Association between Single Nucleotide Polymorphisms of the Major Histocompatibility Complex Class II Gene and Newcastle Disease Virus Titre and Body Weight in Leung Hang Khao Chickens
Article information
Abstract
The aim of the present study was to investigate the effect of single nucleotide polymorphisms in the major histocompatibility complex (MHC) class II gene on resistance to Newcastle disease virus and body weight of the Thai indigenous chicken, Leung Hang Khao (Gallus gallus domesticus). Blood samples were collected for single nucleotide polymorphism analysis from 485 chickens. Polymerase chain reaction sequencing was used to classify single nucleotide polymorphisms of class II MHC. Body weights were measured at the ages of 3, 4, 5, and 7 months. Titres of Newcastle disease virus at 2 weeks to 7 months were determined and the correlation between body weight and titre was analysed. The association between single nucleotide polymorphisms and body weight and titre were analysed by a generalized linear model. Seven single nucleotide polymorphisms were identified: C125T, A126T, C209G, C242T, A243T, C244T, and A254T. Significant correlations between log titre and body weight were found at 2 and 4 weeks. Associations between single nucleotide polymorphisms and titre were found for C209G and A254T, and between all single nucleotide polymorphisms (except A243T) and body weight. The results showed that class II MHC is associated with both titre of Newcastle disease virus and body weight in Leung Hang Khao chickens. This is of concern because improved growth traits are the main goal of breeding selection. Moreover, the results suggested that MHC has a pleiotropic effect on the titre and growth performance. This mechanism should be investigated in a future study.
INTRODUCTION
In Thailand, there are many varieties of indigenous chickens, including the Leung Hang Khao (LHK) variety. In 2001, the Thailand Research Fund and the Department of Livestock Development (DLD) cooperated to collect four lines of Thai indigenous chickens: Phradhu Hang Dum, LHK, Dang, and Chee. All lines were given to the DLD, which has a remit to maintain these original lines, for which random mating is used to produce the replacement herd. Male and female LHK chickens were drawn from the base population by researchers from the Suranaree University of Technology to investigate the potential of this line for genetic development.
Thai indigenous chickens are characterized by a desirable meat texture and meat flavour (Wattanachant et al., 2004; Jaturasitha et al., 2008; Teltathum and Mekchay, 2009;2010). LHK meat is accepted by the high end of the market, and LHKs command a higher price than commercial broiler meat. The skin and shank of the LHK chicken are yellow in colour. Thai indigenous chickens are also highly disease-resistant (Boonyanuwat et al., 2006). Their higher costs, however, are an obvious disadvantage. The higher cost is related to their slow growth: they need 4 months to increase their body weight to 1.5 kg, and they need more feed (about 4 kg) to produce 1 kg of body weight (Na Rangsri et al., 2008). These disadvantages caused fluctuations in their availability in the market. Therefore, from a commercial point of view, genetic improvement in growth performance is needed to maintain the uniqueness of this breed and its ability to resist disease. However, the question remains whether during selection for growth, their disease resistance will be affected, and if so, whether the effect will be positive or negative.
Previous studies showed that there is a relationship between immunocompetence and production in animals. A review by Pieter and Stephen (2014) described the main relationship between immunology and animal production, whereby an animal’s immunocompetence is influenced by that animal’s production potential. Furthermore, the study of Kuhnlein et al. (1997) found an association between a growth hormone gene and egg production traits, resistance to Marek’s disease, and avian leucosis in White Leghorn chickens. Moreover, Rauw et al. (1998) and Masilamani (2003) found a negative relationship between growth performance and the efficiency of the immune system in chickens. These findings are in agreement with the review of Lamont (1998), which reported that some economically important traits, such as embryonic, juvenile and adult mortality, body weight, fertilization rate, hatchability, and egg production are influenced by the major histocompatibility complex (MHC) genotype. Similarly, Lakshmanan et al. (1997) observed that the MHC gene is not only associated with the mechanism of resistance to disease, but also with egg production traits.
