 |
 |
| Anim Biosci > Volume 28(6); 2015 > Article |
|
Abstract
The purpose of this study was to investigate the alternative splicing in equine cordon-bleu WH2 repeat protein-like 1 (COBLL1) gene that was identified in horse muscle and blood leukocytes, and to predict functional consequences of alternative splicing by bioinformatics analysis. In a previous study, RNA-seq analysis predicted the presence of alternative spliced isoforms of equine COBLL1, namely COBLL1a as a long form and COBLL1b as a short form. In this study, we validated two isoforms of COBLL1 transcripts in horse tissues by the real-time polymerase chain reaction, and cloned them for Sanger sequencing. The sequencing results showed that the alternative splicing occurs at exon 9. Prediction of protein structure of these isoforms revealed three putative phosphorylation sites at the amino acid sequences encoded in exon 9, which is deleted in COBLL1b. In expression analysis, it was found that COBLL1b was expressed ubiquitously and equivalently in all the analyzed tissues, whereas COBLL1a showed strong expression in kidney, spinal cord and lung, moderate expression in heart and skeletal muscle, and low expression in thyroid and colon. In muscle, both COBLL1a and COBLL1b expression decreased after exercise. It is assumed that the regulation of COBLL1 expression may be important for regulating glucose level or switching of energy source, possibly through an insulin signaling pathway, in muscle after exercise. Further study is warranted to reveal the functional importance of COBLL1 on athletic performance in race horses.
The Thoroughbred is the most famous breed for horse racing because of its speed and agility. The Thoroughbred industry as a large agribusiness creates annual revenue of around $34 billion in the United States alone, and provides about 470,000 jobs related to the network of farms, training centers and race tracks. Genes related to their athletic phenotypes have not been identified, although physical and physiological adaptations related to the potentials of their superior athleticism are well characterized (Gu et al., 2009).
In a previous study, the whole transcriptome for blood and muscle tissues of six Thoroughbred horses before and after 30 min of exercises were analyzed using RNA-sequencing. From the analysis, 32,361 unigene clusters were identified, of which 20,428 have been newly reported as belonging to the equine gene model. Differentially expressed genes were also found in blood (142) and muscle (1,163). In addition, alternative splicing forms of cytoplasmic dynein 1 (DYNC1), AXL receptor tyrosine kinase (AXL), pleckstrin homology domain containing, family G (with RhoGef Domain) member 1 (PLEKHG1), and the cordon-bleu WH2 repeat protein-like 1 (COBLL1) showed reversed expression patterns before and after exercise in skeletal muscle (Park et al., 2012).
