Predicting the rate of inbreeding in populations undergoing four-path selection on genomically enhanced breeding values
Article information
Abstract
Objective
A formula is needed that is practical for current livestock breeding methods and that predicts the approximate rate of inbreeding (ΔF) in populations where selection is performed according to four-path programs (sires to breed sons, sires to breed daughters, dams to breed sons, and dams to breed daughters). The formula widely used to predict inbreeding neglects selection, we need to develop a new formula that can be applied with or without selection.
Methods
The core of the prediction is to incorporate the long-tern genetic influence of the selected parents in four-selection paths executed as sires to breed sons, sires to breed daughters, dams to breed sons, and dams to breed daughters. The rate of inbreeding was computed as the magnitude that is proportional to the sum of squared long-term genetic contributions of the parents of four-selection paths to the selected offspring.
Results
We developed a formula to predict the rate of inbreeding in populations undergoing four-path selection on genomically enhanced breeding values and with discrete generations. The new formula can be applied with or without selection. Neglecting the effects of selection led to underestimation of the rate of inbreeding by 40% to 45%.
Conclusion
The formula we developed here would be highly useful as a practical method for predicting the approximate rate of inbreeding (ΔF) in populations where selection is performed according to four-path programs.
INTRODUCTION
Deterministic predictions of response to multi-trait genomic selection in a single generation in a population with four-path programs, was developed [1,2]. That is, the selection paths in four-path programs are sires to breed sires (SS), sires to breed dams (SD), dams to breed sires (DS), and dams to breed dams (DD). However, when creating formulas for calculating the asymptotic response to index or single-trait selection in four-path selection programs rather than in a single generation, the initial genetic response in generation 0 overestimated the asymptotic response due to the decrease in equilibrium genetic variance from generation 0 onwards [3]. Consequently, to safeguard the genetic variation of the population over the long term, the rate of inbreeding needs to be restricted to an acceptable level. Therefore, one needs to know the expected rate of inbreeding as well as the equilibrium genetic response before choosing a breeding scheme.
A population with discrete generations under mass selection in a four-path selection program is modeled to predict the rate of inbreeding in the long term. When sires in the SS path are used with constant selection intensity and in equal number throughout the usage period of several years, every SS sire belongs to a single or exclusive category. Similarly, SD, DS, and DD parents each belong to a single or exclusive category when they are used with constant selection intensity and in equal numbers over several years. Consequently generations can be regarded as discrete rather than overlapping. A formula is needed that is practical for current livestock breeding methods and that predicts the approximate rate of inbreeding (ΔF) in populations where selection is performed according to four-path programs.
The rate of inbreeding is proportional to the sum of squared long-term genetic contributions [4]. General predictions of expected genetic contributions were developed by Woolliams et al [5] by using equilibrium genetic variances instead of second-generation genetic variances. Methods were developed by Bijma and Woolliams [6] to predict rates of inbreeding in populations selected on breeding values according to best linear unbiased prediction (BLUP) [7]. A formula was developed for predicting the rate of inbreeding in four-selection path programs [8]; however, this formula ignored the effect of selection. The purpose of the current study was to develop a formula for predicting the rate of inbreeding in four-path selection programs that incorporated the effect of selection and was practical for use under real-life conditions of cattle breeding.
MATERIALS AND METHODS
Prediction of expected long-term genetic contributions
Our prediction method is based on the concept of long-term genetic contributions. The long-term genetic contribution of individual i (ri) in generation t1 is defined as the proportion of genes from individual i that are present in individuals in generation t2 deriving by descent from individual i, where (t2–t1) →∞ [5]. That is, after several generations, the genetic contributions of ancestors stabilize and become equal for all descendants, i.e., the ultimate proportional contribution of an ancestor to its descendants is reached.
