is hard to explain the contradictory genetic correlations in both parities. However, the trace of milk BHBA and protein % in second parity had similar shape with that of second parity in Figure 5. Considering that all traces for milk BHBA and acetone had similar shape except for milk BHBA at second parity, low genetic correlation between milk BHBA and acetone early in second parity in Figure 5 seems to be due to milk BHBA.
The correlations of acetone with protein % were generally weak through lactation for all parities. The correlations of milk acetone with protein % were mostly negative in parities 1 and 3, but not in parity 2. The overall correlation was ā0.03 with a range of ā0.21 to 0.32 (Table 3). The average was 0.12 for parity 2, with a range of ā0.05 to 0.32, in which the trace of correlations for parity 2 was greater than those for parities 1 and 3 (ā0.14 and ā0.07 on average, respectively). The correlation of acetone with protein % was not significant through lactation for parity 3. There was no strong genetic propensity for protein % to be correlated with concentration of either ketone bodies. The correlations of both ketone bodies were mostly low (Ā±0.2) through lactation and parities. Koeck et al. (2012) found that correlation of ketosis with milk composition traits was not significant and was close to zero at 31 to 60 DIM.
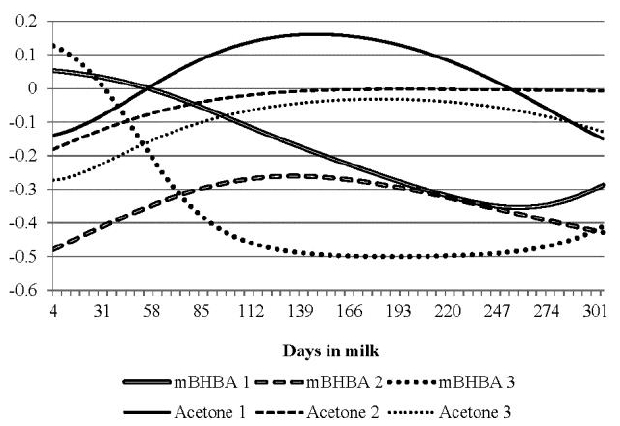
Milk production is given metabolic priority over other physiological processes, and ketosis is a metabolic disorder of fat in the body of milking cows. Koeck et al. (2012) found that ketosis was significantly correlated with fat % (0.33) at 5 to 30 DIM in primiparous cows. The breeding value correlations of both ketone bodies with breeding values of TD fat % were estimated and the traces of correlations were drawn in Figure 7. The correlations for milk BHBA were mostly positive during early lactation in all parities (over 0.4 at 4 DIM for parities 1 and 2). The correlations for parity 2 became negative from 60 DIM and decreased to ā0.44 at 305 DIM, while the correlations for parity 1 became positive during late lactation. In addition, the correlations of milk BHBA for parity 3 were weak and positive through lactation. The overall average of correlations was ā0.02 with a range of ā0.44 to 0.61. The average was 0.06 for parity 1 with a range of ā0.27 to 0.61. In general, the ranges of the correlation were wide in parities 1 and 2, while those in the parity were not. Variations of milk BHBA correlation with fat % were large among parities and through lactation.
The correlations of milk acetone with fat % were mostly negative except for early lactation of primiparous cows. The overall average of correlations of milk acetone with fat % was ā0.20 with a range of ā0.44 to 0.28. The average was ā0.21 for primiparous cows with a range of ā0.41 to 0.28, in which the trace of correlations (0.28) at 4 DIM was greater than any period and parity.
Fat % showed a positive genetic relationship with the concentration of ketone bodies during early lactation, in which the ratio of ketosis occurrence in dairy cows is usually high. Moreover, the correlations of ketone bodies with fat % showed high variations through lactation and over parities. The genetic correlation traces of both milk BHBA and acetone with fat % were similar at the first and third parity, but the traces of first and third parity were different each other. It seems to be consistent with the relation between milk BHBA and acetone as shown in Figure 5.
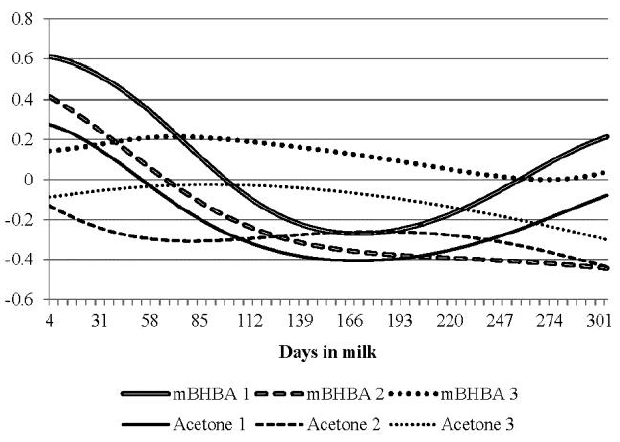
With clinical or subclinical ketosis, milk production tends to decrease because the cow is unable to synthesize sufficient lactose due to the deficiency in glucose (Wood et al., 2004). The breeding value correlations of milk BHBA and acetone concentrations with breeding values of TD milk yield are shown in Figure 8 for each parity. The correlations for milk BHBA were mostly negative, except for parities 1 and 3 during early lactation. Even though those correlations were not high (<0.13), these findings showed a positive genetic relationship with milk yield at early lactation. Eventually, high concentrations of milk BHBA appear to lead to low milk production later. The correlations for parity 3 decreased rapidly to ā0.50 at 150 DIM, while the correlations for parity 1 slowly became negative at 57 DIM and decreased to ā0.35 at 260 DIM. The correlations for parity 2 increased from ā0.48 at 4 DIM to ā0.26 at 135 DIM, which was the peak of the trace. The overall average of the correlations was ā0.30 with a range of ā0.50 to 0.13. The average for parity 3 was ā0.38 with a range of ā0.50 to 0.13. Koeck et al. (2014) found with a set of data including about 3000 records that the Pearson correlations of sire breeding values of milk yield and concentrations of daughter milk BHBA ranged from 0.06 to 0.22 between 5 and 100 DIM, which differ from findings of this study.
The genetic correlations of milk acetone with milk yield were weakly positive for primiparous cows except for the correlation between early and late lactation (Figure 8), while the correlations in parities 2 and 3 were negative and became low at mid lactation. In general, the correlations at early lactation were negative for all parities (ā0.14, ā0.18, and ā0.27 at 4 DIM for parities 1, 2, and 3, respectively) and the correlations of parities 2 and 3 became insignificant from about 40 DIM. The overall average of correlations was ā0.03 with a range of ā0.27 to 0.16. The average for parity 1 was 0.04 with a range of ā0.15 to 0.16, in which there were only positive curves with the peak (0.16) at 148 DIM. There was a genetic propensity in that milk yield showed a negative genetic relationship with concentration of milk acetone during early lactation, when the occurrence ratio of ketosis in dairy cows is high. These correlations became low or positive at mid lactation.
Daily milk yield tends to be reduced in cows with increased concentrations of acetone, but the responses are not entirely consistent (Gustafsson and Emanuelson, 1996). In this study, milk BHBA mostly showed a clear negative correlation with milk yield. However, the correlations of milk acetone with milk yield were positive or negative, but low. A typical symptom of clinical or subclinical ketosis is decreased milk yield (Enjalbert et al., 2001; Oetzel, 2012), in which high concentrations of ketone bodies may be associated with low milk yields. Conversely, Koeck et al. (2014) and van der Drift et al. (2012) suggested that a higher genetic merit for milk yield was associated with higher milk BHBA, and therefore a greater susceptibility to hyperketonemia. These findings suggest that milk yield and concentrations of ketone bodies interact with each other and consequently show a limited genetic relationship. There was no strong genetic propensity for milk yield with concentration for either ketone body. Nonetheless, care must be taken before any conclusions are drawn based on these findings since milk yield can be affected by many factors.
Oetzel (2007) suggested that herd-based testing with animals of about 5 to 50 DIM should be conducted to evaluate ketosis. Unlike milk BHBA, the concentration of milk acetone showed a continuous bell shape frequency for cows through 305 DIM or for the interval between 5 and 50. Further variation of milk acetone may provide a more sensitive indication of ketosis than that of milk BHBA, and the heritability traces through lactations were higher in milk acetone than in milk BHBA. Even though milk production traits with milk ketone bodies showed limited genetic association, concentrations of BHBA and acetone in milk can be evaluated as part of the set of procedures now used to evaluate concentrations of fat, protein, and somatic cells in milk. Additionally, a breeding scheme should be established to enable selection that reduces the occurrence of ketosis.
ACKNOWLEDGMENTS
This work was carried out with the support of the Development of Systems and Techniques for Improving Resistance and Early Diagnosis of Ketosis in Dairy Cattle project of the National Institute of Animal Science, RDA, Korea (Project No PJ01046501).
FigureĀ 2
Distribution of the concentrations of milk Ī²-hydroxybutyric acid (Avg = 33.8, SD = 56.3), milk acetone (Avg = 125.8, SD = 87.8), milk acetone of cows with zero milk Ī²-hydroxybutyric acid (MBHBA) at 305 days in milk, and milk acetone of cows at 5 to 50 DIM. Avg, average; SD, standard deviation; DIM, days in milk.
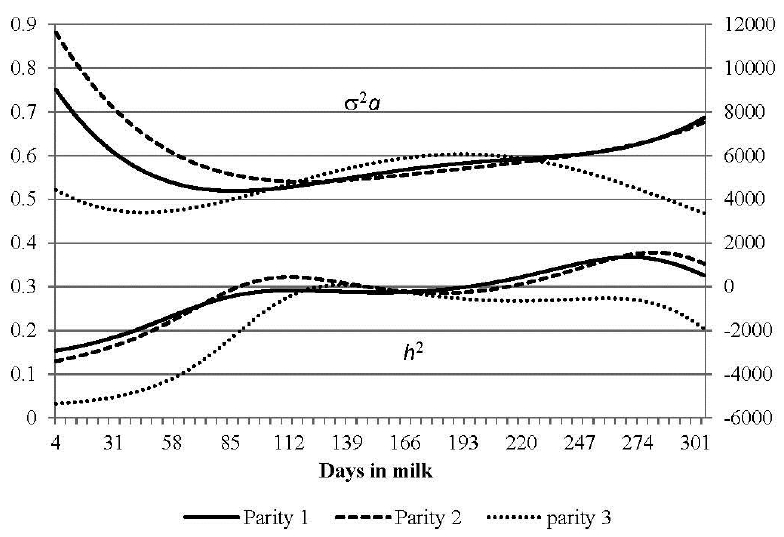
FigureĀ 3
Estimates of heritability (h2) and additive genetic variance (Ļ2a) for milk Ī²-hydroxybutyrateacid according to parities.

