 |
 |
|
|
Abstract
The complex analysis of the pedigree records of Czech Landrace (CLA), Czech Large White-dam line (CLWd), Czech Large White-sire line (CLWs), Duroc (DC), and Pietrain (PN) was performed to determine trends of genetic diversity (GD), and to find the main sources of the GD loss. The total size of the pedigree was 132,365, 391,151, 32,913, 13,299, and 7,160 animals in CLA, CLWd, CLWs, DC, and PN, respectively. Animals born in the years 2011 through 2013 were assumed as the reference population. The average pedigree completeness index for one generation back was 95.9%, 97.4%, 91.2%, 89.8%, and 94.2% for appropriate breeds. Number of ancestors explaining 100% of gene pool was 186, 373, 125, 157, and 37 in CLA, CLWd, CLWs, DC, and PN, respectively. The relative proportion of inbred animals (58%, 58%, 54%, 47%, and 25%), the average inbreeding (2.7%, 1.4%, 2.5%, 3.6%, and 1.3%) and the average co-ancestry (3.1%, 1.6%, 3.3%, 4.2%, and 3.3%) were found over the past decade in analysed breeds. The expected inbreeding under random mating increased during the last 10 years in CLWs and PN and varied from 1.27% to 3.2%. The effective population size computed on the basis of inbreeding was 76, 74, 50, 35, and 83 in 2012 in CLA, CLWd, CLWs, DC, and PN, respectively. The shortest generation interval (1.45) was observed for CLWd in sire to son selection pathway. The longest generation interval obtained PN (1.95) in sire to daughter pathway. The average relative GD loss within last generation interval was 7.05%, 4.70%, 9.81%, 7.47%, and 10.46%, respectively. The relative proportion of GD loss due to genetic drift on total GD loss was 85.04%, 84.51%, 89.46%, 86.19%, and 83.68% in CLA, CLWd, CLWs, DC, and PN, respectively. All breeds were characterized by a high proportion of inbred animals, but the average inbreeding was low. The most vulnerable breeds to loss of GD are DC and PN. Therefore, a breeding program should be more oriented to prevent the increase of GD loss in these breeds.
Numbers of pure-bred pigs were continuously reduced in the Czech Republic during the last years. This, together with the intensive selection in the economically oriented breeding schemes, can leads to the decreasing of genetic variability. Finally, it can cause in the higher risk of the inbreeding depression. As mentioned Koening and Simianer (2006), inbreeding can be evaluated on the basis of the pedigree records, where the inbreeding coefficient is defined as the probability that the two alleles of an individual are identical by descent. The expected heterozygosity, also called gene diversity, is the basic criterion for evaluating the genetic variability (Nei, 1973). Falconer and Mackay (1996) described it as a representation of the expected proportion of heterozygotes if the population is in Hardy-Weinberg equilibrium. The main manifestation of the inbreeding is the inbreeding depression, which is usually associated with negative influence on fitness-related traits (Fernández et al., 2005). Therefore, knowledge of the genetic diversity (GD) in the population under intensive selection is a basic premise to make a good decision to minimize the impact of inbreeding in the long-term. Fortunately, many studies have been performed in the evaluation of the livestock GD based on pedigree records (Norberg and Sørensen, 2007; Pjontek et al., 2012; Oravcová, 2013; Tang et al., 2013). This study represents the first comprehensive attempt to describe the trend of inbreeding, the risks associated with the increasing capabilities in main purebred pig population in the Czech Republic. The objective of the presented study was to determine the historical trend of inbreeding, to predict the value of expected inbreeding and to find the main causes of GD loss in pedigree information of the five pig breeds in Czech Republic.
Historical data were provided by the Pig Breeders Association of the Czech Republic. Pedigree information of the Czech Large White–dam line (CLWd), Czech Landrace (CLA), both dam breeds, Czech Large White–sire line (CLWs), Duroc (DC), and Pietrain (PT) breed containing 132,365, 391,151, 32,913, 13,299, and 7,160 records, respectively, were used in the analyses (Table 1). The latter three breeds are used in sire position in the Czech national pig breeding program. Pedigree records of the Hampshire breed were also analysed, however breeding program of this breed was stopped in 2012, and it is no longer used in Czech Republic. Animals born between 1980 and 2013 with known sex were only included into the analyses. The group of animals born within the years 2011 through 2013 was assumed as the reference population for analysed breeds. This population participated by 6.5, 5.5, 8.6, 12.6, and 9.4 per cent in CLWd, CLA, CLWs, DC, and PN breed, respectively.