Selection is performed in four categories of selection path (SS, SD, DS, and DD). Rates of inbreeding can be expressed in terms of the expected contributions of these categories [6, 9–11]:
where 1′ = (1 1 1 1), N is a 4×4 diagonal matrix containing the number of selected parents for element (i, i) as Ni,i, N1,1 is the number of sires in SS and is referred to as NSS, N2,2 is the number of sires in SD and is referred to as NSD, N3,3 is the number of dams in DS and is referred to as NDS, and N4,4 is the number of dams in DD and is referred to as NDD. In addition,
The selective advantage of the ith sire in SS (Si,SS) and in SD (Si,SD) in the linear model is:
where Ai,SS is the breeding value of sire i in SS or SD, Āi,DS and DD is the average breeding value of dams mated to the ith sire in SS and SD, respectively; the dams mated to the ith sire in SS belong to the DS category, and the dams mated to the ith sire in SD belong to the DD category; and Āi,SS, Āi,SD, Āi,DS, and Āi,DD, are the average breeding values of the individuals in the SS, SD, DS, and DD categories.
The selective advantage of the ith dam in DS (Si,DS) and in DD (Si,DD) in the linear model is:
where Ai,DS and DD is the breeding value of dam i in DS and DD, respectively; Ai,SS and SD is the breeding value of a sire mated to the ith dam in DS and DD, respectively; the sires mated to the ith dam in DS belong to the SS category; and the sires mated to the ith dam in DD belong to the SD category.
Expected contributions (ui,SS,SD,DS,or DD) are predicted by linear regression on the selective advantage. That is,
where αx is the expected contribution of an average parent in x, and βx is the regression coefficient of the contribution of i on its selective advantage (Si,x). In addition, αx can be obtained according to Woolliams et al [9]:
where G is a 4×4 matrix representing the parental origin of genes of selected offspring in the order of SS, SD, DS, and DD category, i.e., representing rows offspring and columns parental categories. That is,
However,
where α′N is the left eigenvector of G with eigenvalue 1; the left eigenvector is obtained according to Bijma and Woolliams [11] and is equal to (0.25 0.25 0.25 0.25).
Solutions for βx are obtained according to Woolliams et al [9]:
note that the right hand side of (1) is unaffected by the number of parents, so that βx is inversely proportional to the number of parents (that is,
In addition, Λ is a 4×4 matrix of regression coefficients, with λxy being the regression coefficient of the number of selected offspring of category x on Sj,y of its parent j of category y. In the same way as Π, we have non-zero elements, λSS,SS and λSD,SS, λDS,SD and λDD,SD, λSS,DS and λSD,DS, and λDS,DD and λDD,DD in Λ as elements (1,1) and (2,1), (3,2) and (4,2), (1,3) and (2,3), and (3,4) and (4,4), respectively. Consequently,
representing rows as offspring and columns as parental categories.
In our current study, elements in matrices Π and λ were calculated from Woolliams et al [9] and Bijma and Woolliams [11], as outlined in Appendices A and B.
The sires in the SS category are included among the sires in SD category. That is, the sires in the SS category are selected not only to breed sons but as sires in the SD category to breed daughters. Similarly the dams in the DS category are included among the dams in the DD category. The dams in the DS category are selected not only to breed sons but as dams in the DD category to breed daughters. Therefore, after applying the procedure of Bijma and Woolliams [6], the number of sires in SD is larger than that of sires in SS, and the number of dams in DD is larger than that of dams in DS. Therefore,
E denotes the expectation with respect to the selective advantage,
and E(ĀDS – ĀDD) = (iDS – iDD)σA,f,
note that variance of selective advantage (
The accounting percentage derived from SS, SD, DS, and DD for the rate of inbreeding (ΔF) is obtained,
When the effect of selection on inbreeding is ignored, i.e., β = 0,
This result is in agreement with the formula from Gowe et al [8], which likewise neglects the effects of selection on ΔF.
Correction of E (ΔF) from Poisson variances
The correction for deviations of the variance of the family size from independent Poisson variances in the selected offspring from SS, SD, DS, and DD parents, i.e., δSS, δSD, δDS, and δDD, can be approximated by Woolliams and Bijma [10].