FigureĀ 4
Estimates of heritability (h2) and additive genetic variance (Ļ2a) for milk acetone according to parities.
FigureĀ 5
Breeding correlations of milk Ī²-hydroxybutyrate acid with milk acetone according to parities.
FigureĀ 6
Breeding value correlations of milk Ī²-hydroxybutyrateacid (mBHBA) and milk acetone with protein % according to parities (numbers in the abbreviates).
FigureĀ 7
Breeding value correlations of milk Ī²-hydroxybutyrate acid (mBHBA) and milk acetone with fat % according to parities (numbers in the abbreviates).
FigureĀ 8
Breeding value correlations of milk Ī²-hydroxybutyrateacid (mBHBA) and milk acetone with milk yield according to parities (numbers in the abbreviation)
TableĀ 1
Basic statistics of the analyzed data
TableĀ 2
Estimates of genetic (G), permanent environmental (P) variance components and heritability (h2) for milk Ī²-hydroxybutyrate acid and milk acetone at different days in milk of the parities in Holstein cattle
TableĀ 3
Basic statics of correlations of the breeding values of milk Ī²-hydroxybutyrate acid (BHBA) and milk acetone with breeding values of milk yield, protein % and fat % according to days in milk and parities in Holstein cattle
REFERENCES
Andersson L. 1984. Concentrations of blood and milk ketone bodies, blood isopropanol, and plasma glucose in dairy cows in relation to the degree of hyperketonaemia and clinical signs. Zentralbl Veterinarmed A 31:683ā693.