Quality and integrity of the pedigree information were evaluated by four parameters: by maximum generations traced back, average complete equivalent generations, percentage of known ancestors and by pedigree completeness index (PCI).
Maximum generations traced back were assumed as the number of generations between an animal and its earliest ancestor. Average complete equivalent generations (i.e. number of generations in a comparable complete pedigree) were computed by Maignel et al. (1996) using the formula:
where N is number of animals in reference population, nj is the total number of ancestors of animal j in the population under study, and gij is the number of generations between an animal j and its ancestor i. Percentage of known ancestors was relative expression of all known ancestors per generation. Pedigree completeness index was computed following the MacCluer et al. (1983) algorithm. This procedure summarized the proportion of known ancestors in each ascending generation. It quantifies the change of detecting inbreeding in the pedigree (Sørensen et al., 2005). Followed equations were used:
and
where k represent maternal or sire line of an individual, ai is proportion of known ancestors in generation i and d is number of generations found. The values can vary between 0 and 1. If all ancestors are known Id = 1, if one of the parents is unknown Id = 0.
The generation interval (GI) was computed for four selection pathways (sire to son, sire to daughter, dam to son, dam to daughter) according to Falconer and Mackay (1996) definition. They defined the GI as an average age of the parents at the birth of their selected offspring. The average GI for whole population was also calculated by the same method.
The standard equation for estimation of the effective population size (Ne = 1/2ΔF) was used, where the ΔF was based on the average inbreeding coefficients of offspring and their direct parents (Falconer and Mackay, 1996).
Inbreeding coefficient of an individual was calculated by Meuwissen and Luo (1992) algorithm. Expected inbreeding coefficient, on the other side, was computed as the co-ancestry of the breeding animals assuming random mating (Falconer and Mackay, 1996). The co-ancestry coefficient was obtained by applying of Colleau (2002) algorithm. Total number of founders (ft) was defined as ancestors with unknown parents. Effective number of ancestors computed by Boichard et al. (1997) was represented as the minimum number of ancestors necessary to explain the complete GD of the population under study and was calculated as follows:
where pk is the genetic contribution of ancestor k, not yet explained by the n−1 already selected ancestors. Effective number of founders (founder equivalent), fe (Lacy, 1989), was defined as the number of equally contributing founders that would be expected to generate a similar amount of GD as in the studied population. It can be calculated from the genetic contribution of founders by the following formula:
where qi is the genetic contribution of ith founder to the reference population and ft is total number of founders. If the genetic contribution of all founders is equal, then ft = fe. Typically, the fe is smaller than ft mainly due to different selection intensity of appropriate pathways. As mentioned Bijma and Woolliams (1999), the fe is not useful parameter for measuring of GD. It is done by the convergence of genetic contributions of founders after some generations. Effective number of founder genomes (founder genome equivalent), fge, was defined as the number of equally contributing founder with no random loss of founder alleles that would give the same amount of GD as is presented in population under study. Value of fge was calculated by the algorithm of Caballero and Toro (2000) as follows:
where fg is the average co-ancestry coefficient of the group considered. Effective number of non-founders (non-founder equivalent), fne, was derived from the equation:
The loss of GD was derived from fe, fge, and fne. Total GD of the reference populations in each breed was calculated by Lacy (1995):
Genetic diversity loss in the population due to bottleneck and genetic drift since the founder generation is then expressed as 1−GD. It was assumed that the number of founders in the base population is large enough and GD in base population is close to 1, as recommended by Melka and Schenkel (2010). Amount of the GD in the reference population considered for loss of GD due to unequal number of founders (GD*) was obtained by Lacy (1995):
Similarly, loss of the GD due to unequal number of founders was expressed as 1−GD* (Caballero and Toro, 2000). The loss of GD due to the random genetic drift was derived from equation GD−GD* and was calculated as the inverse of 2fne (Caballero and Toro, 2000; Honda et al., 2004).