The MHC class II gene is well known for its close relationship with disease resistance mechanisms (Chen et al., 1997) and is located on chromosome 16 (Li et al., 1997). Previous studies reported that MHC genes have a significant relationship with the trait of disease resistance in various breeds of chicken (Lakshmanan et al., 1997; Weigend and Lamont, 1999; Boonyanuwat et al., 2006).
The MHC class II gene may not only be involved in disease resistance, but also in the growth of LHKs. Therefore, it is reasonable to investigate the effect of MHC class II on resistance to disease during the process of genetic selection for growth. The objective of this study, therefore, was to investigate the association between the single nucleotide polymorphisms (SNPs) of the MHC gene and the titre of Newcastle disease virus, and the bodyweight traits of LHK chickens. The results will be useful in the design of breeding programs to increase the body weight of LHK chickens.
MATERIALS AND METHODS
Animal and data collection
Thai indigenous chicken, LHK, was used for this study. In 2001, the Thailand Research Fund (TRF) and the DLD cooperated to collect four lines of Thai indigenous chickens from around the country. LHK is one of four breeds which was collected and was given to the DLD, which has a remit to maintain these original lines. Random mating was used to produce the replacement herd. In 2009, 60 LHK males, and 300 LHK females were drawn from the herd at the DLD station and were moved to Suranaree University of Technology. Random mating between 60 LHK males and 300 LHK females was used to produce 485 LHK chicks.
They were raised by a conventional raising system, with free access to feed and water from hatching to 3 weeks. Thereafter, they received a starter feed (21% crude protein, and 3,100 Kcal/ME kg) from 3 to 6 weeks, and from 6 weeks to slaughter, they were fed with the first (19% crude protein, 3100 Kcal/ME kg) and the last step of the grower diet (17% crude protein, 3,100 Kcal/ME kg), respectively.
All chickens were tagged with an ID for individual data collection. Individual body weights were determined at hatching 0, 2, 4, 6, 8, 10 weeks, and 3, 4, 5, 6, and 7 months. On the 7th day, 28th day, 8th week, and 22nd week they were vaccinated with Newcastle disease’s virus (NDV) attenuated vaccine. When the chickens reached 2 and 4 weeks, and 3, 4, 5, 6, and 7 months, their blood was drawn, and NDV titre was measured using an enzymebmdson mii dnknil-ada tkktn kit (BioChek Smart Veterinary Diagnostics, Scarborough, ME, USA).
Genotyping
Blood samples were collected from all of the chickens for genotyping. Genomic DNA was extracted from whole blood using a DNA Mini Kit Protocol-Blood (Geneaid Biotech Ltd, New Taipei City, Taiwan). The DNA solution was quantified spectrophotometrically and diluted to 10 μg/μL. The genotyping of MHC class II was carried out using polymerase chain reaction (PCR) with primers reported by (Boonyanuwat et al., 2006; Liu et al., 2009).
The PCR products were purified and sequenced. All of the sequences were aligned with the reference sequence of the gene (GenBank accession NM-001044679.1) using BLAST 2.2.22. The sequence used in this study had to have a similarity ≥85%. Clustal W2 compared all of the sequences with each other to identify SNPs.
Statistical analyses
The titre data showed high variation (Table 1); therefore, they were transformed by natural logs, and the transformed data were used for statistical analyzes.

