Supplementing Rhodobacter sphaeroides in the diet of lactating Holstein cows may naturally produce coenzyme Q10-enriched milk
Article information
Abstract
Objective
To examine the effects of Rhodobacter sphaeroides (R. sphaeroides) supplementation as a direct-fed microbial (DFM) on rumen fermentation in dairy cows and on coenzyme Q10 (CoQ10) transition into milk, an in vitro rumen simulation batch culture and an in vivo dairy cow experiment were conducted.
Methods
The characteristics of in vitro ruminal fermentation were investigated using rumen fluids from six cannulated Holstein dairy cows at 2 h post-afternoon feeding. A control treatment was included in the experiments based on a typified total mixed ration (TMR) for lactating dairy cows, which was identical to the one used in the in vivo study, plus R. sphaeroides at 0.1%, 0.3%, and 0.5% TMR dry matter. The in vivo study employed six ruminally cannulated lactating Holstein cows randomly allotted to either the control TMR (C-TMR) treatment or to a diet supplemented with a 0.5% R. sphaeroides culture (S-TMR, dry matter basis) ad libitum. The presence of R. sphaeroides was verified using denaturing gradient gel electrophoresis (DGGE) applied to the bacterial samples obtained from the in vivo study. The concentration of CoQ10 in milk and in the supernatant from the in vitro study was determined using high performance liquid chromatography.
Results
The results of the in vitro batch culture and DGGE showed that the concentration of CoQ10 significantly increased after 2 h of R. sphaeroides supplementation above 0.1%. When supplemented to the diet of lactating cows at the level of 0.5%, R. sphaeroides did not present any adverse effect on dry matter intake and milk yield. However, the concentration of CoQ10 in milk dramatically increased, with treated cows producing 70.9% more CoQ10 than control cows.
Conclusion
The CoQ10 concentration in milk increased via the use of a novel DFM, and R. sphaeroides might be used for producing value-added milk and dairy products in the future.
INTRODUCTION
Coenzyme Q, also known as ubiquinone, is a hydrophobic lipophilic molecule synthesized by all animal tissues. It is an important component of the mitochondrial electron transport system and its reduced form (ubiquinol) works as an antioxidant [1]. The major coenzyme Q in higher plants and mammals is coenzyme Q10 (CoQ10), which contains 10 isoprenoid units [2]. Most animal-originated foods, such as meat, egg, and dairy products, are critical sources of CoQ10 [3]. Interestingly, recent studies showed that CoQ10 concentration in human maternal milk is higher in early lactation (within a couple of days post-partum) than in later lactation phases (14 days post-partum) and, more importantly, that CoQ10 concentration in human milk is highly correlated with the antioxidant capacity of milk, particularly at early lactation stages [4,5]. Other studies highlighted the age-related variation of CoQ10 (i.e., it declined with age) [6,7], despite its dietary uptake and endogenous synthesis. Therefore, supplying high levels of CoQ10 to aged or young people via food intake may help to maintain or improve their health status, as CoQ10 prevents age-related functional declines in humans.
Several microorganisms belonging to the genera Rhodobacter, Agrobacterium, and Paracoccus have been reported as high CoQ10 producers [8]. Rhodobacter sphaeroides (R. sphaeroides), in particular, produces a higher level of ubiquinone-10 than Agrobacterium tumefaciens and Paracoccus denitrificans [9]. In addition, R. sphaeroides can be cultured under variable conditions, including anaerobic respiration and fermentation conditions [10]. Thus, it can be hypothesized that supplementing these high CoQ10-producing microorganisms to ruminant diets might produce CoQ10-enriched animal products (i.e., meat and milk), if such microorganisms can inhabit, or just survive for a certain period, in the gut of the ruminants and if they can be successfully supplemented (e.g., probiotics) in the diets fed to ruminants. To produce CoQ10-enriched value-added milk, the effects of R. sphaeroides as a feed additive upon ruminal fermentation and CoQ10 transition into milk were assessed in the present study, using in vitro and in vivo experiments.
MATERIALS AND METHODS
Animal care
This study was approved by the Institutional Animal Care and Use Committee, Chung-Ang University, Seoul, Republic of Korea (NO: 2016-00105).