Four different software packages were used to compute parameters of the inbreeding and of GD. Generation intervals, PCI, and effective population size were computed in population report (POPREP) package (Groeneveld et al., 2009). Inbreeding coefficients, co-ancestry coefficients, the expected inbreeding under random mating and the deviation from random mating were calculated using Evolutionary Algorithm for Mate Selection (Berg, 2012). The contribution, inbreeding (F), coancestry (CFC) software package (Sargolzaei et al., 2006) was used to detect the effective number of founders, founder genomes and non-founders. Cumulated marginal contribution and number of ancestors explaining of appropriate proportion of gene pool were obtained using the pedigree analysis suited for large population (PEDIG) software (Boichard, 2002).
The maximum generations traced back varied from 20 (for PN) to 25 (for both dam breeds CLA and CLWd), whereas the CLWs and DC reached the same intermediate values, i.e. 23 generations (Table 1). The average equivalent complete generation was the lowest in PN (5.21) and DC (6.50) breeds. The highest values were obtained for CLWs, CLWd, and CLA ranged from 7.35 to 8.80 generations. Almost all of the animals in the analysed breeds have known ancestors in the first generation. Proportion of known ancestors gradually reduced in subsequent generations and reached the minimum value in the sixth generation. The lowest proportion of known ancestors in each generation was recorded in PN (11.5%) and DC (29.8%) breeds. The Figure 1a to 1e contains trends of PCI in all breeds. Value of PCI is usually a good indicator whether the pedigree data are good enough for the purposes of the following pedigree analysis. At least the last three generations, and the last two years amounted PCI of 100% in CLA, CLWd, and CLWS breeds. The PN breed has not reached 100% of PCI in any generation. Reduction of PCI in period 1990 through 1993 was observed as common feature in all of evaluated breeds, except of PN. The loss in PCI ranged from 4 to 32 percentage points (pp) in the first generation. We assumed that this reduction is a result of market opening after 1989 when intensive import of breeding animals into the Czech Republic started. These animals were assumed as unrelated, because their pedigree records were not known. That led to the temporary reduction in the completeness of pedigrees. Continuous increase in the completeness of pedigrees in the analysed breeds can be seen with the passage of about 1–2 GI given on Table 1.
Variability of the individual inbreeding parameters among the breeds is shown in Figure 2. Proportion of inbred animals reached in CLA, CLWd, CLWs, and DC was higher (57.5%, 57.9%, 54.1%, and 46.9%, respectively) compared to the value (25.1%) obtained in PN breed. Over all of analysed breeds, proportion of inbred animals in dam population was higher compared to the sires (+20 pp on average). Despite of the higher proportion of related animals in analysed breeds, value of the average inbreeding coefficient was low and ranged between 1.3% (PN) to 3.6% (DC). The DC breed also had the higher proportion of animals with inbreeding of F>0.6%. Over the analysed period and breeds, the average increase of inbreeding was low (from 0.17% in CLA to 0.56% in CLWs). The lowest (1.6%) and the highest (4.2%) value of the average co-ancestry were observed for CLWd and DC breed, respectively. However, the higher value of average rate of co-ancestry (0.56%) was found in CLWs. Reduction in average rate of co-ancestry achieved only in DC breed (−0.14%). Based on the average value of expected inbreeding, its value should be lower in CLWs and PN breed (1.27 and 3.2, respectively) in comparison to the observed value of inbreeding (1.4 and 3.6, respectively). Deviation from random mating was negative for all breeds except of CLWd (0.11%) and DC breed (0.28%). The negative values indicate avoidance of inbreeding.
All computed parameters from analysis of gene origin for the reference population are summarized in Table 3. The highest total number of founders was detected for CLWd (3,849), followed by CLA (1,776) and CLWs (1,575). On the other side, the lowest total number of founders achieved PN (714). The similar trend was also recorded for effective number of founders and number of founders equivalent, where the highest values reached CLWd (412 and 64). The lowest effective number of founders and founder genome equivalent was obtained in PN (182 and 30 founders, respectively). Values of fge/fe ratio were comparable with fe/ft ratio and ranged between 0.11 (CLW) to 0.35 (DC). Approximately 17 ancestors were needed to explain a fifty per cent of gene pool in the reference population of CLWd whereas in PN it was only four ancestors. The total number of ancestors needed for explaining of 100% of gene pool differs considerably between analysed breeds. As expected, the highest number was observed in CLWd (373 ancestors) and contrary, only 37 ancestors explain whole genetic pool in PN. The cumulated marginal contribution of 100 major ancestors is illustrated in Figure 3. Eighty nine ancestors explain ninety per cent of gene pool at least. Ten major ancestors explain about 46, 38, 47, 51, and 82 per cent of gene pool in CLA, CLWd, CLWs, DC, and PN, respectively.