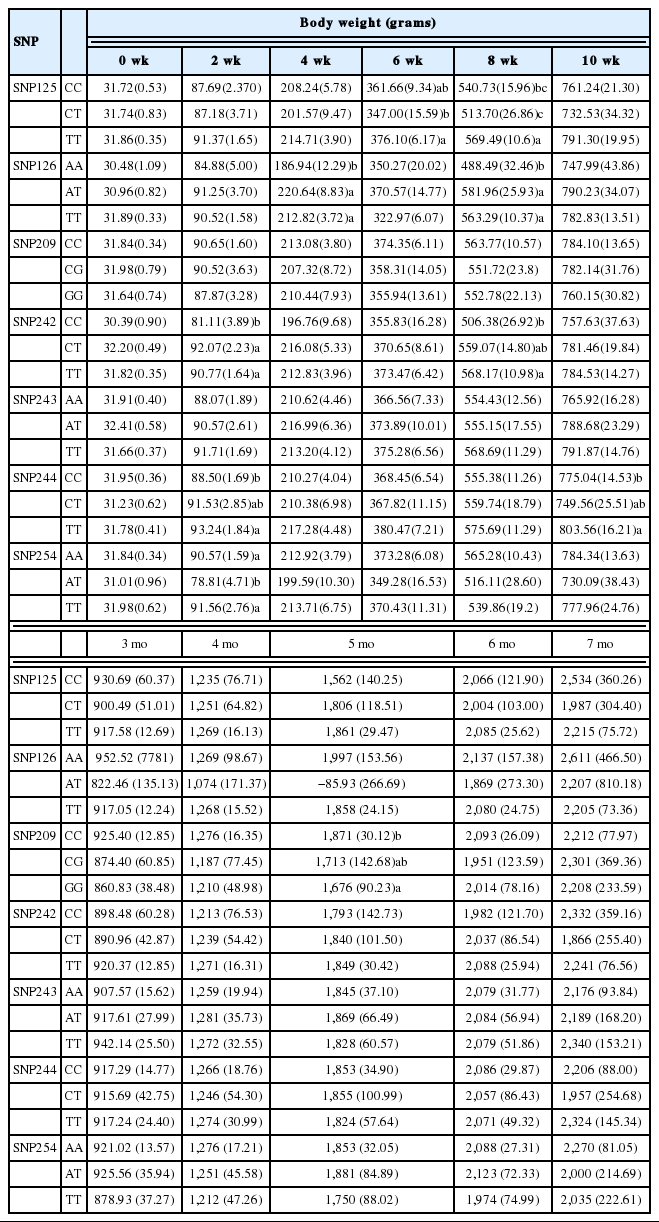
Mean and standard deviation (in parentheses) of Newcastle disease titre and body weight of Leung Hang Khao chickens
Correlations between the log of NDV titre, and body weight at the 2nd and 4th weeks, and 3rd, 4th, 5th, 6th, and 7th months were analyzed.
Linkage disequilibrium (LD) between SNPs was analysed using GENEPOP version 3.4 (Laboratiore de Genetique et Environment, Montpellier, France; Raymond and Rousset, 2003). An association was considered significant when p<0.005. The group of SNPs with significant associations were re-arranged as composite genotype.
The significant effects of composite genotype on the log of NDV titre and body weight were analyzed using a general linear model, which is shown below. The effects of composite genotype on the traits were estimated by the ordinary least squares method. Differences were considered significant at a p-value of 0.1. SPSS for Windows software (Release 10.0; SPSS Inc., Chicago, IL, USA) was used to carry out the analysis.
General linear model:
Where, y is the phenotype data (body weight or the log of NDV titre), X is an incidence matrix that represented the sex and the composite genotype, β is the regression coefficient of sex and genotypes, and ε is the residue effect.
RESULTS
Single nucleotide polymorphisms and linkage disequilibrium
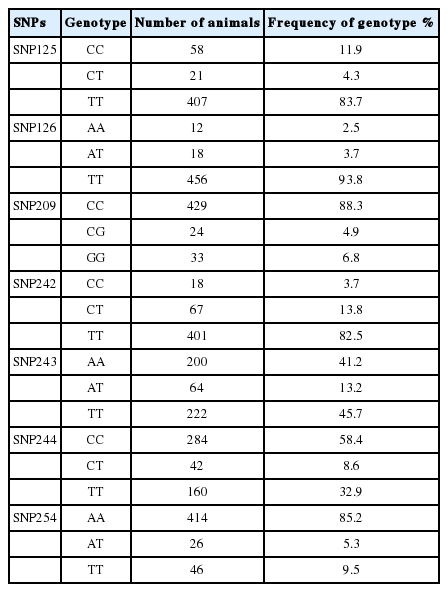
Seven SNPs were identified: C125T, A126T, C209G, C242T, A243T, C244T, and A254T. All of these SNPs have three genotypes (Table 2). Significant deviations from the Hardy-Weinberg Equilibrium (HWE) were detected for all SNPs, and a high frequency of the homozygous genotype was observed for most of the SNPs in this population.