Andersson L. 1988. Subclinical ketosis in dairy cows. Vet Clin North Am Food Anim Pract 4:233ā251.


Baird GD. 1982. Primary ketosis in the high-producing dairy cow: Clinical and subclinical disorders, treatment, prevention, and outlook. J Dairy Sci 65:1ā10.


Cho KH, Park B, Choi J, Choi T, Choy Y, Lee S, Cho C. 2013. Development of international genetic evaluation models for dairy cattle. J Anim Sci Technol 55:1ā6.

Duffield TF, Kelton DF, Leslie KE, Lissemore KD, Lumsden JH. 1997. Use of test day milk fat and milk protein to detect subclinical ketosis in dairy cattle in Ontario. Can Vet J 38:713ā718.


Enjalbert F, Nicot MC, Bayourthe C, Moncoulon R. 2001. Ketone bodies in milk and blood of dairy cows: Relationship between concentrations and utilization for detection of subclinical ketosis. J Dairy Sci 84:583ā589.


Gustafsson AH, Emanuelson U. 1996. Milk acetone concentration as an indicator of hyperketonaemia in dairy cows: The critical value revised. Anim Sci 63:183ā188.

Jamrozik J, Schaeffer LR, Dekkers JCM. 1997. Genetic evaluation of dairy cattle using test day yields and random regression model. J Dairy Sci 80:1217ā1226.