Computed GI averaged over last decade and divided into four selection paths along with overall average GI in all breeds are shown in Table 4. The lowest value of average GI was observed for DC breed (1.59 year). In contrast, the longest GI obtained PN breed (1.86). The shortest GI was found for the sire to sire path, when comparing the different paths of selection, in CLA, CLWd, CLWs, and DC, respectively. As it was expected, the longest GI was detected for the dam to dam path. The highest difference among selection pathways within one breed was observed in PN (0.45). The most balanced GI was detected for CLWs (0.27) when comparing the individual paths of selection. Table 5 contains trend in the effective population size for all breeds. The values are presented only for the years with completed data. Value of Ne varied during last five years in all breeds. Drastically change among the years is usually caused by the low pedigree completeness. Therefore the lowest Ne was observed in DC breed (34 animals), whereas in CLWd and CLWs breeds obtained the highest Ne over the last five years.
The total amount of GD loss in all of analysed breeds during the last 28 years, due to different reasons, is shown in Figure 4a, 4b, and 4c. The continuous increase in the total loss of GD due to the unequal founder contribution and random genetic drift over the last decade has been observed only in CLWs and PN breeds. The average relative GD loss in reference population was 7.05%, 4.70%, 9.81%, 7.47%, and 10.46% for CLA, CLWd, CLWs, DC, and PN, respectively. The highest GD loss in reference population was found in CLWs and PN (12.21% and 13.11%, respectively), whereas the lowest values obtained CLWd (4.22%, respectively). In general, an increase in the total loss of GD can be seen in most breeds since 1996. Furthermore, the value of GD loss due to random genetic drift over the years increased (Figure 4b), whereas the value of the GD loss due to unequal founder contribution diminished (Figure 4c). The average proportion of the random genetic drift on the total GD loss ranged from 83.68 (PN) to 89.46 (CLWs).
Direct comparison of values between studies should always consider the overall complexity of analysed populations and datasets. The main goal of presented study was to analyse an actual status of GD in a population of five commercial pig breeds in the Czech Republic. An accuracy of estimated inbreeding coefficient is depending mainly on the pedigree information quality. Better knowledge of each individual’s ancestry leads to more accurate estimation of the inbreeding, and consequently, the other parameters such as the effective population size. Percentage of known ancestors (e.g. 61% over the analysed breeds in 4th generation) as well as the high quality of pedigree completeness parameters in analysed breeds (Table 1) indicated a precise estimation of other pedigree parameters. Furthermore, the pedigree quality of the Czech pig population is at an good level when compared our results with some recent pedigree analysis published in pigs. Tang et al. (2013) analyzed Chinese Duroc, Landrace and Yorkshire breeds farmed in Sichuan province. They observed the lower values of the PCI, which could be caused by the relative short time of the pedigree information (one decade). For example in the PCI for the third generation it was below 70% for all breeds expect Yorkshire. The fifth generation not reached the 50%. Analyses of South African Landrace and Duroc presented by Groeneveld et al. (2009) showed the relative higher PCI, which exceed 90% in the sixth generation, but these values were calculated for animals born in the last year (2008). Similarly, the Finish Landrace and Yorkshire (Uimari and Tapio, 2011) reached a slightly higher pedigree completeness indices (97% and 99%, respectively). Compared to our study, Melka and Schenkel (2010) published a similar complete pedigree records for the Canadian pig breeds.