Preparation of Rhodobacter sphaeroides culture
R. sphaeroides belonging to the Korean Collection Type Culture 1434 strain (http://kctc.kribb.re.kr/English/index.aspx, Korea) were cultivated on Van Niel’s medium [11], containing 1.0 g K2HPO4, 0.5 g MgSO4, and 10 g yeast extract, at 20°C for 72 h, resulting in a final total concentration of 109 cfu/mL (viable units). Total bacterial counts were determined using the method of Harris and Sommers [12]. Both in vitro and in vivo experiments were conducted using these cultured microorganisms as a direct-fed microbial (DFM).
Experiment 1. In vitro rumen simulation experiment
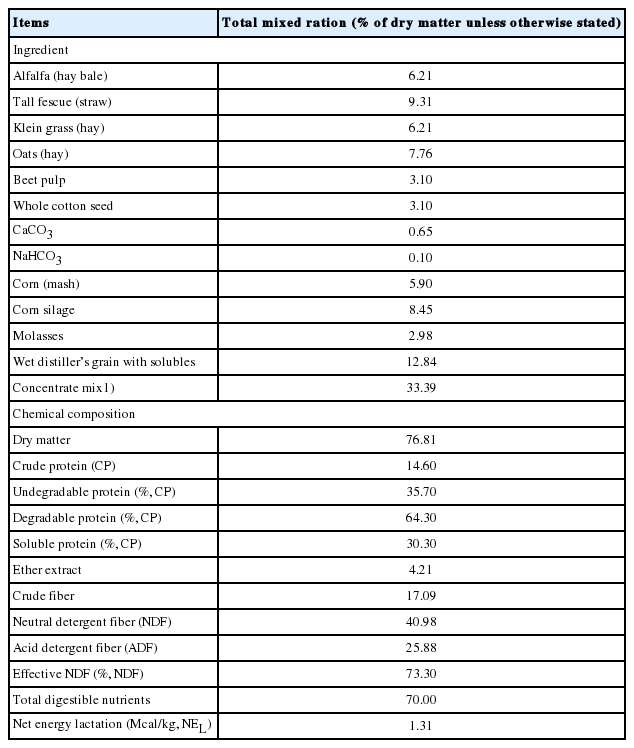
To examine the effect of R. sphaeroides supplementation on the characteristics of ruminal fermentation, an in vitro batch culture experiment was conducted using the rumen contents of six ruminally cannulated Holstein dairy cows, collected 2 h post-afternoon feeding. Approximately 1 L of rumen content from each cow were filtered through four layers of muslin, pooled into a Thermos bottle, and immediately brought to the laboratory. Experimental diets consisted of four dietary treatments, including a control diet based on total mixed ration (TMR) with no supplement and three diets supplemented with R. sphaeroides culture at 0.1%, 0.3%, and 0.5% of TMR dry matter (DM) (v/w) in triplicate. The control diet for the in vitro study was identical to the one used for the in vivo study, which was a typical TMR diet for lactating cows. The chemical composition of the basal experimental diet is presented in Table 1. For the in vitro procedure, 100 mL of artificial rumen saliva [13] was placed in serum bottles containing 0.5 g of experimental diets under anaerobic techniques, in triplicate [14]. The filtered rumen fluid was injected (10%, v/v) into the serum bottles with continuously infused O2-free CO2 gas, and these serum bottles were crimped with butyl rubber stoppers and aluminum seals before being placed in a 39°C shaking water bath (100 rpm) and incubated for 2, 4, 8, 12, and 24 h. The volume of gas produced was measured at each time point by a pressure detector (model PSGH-28PCCA, DECO Co., Seoul, Korea) connected to a digital pressure transducer (DPT-03, Dail Information Co., Seoul, Korea). The supernatant of each incubation bottle was collected for pH determination and stored at −20°C for ammonia nitrogen (NH3-N) [15], volatile fatty acids (VFAs) [16], and microbial protein synthesis [17] analyses. To determine DM digestibility, the incubated residues were transferred to a sintered glass crucible, cleaned, oven-dried, and weighed.