Jamrozik J, Schaeffer LR. 2002. Bayesian comparison of random regression models for test-day yields in dairy cattle. In : Proceedings of the 7th World Congress on Genetics Applied to Livestock Production; Montpellier, France. Communication no. 01ā03
Kessel S, Stroehl M, Meyer HH, Hiss S, Sauerwein H, Schwarz FJ, Bruckmaier RM. 2008. Individual variability in physiological adaptation to metabolic stress during early lactation in dairy cows kept under equal conditions. J Anim Sci 86:2903ā2912.


Kirkpatrick M, Heckman N. 1989. A quantitative genetic model for growth, shape, reaction norms, and other infinite-dimensional characters. J Math Biol 27:429ā450.


Kirkpatrick M, Lofsvold D, Bulmer M. 1990. Analysis of the inheritance, selection and evolution of growth trajectories. Genetics 124:979ā993.




Koeck A, Miglior F, Kelton DF, Schenkel FS. 2012. Health recording in Canadian Holsteins: Data and genetic parameters. J Dairy Sci 95:4099ā4108.


Koeck A, Jamrozik J, Schenkel FS, Moore RK, Lefebvre DM, Kelton DF, Miglior F. 2014. Genetic analysis of milk Ī²-hydroxybutyrate and its association with fat-to-protein ratio, body condition score, clinical ketosis, and displaced abomasum in early first lactation of Canadian Holsteins. J Dairy Sci 97:7286ā7292.


Mantysaari EA, GrĆhn YT, Quaas RL. 1991. Clinical ketosis: phenotypic and genetic correlations between occurrences and with milk yield. J Dairy Sci 74:3985ā3993.


McArt JAA, Nydam DV, Oetzel GR. 2012. Epidemiology of subclinical ketosis in early lactation dairy cattle. J Dairy Sci 95:5056ā5066.


Meyer K, Hill WG. 1997. Estimation of genetic and phenotypic covariance functions for longitudinal or ārepeatedā records by restricted maximum likelihood. Livest Prod Sci 47:185ā200.

Meyer K. 2015. WOMBAT A program for Mixed Model Analyses by Restricted Maximum Likelihood-User Notes. Version 1.0. University of New England; Armidale, Australia:
Nielsen NI, Friggens NC, Chagunda MGG, Ingvartsen KL. 2005. Predicting risk of ketosis in dairy cows using in-line measurements of Ī²-hydroxybutyrate: A biological model. J Dairy sci 88:2441ā2453.


Oetzel GR. 2007. Herd-level ketosis ā diagnosis and risk factors. In : Proceedings of the 40th Annual Conference of Bovine Practitioners; Vancouver, BC, Canada. p. 67ā91.
Oetzel GR. 2012. Understanding the Impact of Subclinical Ketosis. In : Cornell Nutrition Conference Proceeding; Ithaca, NY, USA. p. 12ā21.
Pool MH, Janss LLG, Meuwissen THE. 2000. Genetic parameters of legendre polynomials for first parity lactation curves. J Dairy Sci 83:2640ā2649.


Sakha M, Ameri M, Rohbakhsh A. 2006. Changes in blood b-hydroxybutyrate and glucose concentrations during dry and lactation periods in Iranian Holstein cows. Comp Clin Path 15:221ā226.

Strabel T, Misztal I. 1999. Genetic parameters for first and second lactation milk yields of Polish Black and White cattle with random regression test-day models. J Dairy Sci 82:2805ā2810.


Tveit B, Lingaas F, Svendsen M, Sjaastad OV. 1992. Etiology of acetonemia in Norwegian cattle I. Effect of ketogenic silage, season, energy level, and genetic factors. J Dairy Sci 75:2421ā2432.


van der Drift SGA, van Hulzen KJE, Teweldemedhn TG, Jorritsma R, Nielen M, Heuven HCM. 2012. Genetic and nongenetic variation in plasma and milk Ī²-hydroxybutyrate and milk acetone concentrations of early-lactation dairy cows. J Dairy Sci 95:6781ā6787.


Van Der Werf JHJ, Goddard ME, Meyer K. 1998. The use of covariance functions and random regressions for genetic evaluation of milk production based on test day records. J Dairy Sci 81:3300ā3308.


Van Dorp RT, Martin SW, Shoukri MM, Noordhuizen JP, Dekkers JC. 1999. An epidemiologic study of disease in 32 registered Holstein dairy herds in British Columbia. Can J Vet Res 63:185ā192.


Vosman JJ, de Jong G, Eding H, Knijn H. 2015. Genetic evaluation for ketosis in the Netherlands based on FTIR measurements. In : Proceedings of the 2015 Interbull Meeting; Orlando, FL, USA. Bulletin No. 49





 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print