As we expected, there were found the breed differences in the quality of pedigree parameters in our study. Disparities between breeds, but also between studies are, in our opinion, caused by the dynamics and intensity of individual breeds used in national or commercial breeding programs. Moreover, some differences could be also based on using of artificial insemination, on the impact of imports or on the depth of pedigree knowledge of imported animals. These parameters finally influenced the quality of pedigree and consequently the estimated inbreeding and GD.
Observed values of actual inbreeding coefficients and actual co-ancestry coefficients not exceed 3.6% and 4.2%, respectively over the analysed breeds. Melka and Schenkel (2010) found higher inbreeding coefficients for Canadian Hampshire and Lacombe breeds (approximately 18% and 12%, respectively in the most of evaluated year), while for Canadian Duroc and Landrace breeds it has not exceed 6%. Similarly to results of Welsh et al. (2010), which analyzed five United States pig breeds, we found a higher proportion of inbred animals for all breeds, expect of Pietrain, although the values in the above mentioned study were extremely high (more than 99% of animals were inbred). More than 90% of all inbred animals have inbreeding less than 5% in our study, expect of Duroc, whereas the previously mentioned authors reported proportion 90% of inbred animals with inbreeding varied from 10% (for Hampsihe) to 20% (for Berkshire).
One of the main parameters for monitoring of GD is rate of inbreeding. According to the recommendations of the Food and Agriculture Organization of the United Nations (FAO, 2000), ΔF should not exceed 1%, or had to be less than 0.5% (Nicholas, 1989). In our study, only CLWs and DC breeds have not met the above mentioned thresholds. Generation interval, observed in our study, was larger for female’s pathways, than for male’s pathways in all breeds, expect the PN. Higher difference in GI of selection pathways in PN breed was probably caused by low number of evaluated animals in this breed (given in Table 1).
FAO (2000), cited by Welsh et al. (2010), suggested that the effective population size (Ne) for a breed should be maintained above 50 animals. Meuwissen and Woolliams (1994) recommended maintain Ne value between 31 and 250 animals, for good population fitness. Melka and Schenkel (2010), cited by Tang et al. (2013), reported an effective population size of 50 as a threshold necessary to avoid the negative impact of inbreeding in population. These authors defined at least 500 animals needed for maintainence of GD in population for several generations. From the perspective of current and past effective population size, all breeds analysed in our study required an additional attention, despite the fact that, almost all of them meet the minimum requirements for Ne. The overall trend in Ne is more negative than positive, especially over the last three years. Intensive reduction in the number of animals in the analysed breed populations has been observed in this period in the Czech Republic. Higher value of Ne for PN (83 animals calculated in 2012) was probably caused by the applied methodology, where Ne was based on ΔF. Over the analysed breeds, PN breed generally reached the lowest values of average inbreeding and also values for ΔF. Average number of known ancestors and values of PCI were also the lowest in PN breed (Table 1 and Figure 1e). These facts could finally affect the values of Ne observed for PN breed. Uimari and Tapio (2011) not confirmed this presumption in their study. Mentioned authors investigated changes in Ne considering different number of pedigree generations (5 and 10, respectively), different limits for PCI (0.4 and 0.7, respectively) and different size of reference populations (3 and 5 years, respectively) for Finish Landrace and Yorkshire breeds. Generally, the estimates were not sensitive to a reduced reference population and to decreased PCI value. Only small changes were observed when 5 ancestral generations were used instead of 10 generations in their study.
Parameters derived from probability of gene origin analysis point to increased risk of GD loss within actual breeding program in breeds analysed in our study. A relative small number of major ancestors are needed to explain a whole GD, expect of CLWd breed. As mentioned Melka and Schenkel (2010), the ratio between effective number of founders and total number of founders shows on reduction of GD based on unequal contribution of founders. The smaller number of effective founders than total number of founders indicates the excessive use of some animals as parents or a disequilibrium between the founder contributions. The fge/fe ratio can be used to quantify of only genetic drift influence on amount of GD, excluding the effect of founder contribution. Both of the previous mentioned ratios are inversely proportional. Therefore, the effect of random genetic drift is stronger with lower fge/fe ratio. In our study, the impact