The COBLL1 gene was originally cloned from the human brain cDNA library and is highly expressed in most human tissues including lung, liver, kidney, pancreas, ovary, spinal cord and brain (Nagase et al., 1999). COBLL1 was identified as another class of Cobl (Carroll et al., 2003). COBLL1 has been reported as a negative regulator of apoptosis in tumor cells (Gordon et al., 2011) and has been associated with metabolic diseases related to insulin resistance (Mancina et al., 2013). Despite various studies on COBLL1, the specific roles of both genes are still unclear, especially in horses. In this study, we focused on the analysis of the COBLL1 gene which was one of the alternatively spliced genes during exercise. We confirmed alternative splicing forms of COBLL1 in various horse tissues and examined the expression pattern of each alternative splicing form in response to exercise.
Three male Thoroughbred horses aged 5, 9, and 10 years, which were maintained at Ham-an Racing Horse Resort and Training Center, were used to obtain the blood and skeletal muscle samples before and after exercise. Exercise was performed by trotting at the speed of 13 km/h for 30 min. The National Institute of Subtropical Agriculture, Rural Development Administration, provided three Jeju horses which were used for collecting the tissue samples such as skeletal muscle, kidney, thyroid, lung, appendix, colon, spinal cord and heart. All tissue samples were kept in liquid nitrogen tank until extraction of RNA was done. All procedures were conducted by following the protocol that had been reviewed and approved by the Institutional Animal Care and Use Committee at Pusan National University (protocol numbers: PNU-2013-0417, PNU-2013-0411).
Trizol reagent (Invitrogen, Carlsbad, CA, USA) was used to extract total RNA from tissue samples (skeletal muscle, kidney, thyroid, lung, colon, spinal cord, and heart) and leukocytes after exercise in horse, according to the Invitrogen manual. In order to prevent contamination of genomic DNA, RNase-free DNase kit (Qiagen, Venlo, Netherlands) was used according to the manufacturerŌĆÖs operating manual. Total RNA quantification was performed by using a NanoDrop ND-1000 Spectrophotometer. The cDNAs were synthesized in a reaction with oligo-dT primers, moloney-murine leukemia virus reverse transcriptase (Promega, Madison, WI, USA), RNase inhibitor (Promega, USA) and RNase-free ddH2O, which were incubated at 37┬░C for 4 h.
The horse COBLL1 transcripts were analyzed by reverse transcription polymerase chain reaction (RT-PCR) amplification. The original horse COBLL1 transcripts were amplified by the primer pairs S (5ŌĆ▓-ATTCCCTGGAGTCAGTCCATGCAG-3ŌĆ▓) and AS (5ŌĆ▓-GGGAAGTGCCTAAAGATGAGGCTG-3ŌĆ▓). These primers were designed using the PRIMER3 program (http://frodo.wi.mit.edu/primer3/) and sequence information was obtained from the UCSC genome browser (Accession number: NM-001081812.1). The expected sizes of PCR products were 276 bp and 156 bp for COBLL1a and COBLL1b, respectively. The RT-PCR conditions were as follows: an initial step of 94┬░C for 4 min, 35 cycles of 94┬░C for 40 s, 56┬░C for 1 min, and 72┬░C for 2 min were performed. The RT-PCR products were analyzed after electrophoresis on a 1.5% agarose gel. COBLL1a and COBLL1b were cloned into pGEM-T Easy-cloning vector (Promega, USA) and their nucleotide sequences were determined by Sanger sequencing. The sequenced nucleotides were aligned by Blast (National Center for Biotechnology Information; NCBI, Bethesda, MD, USA).