According to Woolliams and Bijma [10],
Where
ΔVSS, ΔVSD, ΔVDS, and ΔVDD, are 4×4 matrices which are variances of selected family size deviated from Poisson variance by applying binomial distribution to the family size from the parents of SS, SD, DS, and DD, respectively, and sx is the selective advantage of parents in category x (SS, SD, DS, DD). Elements of ΔVSS,SD,DS,or DD are shown in Appendix C.
Example applications of the formula
To demonstrate the application of our formula, we assumed only two quantitative traits: trait 1 was assumed to be moderately heritable, with h2 = 0.3, whereas trait 2 was assumed to have low heritability, with h2 = 0.1. These traits are selected as single traits expressed as GEBV. Furthermore, we assumed an aggregate genotype as a linear combination of genetic values, each weighted by the relative economic weights, which was expressed as a1g1 + a2g2, where g1 is the true genetic value for trait i, ai is the relative economic weight for trait i, and the genetic correlation between traits 1 and 2 was assumed as 0.4. Index selection was performed to select a1g1 + a2g2, that is, breeding goal (H), under the assumption that the relative economic weight between traits 1 and 2 is 1:1. Breeding value (A) was defined as described earlier in the Methods; for example, the breeding value of sire i in SS was defined as Ai,SS. Similarly, the breeding goal value (H) of sire i in SS can be expressed as Hi,SS; note that the formula that we developed in Methods can be applied not only to breeding value (A) but also to breeding goal value (H).
In our example, we assumed the reliabilities of the GEBVs for traits 1 and 2 to be 0.5721 and 0.4836, respectively [3]. Index selection (I) was performed as I = a1 GEBV1 + a1 GEBV2, because GEBVs are assumed to be derived from multiple-trait BLUP (MT BLUP) genetic evaluation methods in the current study (as done for single-step genomic BLUP [13]). We calculated equilibrium genetic variances and reliabilities based on Togashi et al [3]. The initial (generation 0) and equilibrium genetic variances and reliabilities for single-trait selection (h2 = 0.3 or h2 = 0.1) and index selection are shown in Table 1. Rates of inbreeding were calculated based on equilibrium genetic variances and reliabilities, because regression coefficients of the number or breeding value of selected offspring on the breeding value of the parent are equal for the parental and offspring generations under equilibrium genetic variances and reliabilities.
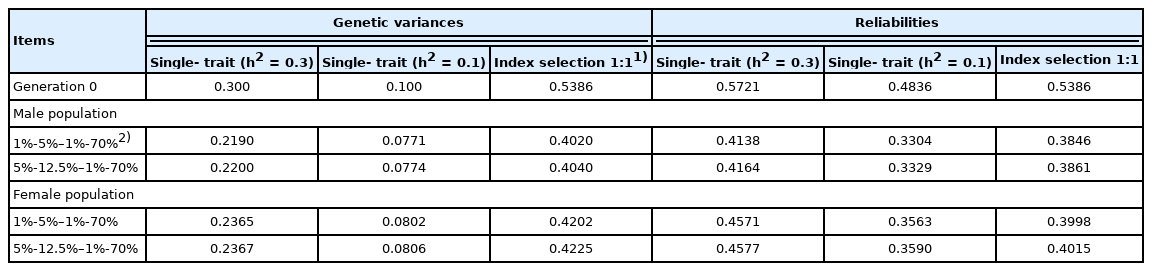
Genetic variances and reliabilities of genomically enhanced breeding values in generation 0 and at the asymptote
We considered two scenarios for the selection percentages for SS, SD, DS, and DD—5%-12.5%–1%-70% and 1%-5%–1%-70%—and three scenarios for the numbers of selected parents of SS, SD, DS, and DD—namely 20-50–100-7,000, 40-100–200-14,000, and 60-150–300-21,000. Therefore, we considered six scenarios (two scenarios of selection percentage and three scenarios of the number of parents in SS, SD, DS, DD) in total. Note that the two scenarios for selection percentage for SS, SD, DS, and DD differ only in the selection percentage along the SS and SD selection paths, because under actual breeding conditions, selection intensity can be adjusted more easily in male selection paths (SS and SD) than in female selection paths (DS and DD). The numbers of male and female offspring from a dam of DS, i.e., fmds and ffds, were set at 4. The number of female offspring from a dam of DD, i.e., ffdd, was set at 1.4. These numbers are derived from the years of usage of a dam and the reproduction method (ovum collection, in vitro fertilization, or embryo transfer). When DS and DD parents are used with constant selection intensity and in equal numbers over several years, they belong to a single or exclusive category. The numbers are used to compute the deviation of the variance of the family size from Poisson variance.