of random genetic drift was substantial for all breeds. The highest value of overall GD lost was observed for PN and CLWs breeds, within last decade. Especially, the continuous increasing of GD loss was observed from 2007 for both breeds. Annual increase in the loss of GD ranged from 0.11% to 3.6% in this period. The loss of GD was increased by 1.5% and 1.3% on average for CLWs and PN, respectively. Melka and Schenkel (2010) reached similar results in their analysis. They found the reduction of GD in four Canadian pig breeds especially in the last two decades of the analysed period. On the other hand, Tang et al. (2013) found a gradual reduction of the GD loss for the three Chinese breeds in the last 3 to 4 years of evaluated. Increase in the total number of animals in this period, and mainly increased import unrelated animals to the populations under studied were the main reasons for the positive trend in the GD loss according to these authors. Opposite situation occurred in the Czech pig population, where the number of animals along with the proportion of imported animals reduced in last period. In totally, GD loss was caused mainly by random genetic drift as this parameter participated by 84% to 90% on the total loss of GD in our study (Figure 4). These findings are in agreement with results of Melka and Schenkel (2010) for Canadian Duroc and Lacombe breeds. The mentioned authors observed the opposite trends for other two pig breeds, the Canadian Hampshire and Landrace, as well as Tang et al. (2013) for all three Chinese pig breeds. One of the main reasons for such a high proportion of the loss of GD caused by random genetic drift (and other reasons like bottleneck) could be very low effective population sizes for all Czech pig breeds. Our results represent the historical and the current situation in genetic variability of pig breeds in Czech Republic. The possible impact on the genetic improvement should be investigated. Independently observed genetic trends for evaluated traits of pig breeds in Czech Republic show their permanent and regular growth (unpublished data). Despite the fact that the loss of genetic variability has been an increasing in recent years, it is not reflected on genetic trends of production and reproduction traits. We assume that this situation could occur with a certain time lag due to GI. It will be necessary to specify whether it is under the influence of the GD loss, or due to other factors (e.g. achieving of the certain genetic gain limits for evaluated traits in the given environment).
The first complex pedigree analysis of five Czech pig breeds was performed. All breed were characterized by sufficient quality of pedigrees, which provides adequately accurate estimate of the parameters needed for measuring of the GD. Presented results demonstrated that the intensive selection and actual breeding programs influenced the loss of GD. Higher proportion of inbred animals and smaller number of founder genomes equivalent were observed for all breeds, expect of PN. High proportion of GD loss due to the random genetic drift was observed. The most vulnerable breed with respect to the loss of GD was PN, CLWs, and DC breed. Therefore, the breeding focused to reduce proportion of inbred animals as well as to increase an effective population size will be important in these breeds. Moreover, an introduction of new genetic material or occasional immigrants into a managed population could be helpful to counter the effect of genetic drift. In the future, the continuous monitoring of the population will be also necessary to secure the GD over the long-term. A possible breeding strategy, especially for CLWs and DC, should incorporate the rules that control the inbreeding and limit the impact of individual dam or sire by limiting number of used progeny in next generation.
ACKNOWLEDGMENTS
Thanks are due to L. Vostrý and R. Prošková for helpful discussions and technical assistance and also to the Pig Breeders Association of the Czech Republic for making the data available. This study was supported by the project QJ1310109 of the Ministry of Agriculture of the Czech Republic.
Figure 1
i) Pedigree completeness index in (a) Czech Landrace (CLA), (b) Czech Large White dam line (CLWd) (c) Czech Large White sire line (CLWs), (d) Duroc (DC, and (e) Pietrain (PN) breed.
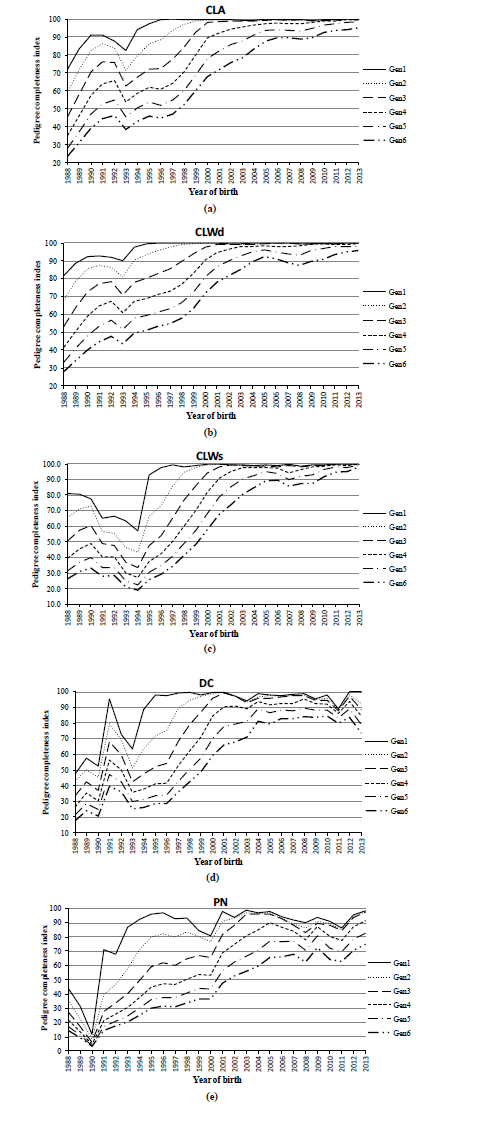
Figure 2
Proportion of inbreeding in evaluated breeds. CLA, Czech Landrace; CLWd, Czech Large White dam line; CLWs, Czech Large White sire line; DC, Duroc; PN, Pietrain,
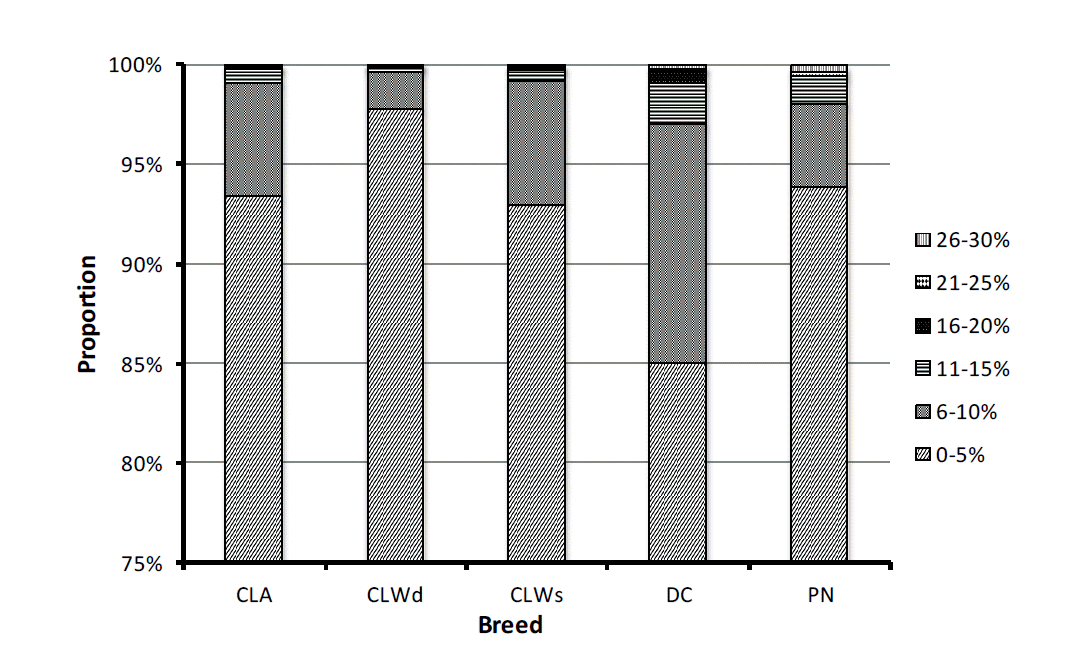
Figure 3
Cumulative marginal contribution of 100 ansectors in evaluated breeds. CLA, Czech Landrace; CLWd, Czech Large White dam line; CLWs, Czech Large White sire line; DC, Duroc; PN, Pietrain.