To analyze the expression level of COBLL1 alternative splicing isoforms in muscle before and after exercise, real-time quantitative polymerase chain reaction (RT-qPCR) was conducted by using the BioRad CFX-96 machine (BioRad, Hercules, CA, USA). Primer sequences were designed to detect each type of alternative splicing forms. Two primer pairs, Cobll1a-S (5ŌĆ▓-ATCAGTAGACCCGCACGGAA-3ŌĆ▓) and Cobll1a-AS (5ŌĆ▓-ACACTGACGGAGGAAACCTC-3ŌĆ▓) for COBLL1a, and Cobll1b-S (5ŌĆ▓-AGGGTCTGTCCAGCCCAGCT-3ŌĆ▓) and Cobll1b-AS (5ŌĆ▓-CCATAACGTGTGGTCCCTGTCC-3ŌĆ▓) for COBLL1b, were used to specifically detect each isoform. Each reaction was executed in a total 25 ╬╝L of mixture containing 14 ╬╝L of SYBR green master mix, 2 ╬╝L of forward primer (5 pmol), 2 ╬╝L of reverse primer (5 pmol), 5 ╬╝L of distilled water, and 2 ╬╝L (50 ng/╬╝L) of cDNA. The PCR conditions were at 94┬░C for 5 min of pre-denaturation step, 39 cycles of 94┬░C for 20 s, 56┬░C for 20 s and 72┬░C for 30 s, and followed by 72┬░C for 10 min as a final step. All samples were measured in triplicate to ensure reproducibility, and Ct value was used to calculate the fold change by using the 2ŌłÆ╬ö╬öCt method (Livak and Schmittgen, 2001). The glyceraldehyde 3-phosphate dehydrogenase gene was used for reference.
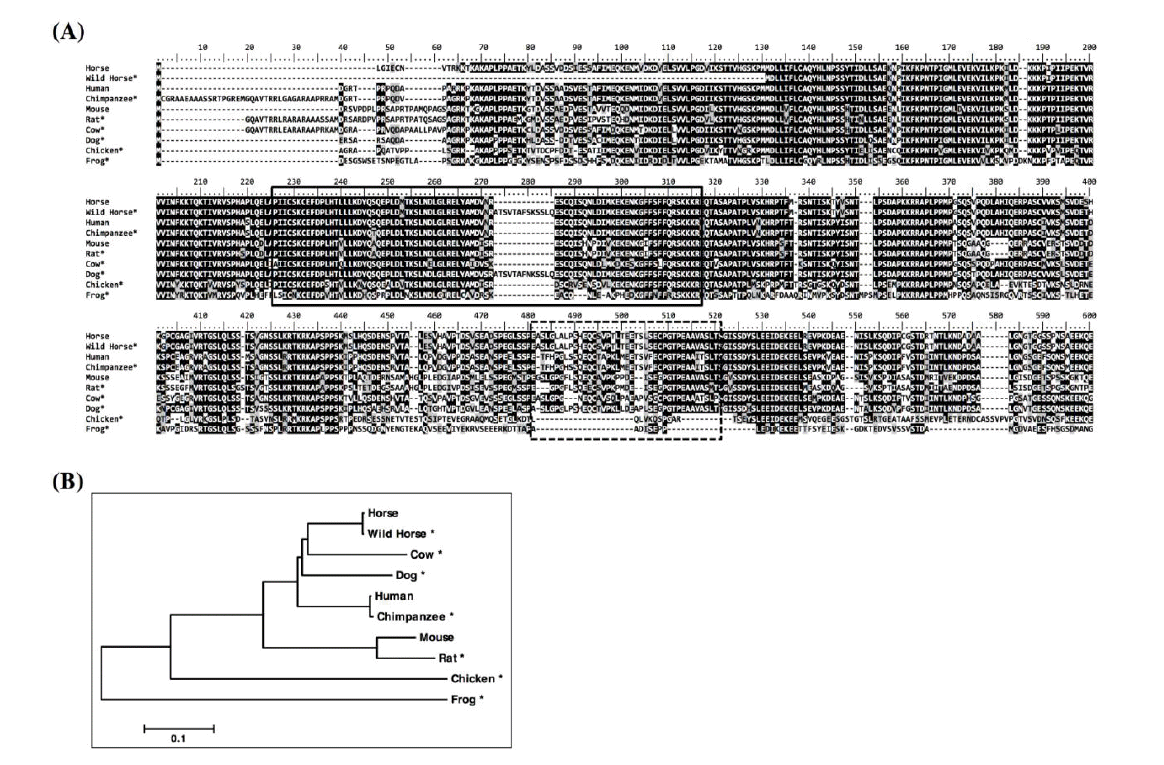
The amino acid sequences of COBLL1 of various species were retrieved from the Gene database of NCBI (wild horse, XP_008531702.1; human, NP_055715.3; chimpanzee, XP_003309326.1; mouse, NP_795999.2; rat, XP_006234363.1; cow, XP_002685420.3; dog, XP_005640312.1; chicken, XP_004942872.1; frog, XP_002939319.2). The amino acid sequences were aligned and used for generating phylogenetic tree by using the MUSCLE and Neighbor-Joining methods, respectively, in MEGA6 program (Tamura et al., 2013). The web-based protein prediction program, PredictProtein (www.predictprotein.org), was used to predict the secondary and tertiary structure (Rost et al., 2004).
The horse COBLL1 gene has 14 exons and 13 introns spanning about 60.5 kb on the chromosome 18 and six transcriptional variants had been predicted in NCBI Gene database. When the amino acid sequences of COBLL1 were compared among various species, COBLL1 showed higher identity at the N-terminal sequences containing a Cordon-bleu ubiquitin-like domain than at the C-terminal (Figure 1A, solid box). Interestingly, one of the alternative splicing forms of COBLL1 gene had been observed within the Cordon-bleu ubiquitin-like domain of various species including horse, wild horse, human, dog, chimpanzee, and chicken. However, this variant did not seem to affect the function of the domain as it was predicted as the Cordon-bleu ubiquitin-like domain. Another alternative splicing was observed between exon 8 and 10 (with skipping of exon 9) in many species including horse, wild horse, human, mouse, rat, dog, and chicken (Figure 1A, dashed box). The similarity of amino acid sequences around these sequences was relatively lower than the other N-terminal region shown in Figure 1A. Horse COBLL1 was evolutionarily close to cow COBLL1 while largely divergent from