Experiment 2. In vivo experiment
Six Holstein lactating cows (body weight 612±27 kg, milk yield 28.4±2.3 kg/d, and parity 2nd) equipped with permanent ruminal cannulae were used in the in vivo experiment, to examine if supplementing diets with R. sphaeroides cultures affected milk production and milk composition, particularly the concentration of CoQ10 in milk. Cows were randomly allocated to one of the two dietary treatments (n = 3): control TMR (C-TMR), identical to that used in the in vitro study, and TMR supplemented (S-TMR) with 0.5% of R. sphaeroides culture (TMR dry matter basis, v/w). Cows were allowed to adapt to the experimental diets for 20 days and at the end of the adaptation to the experimental diets an aliquot of milk samples (30 mL) were then collected at the same time, over three consecutive days and pooled per cow, immediately stored at −20°C, and freeze-dried prior to the CoQ10 analysis. Milk composition was analyzed using 50 mL of milk and Milko-Scan (FOSS-4000, FOSS, Denmark).
Isolation and purification of rumen microbial DNA
For the molecular analyses, R. sphaeroides genomic DNA was isolated from the rumen fluid of the cows fed on C-TMR and S-TMR, using a previously described method with minor modifications [18]. Briefly, genomic DNA was extracted by bead-beating (BioSpec Products, Bartlesville, OK, USA) for 4 min at full speed (2,000 strokes/min) in the presence of zirconium beads (weight 0.7 g, diameter 0.1 mm), 282 μL buffer A (NaCl 0.2 M, Tris 0.2 M, ethylenediaminetetraacetic acid 0.02 M; pH 8), 268 μL buffer B (QIAquick 96 PCR purification kit, Qiagen, Hileden, Germany), and 200 μL phenol-chloroform-isoamyl alcohol (25:24:1, pH 8). After centrifugation (16,000×g for 20 min at 4°C), the supernatant was thoroughly mixed with 650 μL buffer PB (Qiagen, Germany), and DNA was purified from the sample using the Qiagen PCR purification kit following the manufacturer’s protocol.
Denaturing gradient gel electrophoresis analysis
To conduct the denaturing gradient gel electrophoresis (DGGE) analysis, the photosynthetic reaction center M subunit gene, which is specific to photosynthetic bacteria such as R. sphaeroides, was amplified using the primers GC-clamp-557f (5′-CGC ACC TGG ACT GGA C -3′) and 750r (5′-CCC ATG GTC CAG CGC CAG AA -3′) as previously described [19]. The polymerase chain reaction (PCR) was performed in a TaKaRa Bio Ins. PCR Thermal Cycler (Kusatsu, Shiga, Japan) in a 25 μL final volume containing EmeraldAmp GT PCR MASTER mix (TaKaRa Bio Ins., Japan), 1 μL each primer (GC-cramp 557f and 750r), 2 U Taq polymerase (Ex Taq, TaKaRa Bio Inc., Japan), and 1 μL template DNA. Amplification cycles started with denaturation at 95°C for 5 min, followed by 35 cycles of denaturation at 94°C for 30 s, annealing at 55°C for 30 s, and extension at 72°C for 30 s, and ended with a final extension at 72°C for 10 min. Amplification success was checked by using 2% agarose gel electrophoresis and by visualizing PCR products in a Gel Doc XR+ system (Bio-Rad, Hercules, CA, USA). Following the purification of PCR products with the QIAquick PCR purification kit (Qiagen, Germany), DGGE was conducted using a D-Code system (Bio-Rad, USA) with 8% (w/v) polyacrylamide gels containing 20% to 80% denaturant gradient in 1×TAE buffer containing a mixture of Tris base, acetic acid and ethylenediaminetetraacetic acid. Equal amounts of purified PCR products were loaded on the gel and electrophoresis was performed at 130 V for 8 h. The gel was stained in 250 mL running buffer (ethidium bromide, Bio-Rad, USA) for 20 min and stained gels were photographed under UV using the Gel Doc XR+ documentation system (Bio-Rad, USA).