Figure 4
The genetic diversity loss due to (a) unequal founder contribution and random genetic drift (1 − GD), (b) random genetic drift (GD−GD*), and (c) unequal founder contribution (1−GD*) in evaluated breeds. CLA, Czech Landrace; CLWd, Czech Large White dam line; CLWs, Czech Large White sire line; DC, Duroc; PN, Pietrain.

Table 1
Basic characteristic of the pedigree datasets and the parameters of pedigree completeness in the analysed breeds
| Breed1 | CLA | CLWd | CLWs | DC | PN |
|---|---|---|---|---|---|
| Number of animals in pedigree | 132,365 | 391,151 | 32,913 | 13,299 | 7,160 |
| Number and percentage2 of animals in reference population | 8,630 (6.5) | 21,370 (5.5) | 2,840 (8.6) | 1,679 (12.6) | 671 (9.4) |
| Number and percentage2 of animals born in last decade | 44,336 (33.5) | 127,744 (32.7) | 13,750 (41.8) | 5,748 (43.2) | 3,435 (48.0) |
| Total number of herds/number of breeding herds in 2013 | 9/4 | 20/6 | 6/5 | 4/4 | 4/4 |
| Proportion of animals in breeding herds in 2013 (%) | 72.9 | 67.8 | 96.7 | 100.0 | 100.0 |
| Proportion of imported parents from all known parents in 2013 (%) | 1.13 | 0.14 | 1.76 | 3.75 | 4.88 |
| Maximum generation traced | 25 | 25 | 23 | 23 | 20 |
| Average equivalent complete generation | 7.35 | 7.37 | 8.80 | 6.50 | 5.21 |
| Percentage of known ancestors in | |||||
| 1st generation | 100.0 | 100.0 | 99.7 | 99.8 | 99.3 |
| 2nd generation | 94.1 | 97.2 | 92.3 | 90.2 | 89.1 |
| 4th generation | 66.3 | 70.9 | 66.7 | 58.0 | 44.4 |
| 6th generation | 33.6 | 38.7 | 35.3 | 29.8 | 11.5 |
Table 2
Parameters of inbreeding analysis in all animals in percent (%)
Table 3
Parameters of gene origin for the reference population in evaluated breeds
Table 4
Average generation intervals in evaluated breeds
| Breed | CLA | CLWd | CLWs | DC | PN |
|---|---|---|---|---|---|
| Average generation interval | 1.77 | 1.76 | 1.71 | 1.59 | 1.86 |
| GI for s-s1 path | 1.54 | 1.45 | 1.54 | 1.40 | 1.73 |
| GI for s-d2 path | 1.72 | 1.70 | 1.66 | 1.57 | 1.95 |
| GI for d-s3 path | 1.68 | 1.72 | 1.72 | 1.55 | 1.50 |
| GI for d-d4 path | 1.86 | 1.89 | 1.81 | 1.69 | 1.84 |
REFERENCES
Berg P. 2012. EVA version 1.75. Evolutionary algorithm for mate selection. User’s Guide. Institute of Arhus, Institute of Genetic and Biotechnology; Tjele, Denmark:
Bijma P, Woolliams JA. 1999. Prediction of genetic contributions and generation intervals in populations with overlapping generations under selection. Genetics 151:1197–1210.