RESULTS AND DISCUSSION
Rates of inbreeding
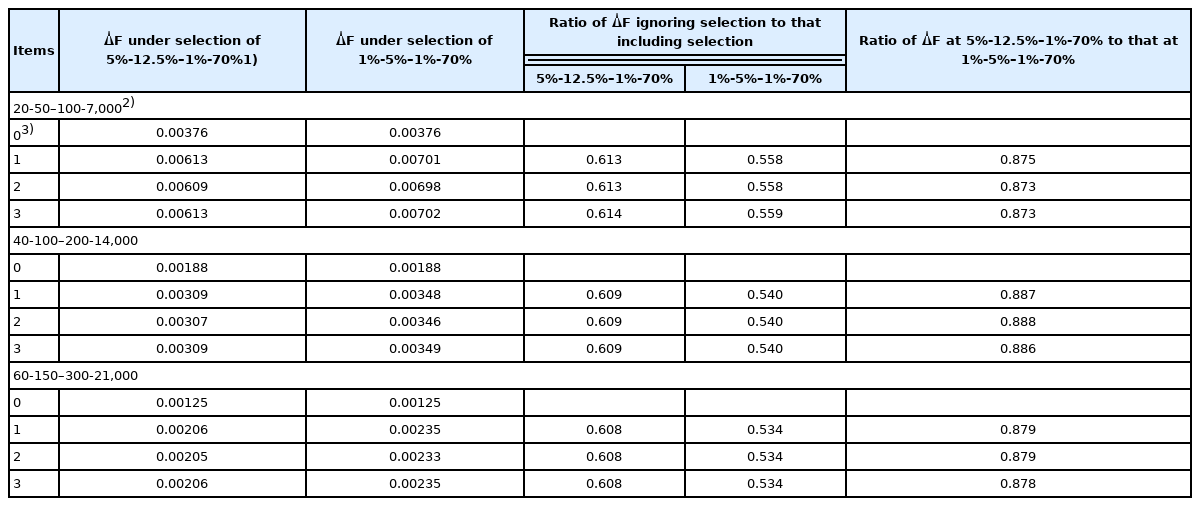
The rates of inbreeding without correction for deviation from Poisson variances (that is, the rates of inbreeding with Poisson family size) are shown in Table 2. Because the rates from Gowe et al [8] do not account for selection, ΔF is the same between two selection percentages in SS, SD, DS, and DD, i.e., 1%-5%–1%-70% and 5%-12.5%–1%-70%. In contrast, ΔF derived from the method developed in the current study increased with the increase in selection intensity. When we applied our formula, ΔF was lower when selection was ignored than when it was included, suggesting that ΔF was underestimated when selection was ignored. The ratio of ΔF when selection was ignored to that when it was included was about 0.61 under the selection percentages of 5%-12.5%–1%-70% for the SS, SD, DS, and DD selection paths, whereas the ΔF ratio was 0.53 to 0.56 under the selection percentage condition of 1%-5%–1%-70%. That is, calculation according to Gowe et al [8] underestimated ΔF by approximately 40% and 45% under selection percentages of 5%-12.5%–1%-70% and 1%-5%–1%-70% for the SS, SD, DS, and DD selection paths, respectively. In contrast, the rates of inbreeding under selection estimated by using our formula were 63% to 87% greater than those calculated according to the current working formula, which does not consider selection [8]. The ratio of ΔF for 5%-12.5%–1%-70% to that for 1%-5%–1%-70% was 0.88 to 0.89, resulting in an approximately 12% decrease in ΔF due to increasing the selection percentage or decreasing the selection intensity for SS and SD for all three scenarios compared in the numbers of parents in SS, SD, DS, and DD (20-50–100-7,000, 40-100–200-14,000, and 60-150–300-21,000). In contrast, the decrease in ΔF due to the increase in the number of parents was proportional to the numbers. The ΔF under the number of parents in SS, SD, DS, and DD (40-100–200-14,000 and 60-150–300-21,000) was approximately half and one third of the ΔF under the number of parents (20-50–100-7,000), respectively, for all two scenarios compared in the selection percentage of parents in SS, SD, DS, and DD (5%-12.5%–1%-70% and 1%-5%–1%-70%). Consequently, the decrease in the rate of inbreeding likely would be greater with an increase in the number of parents than with a decrease in selection intensity; however, we need to perform more trials at different selection intensities to confirm this association.