Sequence analysis
After electrophoresis, bands of interest were carefully excised with a sterile razor blade under UV illumination and then placed in 100 μL TE buffer containing Tris and ethylenediamintetraacetic acid for 24 h at 4°C. This solution containing DNA was again amplified using the primer pairs mentioned above, purified, and sent to Genotech (Daejeon, Korea) for sequencing. The nucleotide sequences obtained were compared to that deposited in the GenBank database (https://www.ncbi.nlm.nih.gov/genbank/) using the basic local alignment search tool (BLAST) algorithm [20]. Sequence identities were determined based on the highest identity score.
Coenzyme Q10 analysis
The concentrations of CoQ10 in rumen contents and milk were determined in a high performance liquid chromatograph (920-LC, Varian Inc., Palo Alto, CA, USA) equipped with an ultraviolet detector [21] and a liquid chromatography column (Zobax Eclipse Plus C18, 4.6×100 mm, 5.0 μm packing; Agilent technologies, Santa Clara, CA, USA). Rumen contents (10 mL) from the in vitro study were centrifuged at 500×g for 15 min to remove feed particles; 1 mL of the resulting supernatant was then re-centrifuged at 16,000×g for 20 min to isolate rumen microbial cells. The concentration of CoQ10 in the collected cells and cow milk samples was then analyzed as previously described [22,23].
Statistical analysis
Ruminal fermentation characteristics and milk composition were subjected to analysis of variance, with diet as the main effect, using the PROC MIXED procedure in the SAS program package [24]. Where necessary, the multiple comparison was performed by Duncan’s multiple range test [25]. Significance of the treatment was tested at 5% level. Orthogonal polynomial contrasts were performed to determine linear and quadratic effects of R. sphaeroides supplement.
RESULTS
The effects of R. sphaeroides on in vitro ruminal fermentation are shown in Table 2. Gas production significantly increased (p <0.05) with increasing R. sphaeroides levels supplemented to the diet. Microbial protein synthesis also increased (p<0.05) with supplementation above 0.1% R. sphaeroides but NH3-N concentration decreased (p<0.05) with increasing R. sphaeroides concentration. Final pH, DM digestibility, total VFAs concentration, and acetate/propionate (A/P) ratio