frog (Figure 1B). COBLL1 has been known to regulate apoptosis or to be associated with metabolic diseases related to insulin resistance. It has been reported that alternative splicing forms of a gene may be differentially expressed under conditions of wound response, tumorogenesis, embryonic development and response to external stimuli (Vitale et al., 1994; Shin and Manley, 2004; Huang et al., 2005; Gardina et al., 2006; Lynch, 2007). Therefore, we assumed that COBLL1 was evolutionally conserved through vertebrates, and the expression patterns of the transcript variants were different according to conditions during stimulation.
In a previous study, two alternative splicing forms of COBLL1 transcripts were identified in horses (Park et al., 2012). And they are found in both NCBI (XM_005601553.1 and XM_005601555.1) and Ensembl (ENSECAT00000027104 and ENSECAT00000027105) data base and designated to COBLL1a and COBLL1b, respectively (Figure 2A). Specific primers were designed for binding to exon 7 and 8 forward primers and exon 10 reverse primer to confirm the expression of COBLL1a and COBLL1b in horse. When PCR for horse COBLL1 transcripts in blood was performed, two transcriptional variants with different sizes (277 bp and 157 bp) were detected (Figure 2B). Each amplicon for COBLL1a and COBLL1b transcript was detected with 120 bp difference in size by gel electrophoresis. By sequencing analysis with the amplicons, the deletion of exon 9 was observed in COBLL1b as expected (Figure 2C).
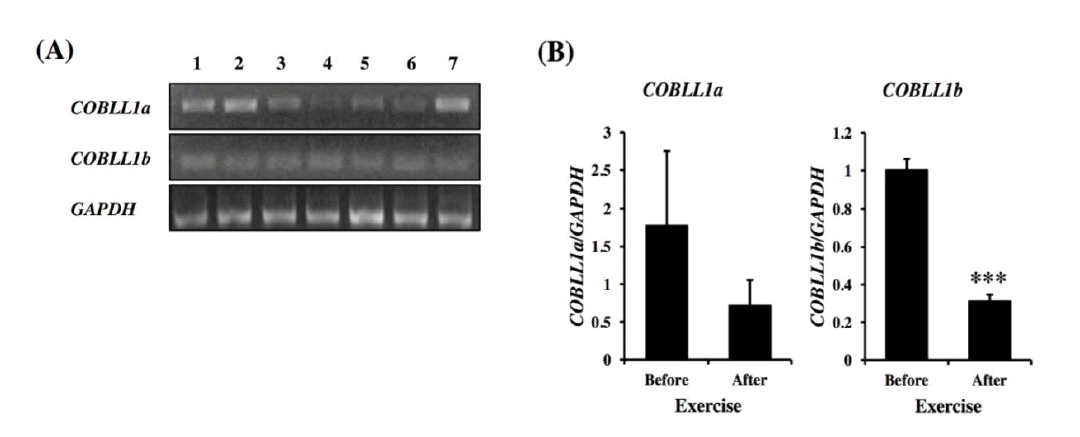
To investigate the expression patterns of COBLL1 variants in horse tissues, RT-PCR was conducted with cDNAs from the various tissues including kidney, spinal cord, heart, thyroid, skeletal muscle, colon and lung (Figure 3A). The COBLL1a showed strong expression in the kidney, spinal cord and lung, moderate expression in the heart and skeletal muscle, and low expression in the thyroid and colon. Unlike COBLL1a expression, COBLL1b was expressed evenly in all tissues. The expression patterns analysis of COBLL1a and COBLL1b in muscle tissues before and after exercise revealed that the expression of both COBLL1a and COBLL1b decreased after exercise, but only COBLL1b showed significant decrease (Figure 3B). Thus, it is speculated that COBLL1a and COBLL1b responded similarly to the effects of exercise in skeletal muscle.
According to a recent genome-wide association study, the human COBLL1 gene was found to be located at the genomic region associated with a metabolic traits such as fasting insulin level, type 2 diabetes, and triacylglycerol metabolism (Manning et al., 2012; Albrechtsen et al., 2013; Desmarchelier et al., 2014). In addition, the C allele of COBLL1 is functionally linked to lower serum insulin levels and insulin resistance in humans (Mancina et al., 2013). These two reports suggest that COBLL1 may play an important role in metabolic pathways, especially in the regulation of glucose uptake by insulin. Generally, exercise increases the uptake of glucose mediated by up-regulation of GLUT4 protein content and leads to enhanced insulin resistance in muscle (Goodyear and Kahn, 1998; Borghouts and Keizer, 