Boichard D, Maignel L, Verrier E. 1997. The value of using probabilities of gene origin to measure genetic variability in a population. Genet Sel Evol 29:5–23.


Boichard D. 2002. PEDIG: a fortran package for pedigree analysis suited for large populations. In : Proc. of the 7th World Congress on Genetics Applied to Livestock Production (WCGALP); INRA; Castanet-Tolosan, France [CD-Rom]: p. 19–23.
Caballero A, Toro MA. 2000. Interrelations between effective population size and other tools for management of conserved populations. Genet Res 75:331–343.


Colleau JJ. 2002. An indirect approach to the extensive calculation of relationship coefficients. Genet Sel Evol 34:409–421.



Falconer DS, Mackay TFC. 1996. Introduction to Quantitative Genetics. 4th EdLongman Scientific and Technical; Harlow, UK:
FAO. 2000. Secondary guidelines for development of farm animal genetic resources management plans. Management of small populations at risk. FAO; Rome, Italy:
Fernandez J, Villanueva B, Pong-Wong R, Toro MA. 2005. Efficiency of the use of pedigree and molecular marker information in conservation programs. Genetics 170:1313–1321.



Groeneveld E, Van der Westhuizen B, Maiwashe A, Voordewind F, Ferraz JBS. 2009. POPREP: A generic report for population management. Genet Mol Res 8:1158–1178.


Honda T, Nomura T, Yamaguchi Y, Mukai F. 2004. Monitoring of genetic diversity in the Japanese Black cattle population by the use of pedigree information. J Anim Breed Genet 121:242–252.

Koenig S, Simianer H. 2006. Approaches to the management of inbreeding and relationship in the German Holstein dairy cattle population. Livest Sci 103:40–53.

Lacy RC. 1995. Classification of genetic terms and their use in the management of captive populations. Zoo Biol 14:565–577.

MacCluer JW, Boyce AJ, Dyke B, Weitkamp LR, Pfennig DW, Parsons CJ. 1983. Inbreeding and pedigree structure in Standardbred horses. J Hered 74:394–399.

Maignel L, Boichard D, Verrier E. 1996. Genetic variability of French dairy breeds estimated from pedigree information. Interbull Bulletin 14:49–54.
Melka MG, Schenkel F. 2010. Analysis of genetic diversity in four Canadian swine breeds using pedigree data. Can J Anim Sci 90:331–340.

Meuwissen THE, Luo Z. 1992. Computing in breeding coefficients in large populations. Genet Sel Evol 24:305–313.


Meuwissen THE, Woolliams JA. 1994. Effective sizes of livestock populations to prevent a decline in fitness. Theor Appl Genet 89:1019–1026.


Nicholas FW. 1989. Incorporation of new reproductive technology in genetic improvement programmes. Evolution and Animal Breeding. Hill WG, Mackay TFC, editorsCAB Int.; Wallingford, UK: p. 201–209.
Norberg E, Sørensen AC. 2007. Inbreeding trend and inbreeding depression in the Danish populations of Texel, Shropshire, and Oxford Down. J Anim Sci 85:299–304.


Oravcová M. 2013. Pedigree analysis in White Shorthaired goat: First results. Arch Tierz 56:547–554.

Pjontek J, Kadlečík O, Kasarda R, Horný M. 2012. Pedigree analysis in four Slovak endangered horse breeds. Czech J Anim Sci 57:54–64.

Sargolzaei M, Iwaisaki H, Colleau JJ. 2006. CFC: A tool for monitoring genetic diversity. In : Proc. of the 8th World Congress on Genetics Applied to Livestock Production (WCGALP); August 13–18 2006; Belo Horizonte, Brazil. p. 27–28.
Sørensen AC, Sørensen MK, Berg P. 2005. Inbreeding in Danish dairy cattle breeds. J Dairy Sci 88:1865–1872.


Tang GQ, Xue J, Lian MJ, Yang RF, Liu TF, Zeng ZY, Jiang AA, Jiang YZ, Zhu L, Bai L, Wang Z, Li XW. 2013. Inbreeding and genetic diversity in three imported swine breeds in china using pedigree data. Asian Australas J Anim Sci 26:755–765.



- TOOLS
-
METRICS

- Related articles
-
Evaluation of the genetic diversity of six Chinese indigenous chickens2020 October;33(10)
Estimation of Genetic Parameters of Body Weight Traits in Ghezel Sheep2010 February;23(2)
Evaluation of Genetic Variability in Kenkatha Cattle by Microsatellite Markers2006 December;19(12)
Estimation of Additive and Dominance Genetic Variances in Line Breeding Swine2001 January;14(1)
Reevaluation of the Metabolic Essentiality of the Vitamins - Review -2000 January;13(1)





 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Print
Print