were not affected by R. sphaeroides supplementation. The concentration of CoQ10 increased (p<0.05) after 2 h for R. sphaeroides supplementation above 0.1% and CoQ10 accumulated in culture media during in vitro ruminal fermentation (Figure 1). After 12 h, CoQ10 concentration obtained for 0.5% R. sphaeroides supplementation (115.78 μg/g) was about 10-fold greater than that of the control (12.96 μg/g).
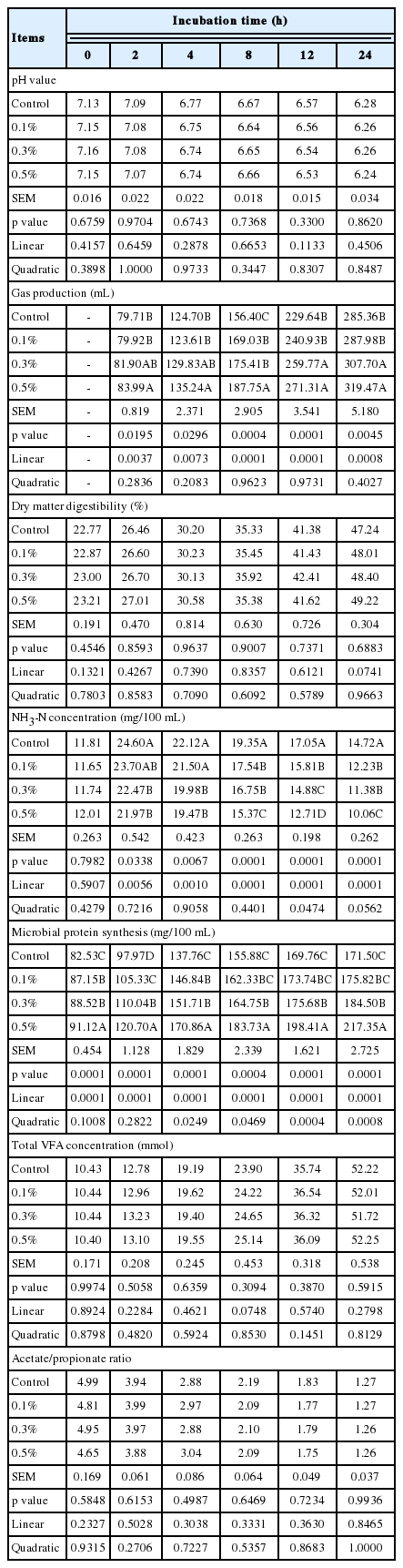
Effect of Rhodobacter sphaeroides KCTC 1434 supplementation in total mixed ration (TMR) on ruminal fermentation characteristics in vitro
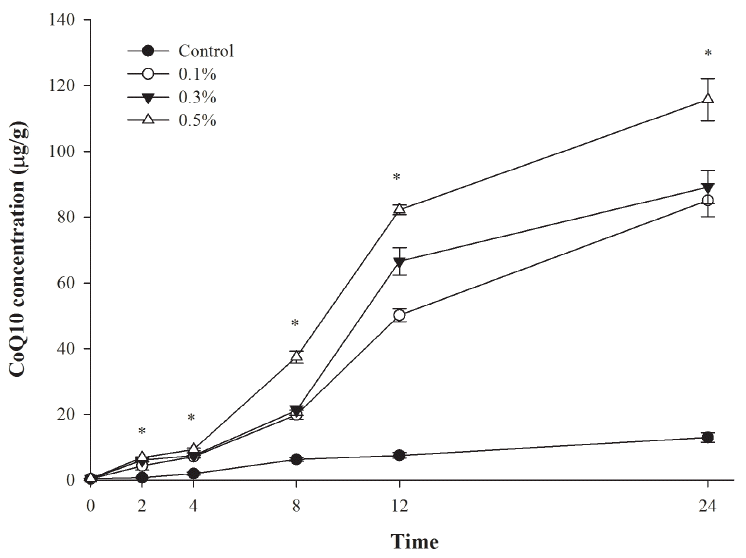
Effect of Rhodobacter sphaeroides KCTC 1434 supplementation in total mixed ration (TMR) on CoQ10 concentration in vitro. Control = TMR with no supplement; 0.1%, 0.3%, and 0.5% indicates TMR supplemented with 0.1%, 0.3%, and 0.5% R. sphaeroides). * Values differ significantly between treatments (p<0.05).
The viability of the supplemented R. sphaeroides was examined using a molecular biological technique (Figure 2). As evidenced, a unique band appeared in the rumen samples obtained from cows fed on diets supplemented with 0.5% R. sphaeroides (Figure 2C), which was hardly perceptible for the samples obtained from cows fed with the control diet (Figure 2B). This band was very intense and migrated to the same position as the band obtained for the PCR product of pure R. sphaeroides cultures. Indeed, sequencing this band revealed that it was closely related to the R. sphaeroides (98.4%) sequences deposited in GenBank. Supplementing R. sphaeroides to the diet of Holstein dairy cows did not affect DM intake, 3.5% fat-corrected milk, and somatic cell count; however, it significantly increased (p<0.05) CoQ10 concentration in milk (Table 3) and cows supplemented with R. sphaeroides produced 70.9% more CoQ10 than control cows (p<0.05).
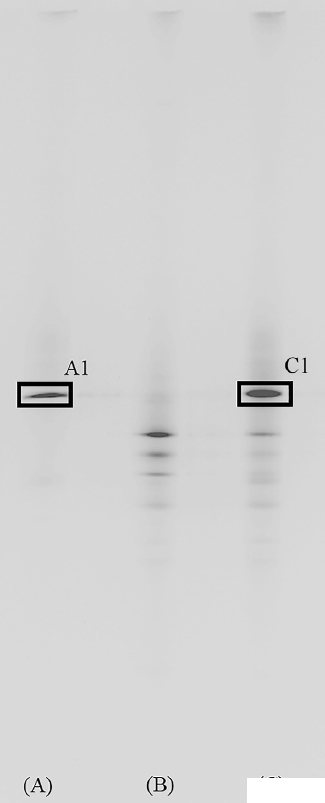
Denaturing gradient gel electrophoresis of rumen microbiota isolated at 12 h post feeding from in vivo study. (A) DNA from Rhodobacter sphaeroides (R. sphaeroides) pure culture, (B) control TMR, (C) 0.5% R. sphaeroides supplemented TMR. A1 (boxed) indicates the band amplified with photosynthetic reaction center M subunit (pufM) gene of R. sphaeroides and C1 (boxed) had 98.4% similarity with R. sphaeroides.