2000). Given the importance of glucose metabolism in exercise, it is of interest that the regulation of COBLL1 expression maybe important for regulating glucose level or switching of energy source, possibly through an insulin signaling pathway, in muscle after exercise. Taken together, this information may be helpful to investigate the role of equine COBLL1 on the metabolic changes related to the athletic performance of racehorses and to develop COBLL1 as a useful biomarker for athletic performance.
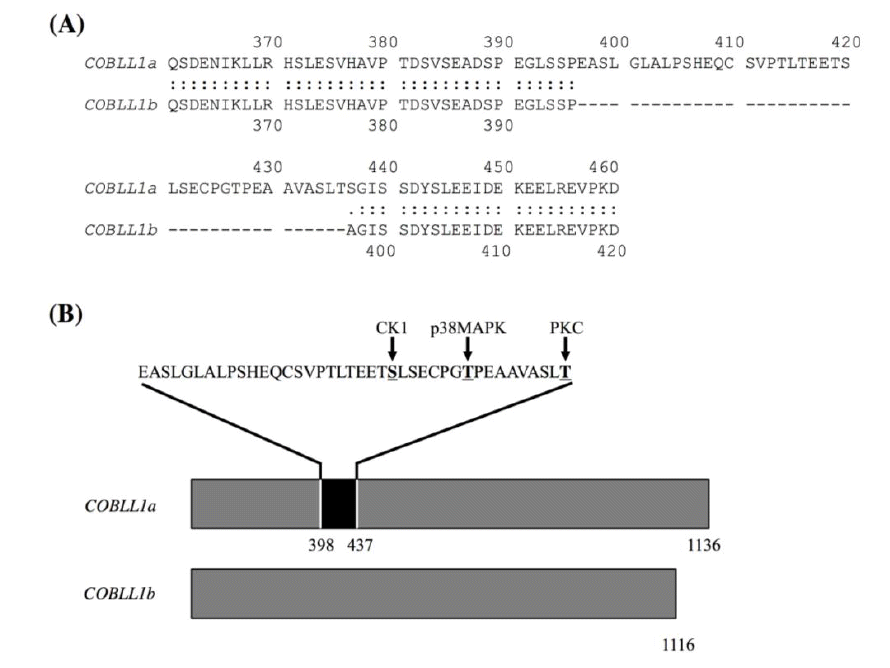
In analyzing the amino acid sequences of COBLL1 splicing forms, COBLL1b has a deletion of 40 amino acids due to the exclusion of exon 9 by alternative splicing compared to COBLL1a (Figure 4A). These sequences were predicted to include three putative phosphorylation sites by casein kinase 1, p38 mitogen-activated protein kinase, and protein kinase C (Figure 4B). In many cases, protein phosphorylation is closely related to the control of protein stabilization, serving as a marker that triggers subsequent ubiquitination, in particular where ubiquitination leads to degradation (Treier et al., 1994; Fuchs et al., 1996; Magnani et al., 2000). Therefore, COBLL1a could be elaborately controlled in each tissue, whereas COBLL1b could be stable in various tissues due to lack of these phosphorylation sites by alternative splicing (Figure 3A).
ACKNOWLEDGMENTS
This work was supported by a grant from the Next-Generation BioGreen 21 Program (No. PJ011173, PJ011044), Rural Development Administration, Republic of Korea. Authors thank Ms S.H. Moon, MSc, for editorial help.
Figure┬Ā1
Analysis of amino acid sequences and phylogenetic tree of COBLL1 among various species. (A) Alignment of N-terminal amino acid sequences of COBLL1 from various species. The sequences were aligned by the MUSCLE method in MEGA6 program. The N-terminal sequences showed higher similarity among the species compared to C-terminal sequences. The Cordon-bleu ubiquitin-like domain is marked by solid box and the sequences deleted by alternative splicing are marked by dashed box. (B) Phylogenetic tree of COBLL1. The phylogenetic tree was made with the full amino acid sequence of each species by Neighbor-Joining method after aligned by the MUSCLE method in MEGA6 program. Horse COBLL1 was similar to cow and dog while divergent from frog and chicken. COBLL1, cordon-bleu WH2 repeat protein-like 1.
Figure┬Ā2
Alternative splicing of equine COBLL1 gene. (A) Genomic structures of COBLL1 alternative splicing variants. Horse COBLL1 gene, which has 14 exons, has two major alternative splicing forms, COBLL1a and COBLL1b. The COBLL1b has a deletion of exon 9 by alternative splicing. (B) Confirmation of COBLL1 alternative splicing variants. The alternative splicing variants of COBLL1 gene, COBLL1a and COBLL1b, were amplified by RT-PCR with a primer set flanking the alternative splicing region. (C) Sequencing of alternative splicing region. The RT-PCR products were sequenced and confirmed that exon 9 was deleted in COBLL1b by alternative splicing. COBLL1, cordon-bleu WH2 repeat protein-like 1; RT-PCR, reverse transcription polymerase chain reaction.