In general, both genetic gain and ΔF increase with an in crease in selection intensity. However, because the number of parents has a greater effect on inbreeding than does selection intensity, increasing the number of parents is one option for offsetting the increase in ΔF due to an increase in selection intensity.
The rate of inbreeding was slightly lower in single-trait se lection with a low heritable trait (h2 = 0.1) than the other selection methods (i.e., single-trait selection with a trait (h2 = 0.3) and index selection [Table 2]). However, the difference was not so remarkable. Consequently, we consider the major factors in the rate of inbreeding to be the number of parents and the selection intensity in each of the four selection paths.
Values for effective population size expressed as
The expectation of the square of long-term contribution of an individual (that is,
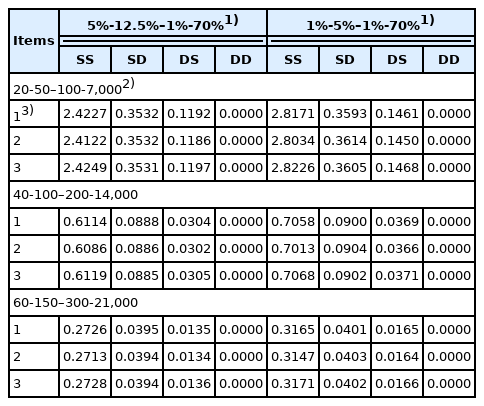
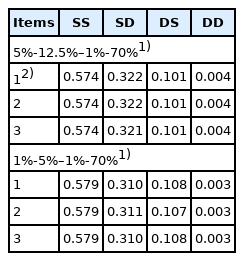
The accounting percentage derived from SS, SD, DS, and DD for the rate of inbreeding (ΔF) when the numbers of parents in SS, SD, DS, and DD are 40-100–200-14,000 is shown in Table 5. The accounting percentage in SS was the greatest of all the four selection paths for all two scenarios compared in the selection percentage in SS, SD, DS, and DD (that is, 5%-12.5%–1%-70% and 1%-5%–1%-70%), because the expectation of the square of lifetime long-term contribution of an individual was the greatest in SS of all the four selection paths (Table 4). The sum of accounting percentage in SS and SD was approximately 90% for ΔF, because the number of male parents in SS and SD was smaller than that of female parents in DS and DD and selection intensity in male parents is generally higher than that in female parents. In addition, the accounting percentage in each of the four selection paths when the numbers of parents in SS, SD, DS, and DD were 40-100–200-14,000 (Table 5) was approximately the same as the other scenario when the numbers of parents in SS, SD, DS, and DD were 20-50–100-7,000 or 60-150–300-21,000, although the accounting percentage in SS, SD, DS, and DD in the other scenarios was not shown. This is mainly because the expected contribution of an average parent (α) and the regression coefficient of the contribution of an individual on its selective advantage (β) are inversely proportional to the number of parents as explained previously in equation (1). Consequently, the accounting percentage derived from SS, SD, DS, and DD for the rate of inbreeding (ΔF), (that is, the relative magnitude of ΔF in SS, SD, DS, and DD), resulted in almost the same for all three scenarios compared in the numbers of parents in SS, SD, DS, and DD, even if the absolute magnitude of ΔF derived from each of the four selection paths differed in the number of parents in each of the four selection paths.