DISCUSSION
To the best of our knowledge, the present study is the first to demonstrate CoQ10 production in the rumen and its transition into milk, using R. sphaeroides as a DFM. Supplementation of R. sphaeroides did not show any detrimental effects on ruminal fermentation characteristics in vitro nor on animal performance in vivo, including DM intake, and milk yield and composition. However, cumulative gas production, CH4 production and CoQ10 concentration increased and NH3-N concentration decreased with increasing levels of R. sphaeroides in in vitro batch cultures, suggesting that R. sphaeroides might adapt to and inhabit the ruminal environment. Because there are limited studies available in the literature to compare with, it is difficult to explain the observed changes in ruminal fermentation characteristics when R. sphaeroides cultures were supplemented. Several scientists [26, 27] have reported that dietary supplementation of DFM, such as Saccharomyces cerevisiae, stimulates the activities of rumen microorganisms in vitro and alters the rumen environment favorably. Therefore, it is speculated that R. sphaeroides cultures might have played a role along with ruminal microorganisms, particularly with respect to the N metabolism in the rumen, as indicated by the reduced NH3-N concentration with increase in the levels of R. sphaeroides supplementation. Further investigation is warranted to examine the role of R. sphaeroides cultures on N metabolism in the rumen.
With CoQ10, previous studies [8,9] have reported that certain strains of R. sphaeroides have the ability to produce high levels of CoQ10 (2.5 mg/g of cell) in low-aeration conditions. Therefore, R. sphaeroides could produce CoQ10 under complex anoxic conditions such as the ruminal environment. This hypothesis was further confirmed by the results of the DGGE analysis, wherein R. sphaeroides specific genes were identified 12 h after their supplementation in the diet. Although further analyses, such as quantitative PCR or plate-based enumeration, were not performed to confirm the number of R. sphaeroides in the rumen, their viability was evidenced by the increased band intensity obtained in the DGGE conducted for the genomic DNA isolated from ruminal bacteria of the experimental cows. Also noteworthy were the CoQ10 concentrations that accumulated in both control and supplemented groups over time, which suggest that CoQ10 might not be degraded by ruminal microbes and thus might have the potential to be by-passed to the lower digestive tract and be absorbed. As a lipophilic substance, CoQ10 connects to the chylomicrons of the small intestine for absorption and passes through via lymphatic vessels and glands [28]. Exogenous CoQ10 supplemented to humans and most animals is non-linearly absorbed at the small intestine [29] and can be transferred from plasma to milk in humans [4]. Thus, the post-rumen CoQ10 absorption mechanism of Holstein dairy cows seems to be similar to that of humans.
In conclusion, the present study is the first to demonstrate the effect of R. sphaeroides supplementation on rumen fermentation and the transference of CoQ10 into milk. Based on in vitro and in vivo results, R. sphaeroides might be able to adapt, survive, and produce CoQ10 in the rumen environment. The produced CoQ10 might be absorbed via rumen wall or via small intestine and then be transferred to milk, although this remains to be determined. Although the present study employed a limited number of animals to test its hypothesis, it clearly demonstrated that the concentration of CoQ10 in milk can be naturally increased by direct-fed microorganisms, and therefore, it might be possible to use this beneficial microorganism for the production of value-added milk and related dairy products in the future.
ACKNOWLEDGMENTS
This research was supported by the Chung-Ang University Graduate Research Scholarship in 2015.
Notes
CONFLICT OF INTEREST
We certify that there is no conflict of interest with any financial organization regarding the material discussed in the manuscript.