Figure┬Ā3
Expression patterns of equine COBLL1 alternative splicing variants. (A) Expression of COBLL1a and COBLL1b genes were analyzed in various tissues. The numbers shown in each lane represent tissues in order of kidney, spinal cord, heart, thyroid, skeletal muscle, colon, and lung. (B) Expression of COBLL1 alternative splicing variants was analyzed by RT-qPCR in skeletal muscle before and after exercise. COBLL1, cordon-bleu WH2 repeat protein-like 1; RT-qPCR, real-time quantitative polymerase chain reaction.
Figure┬Ā4
The comparison of amino acid sequences between COBLL1a and COBLL1b proteins. The amino acid sequences of COBLL1a and COBLL1b proteins were aligned (A), and the putative phosphorylation positions within the alternative splicing region were predicted (B). COBLL1, cordon-bleu WH2 repeat protein-like 1.
REFERENCES
Albrechtsen A, Grarup N, Li Y, Sparso T, Tian G, Cao H, Jiang T, Kim SY, Korneliussen T, Li Q, et al. 2013. Exome sequencing-driven discovery of coding polymorphisms associated with common metabolic phenotypes. Diabetologia 56:298ŌĆō310.



Borghouts LB, Keizer HA. 2000. Exercise and insulin sensitivity: A review. Int J Sports Med 21:1ŌĆō12.