Correction derived from deviation from Poisson variance
Corrections for deviations in the variance of the family size from independent Poisson variances (×10−4) approximated by binomial distribution are shown in Table 6. The magnitude approximated by binomial distribution under the assumed selection percentages in the SS, SD, DS, and DD selection paths of 5%-12.5%–1%-70% and 1%-5%–1%-70% varied from −0.29×10−4 to −0.88×10−4, and −0.04×10−4 to −0.12×10−4, respectively. In comparison, the rates of inbreeding with Poisson family size without correction shown in Table 2 varied from 0.2×10−2 to 0.7×10−2. Therefore, because the magnitude of correction was much smaller than that of the rates of inbreeding with Poisson family size without correction, correction is unnecessary; thus the rates of inbreeding without correction (Table 2) are reasonable rates of inbreeding. However, the method in terms of the factorial moments [10] should be examined to confirm that the magnitude of correction is much smaller than those of ΔF with Poisson family size without correction.
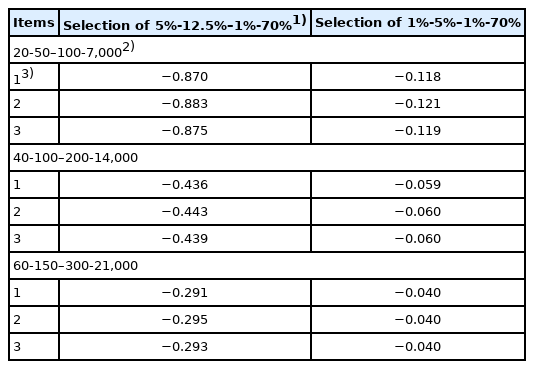
Correction factors for deviations of the variance of the family size from independent Poisson variances (×10−4) approximated by binomial distribution
Selection intensities and variance reduction coefficients should be adjusted by using the procedure from Wray and Thompson [4] in situations of few families with numerous candidates per family, for example, when the number of selected parents is only 5 or 10 [6]. Because we set the number of parents in SS at 20, 40, and 60, we did not adjust the selection intensity in the SS path. In addition, selection intensity in DD generally is much smaller than those in SS, SD, and DD selection paths. Consequently, when selection in DD is not performed, the selection intensity and reduction factor of the variance need to be set at zero in the DD selection path in the formula developed in the current study.
CONCLUSION
We here developed a formula for calculating the rates of inbreeding in populations under selection based on GEBV. The population is selected along the four selection paths of SS (sires to breed sons), SD (sires to breed daughters), DS (dams to breed sons), and DD (dams to breed daughters). Assuming that the number and selection intensity of parents remained the same over the period of usage (several years) enabled us to regard generations as discrete generations. The effect on decreasing the rate of inbreeding was greater when the number of parents was increased than when the selection intensity was decreased, and both number of parents and the selection intensity in four-path selection emerged as major factors affecting the rate of inbreeding. In general, both genetic gain and ΔF tended to increase in line with any increase in selection intensity. Therefore, increasing the number of parents is one option for offsetting the increase in ΔF due to an increase in selection intensity. Especially, increasing the number of male parents would be effective, since the accounting percentage for the increase in ΔF from male parents is greater than that from female parents. When applied without correction for deviation of family size from Poisson distributions, the formula we developed here would be highly useful as a practical method for predicting the approximate rate of inbreeding (ΔF) in populations where selection is performed according to four-path programs.
Notes
CONFLICT OF INTEREST
We certify that there is no conflict of interest with any financial organization regarding the material discussed in the manuscript.
FUNDING
The authors received no financial support for this article.
SUPPLEMENTARY MATERIAL
Supplementary file is available from: https://doi.org/10.5713/ab.21.0350
Appendice A. Elements in matrix Π
Appendice B. Elements in matrix and λ
Appendice C. Elements of ΔVSS,D,DS,or DD
ab-21-0350-suppl.pdf