Carroll EA, Gerrelli D, Gasca S, Berg E, Beier DR, Copp AJ, Klingensmith J. 2003. Cordon-bleu is a conserved gene involved in neural tube formation. Dev Biol 262:16ŌĆō31.


Desmarchelier C, Martin JC, Planells R, Gastaldi M, Nowicki M, Goncalves A, Valero R, Lairon D, Borel P. 2014. The postprandial chylomicron triacylglycerol response to dietary fat in healthy male adults is significantly explained by a combination of single nucleotide polymorphisms in genes involved in triacylglycerol metabolism. J Clin Endocrinol Metab 99:E484ŌĆō488.


Fuchs SY, Dolan L, Davis RJ, Ronai Z. 1996. Phosphorylation-dependent targeting of c-Jun ubiquitination by Jun N-kinase. Oncogene 13:1531ŌĆō1535.

Gardina PJ, Clark TA, Shimada B, Staples MK, Yang Q, Veitch J, Schweitzer A, Awad T, Sugnet C, Dee S, Davies C, Williams A, Turpaz Y. 2006. Alternative splicing and differential gene expression in colon cancer detected by a whole genome exon array. BMC Genomics 7:325



Goodyear LJ, Kahn BB. 1998. Exercise, glucose transport, and insulin sensitivity. Annu Rev Med 49:235ŌĆō261.


Gordon GJ, Bueno R, Sugarbaker DJ. 2011. Genes associated with prognosis after surgery for malignant pleural mesothelioma promote tumor cell survival in vitro. BMC Cancer 11:169



Gu J, Orr N, Park SD, Katz LM, Sulimova G, MacHugh DE, Hill EW. 2009. A genome scan for positive selection in thoroughbred horses. PLoS One 4:6e5767



Huang X, Guo Y, Shui Y, Gao S, Yu H, Cheng H, Zhou R. 2005. Multiple alternative splicing and differential expression of dmrt1 during gonad transformation of the rice field eel. Biol Reprod 73:1017ŌĆō1024.


Livak KJ, Schmittgen TD. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2(ŌłÆDelta Delta C(T)) Method. Methods 25:402ŌĆō408.


Lynch KW. 2007. Regulation of alternative splicing by signal transduction pathways. Adv Exp Med Biol 623:161ŌĆō174.


Magnani M, Crinelli R, Bianchi M, Antonelli A. 2000. The ubiquitin-dependent proteolytic system and other potential targets for the modulation of nuclear factor-kB (NF-kB). Curr Drug Targets 1:387ŌĆō399.


Mancina RM, Burza MA, Maglio C, Pirazzi C, Sentinelli F, Incani M, Montalcini T, Pujia A, Congiu T, Loche S, Pilia S, Wiklund O, Boren J, Romeo S, Baroni MG. 2013. The COBLL1 C allele is associated with lower serum insulin levels and lower insulin resistance in overweight and obese children. Diabetes Metab Res Rev 29:413ŌĆō416.



Manning AK, Hivert MF, Scott RA, Grimsby JL, Bouatia-Naji N, Chen H, Rybin D, Liu CT, Bielak LF, Prokopenko I, et al. 2012. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat Genet 44:659ŌĆō669.



Nagase T, Ishikawa K, Suyama M, Kikuno R, Hirosawa M, Miyajima N, Tanaka A, Kotani H, Nomura N, Ohara O. 1999. Prediction of the coding sequences of unidentified human genes. XIII. The complete sequences of 100 new cDNA clones from brain which code for large proteins in vitro. DNA Res 6:63ŌĆō70.


Park KD, Park J, Ko J, Kim BC, Kim HS, Ahn K, Do KT, Choi H, Kim HM, Song S, Lee S, Jho S, Kong HS, Yang YM, Jhun BH, Kim C, Kim TH, Hwang S, Bhak J, Lee HK, Cho BW. 2012. Whole transcriptome analyses of six thoroughbred horses before and after exercise using RNA-Seq. BMC Genomics 13:473



Shin C, Manley JL. 2004. Cell signalling and the control of pre-mRNA splicing. Nat Rev Mol Cell Biol 5:727ŌĆō738.


Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. 2013. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725ŌĆō2729.








 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print