Correlation between body weight and the log of the Newcastle disease’s virus titre
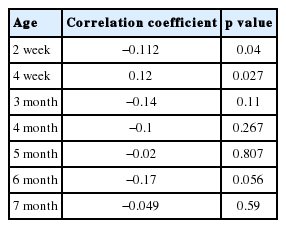
The correlation was used to analyse the relationship between body weight and the log of titre of NDV (Table 3). A low, negative correlation coefficient was observed for most ages of the chickens. These results indicated that these two traits have little relationship with each other. After the genetic selection, there was a small negative impact.
The relationship between single nucleotide polymorphisms of MHC class II and the log of titre of Newcastle disease and body weight
Significant associations between the gene and the NDV titre were detected for SNPs C209G and A254T at 3, 4, 5, 6, and 7 months. There was no significant association in younger chickens (Table 4).
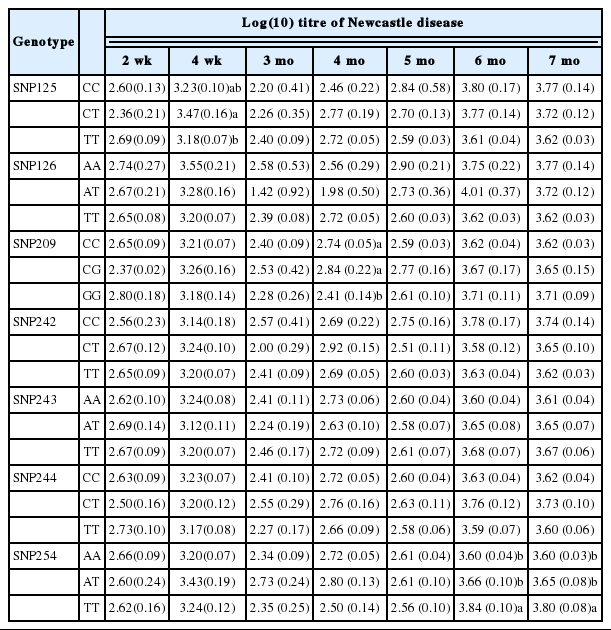
Effect of single nucleotide polymorphisms on log(10) titre of Newcastle disease in Leung Hang Khao chickens
A significant relationship between SNPs and body weight was detected for most of the SNPs, except for SNP243. The most significant association was observed at an early age in LHK chickens (Table 5).
Interestingly, SNP209 and SNP254 showed a significant association between NDV titre and body weight. SNP209 was associated with titre and body weight at 4 and 5 months, respectively. SNP254 had an association with titre at 6 and 7 months of age, and with body weight at 2 weeks. SNP209 genotypes CC and CG had a greater effect on both titre and body weight than genotype GG. Similarly, SNP254 genotype TT had the greatest effect on both titre and body weight. These results suggest that SNP209 and SNP254 of MHC class II could be used in breeding programs where the goals include increased growth and resistance to Newcastle disease virus. A pleiotropic effect might explain the results; however, the mechanism remains unclear.
DISCUSSION
We identified seven SNPs in the current study, but only SNP A126T was also detected by Li et al. (2012) and Liu et al. (2009), who investigated a population of Chinese indigenous chickens. A comparison of the populations could provide information about the common ancestor of the populations and their allelic diversity.
All SNPs deviated from HWE. In theory, in a large, random-mating population with the absence of migration, mutation and selection, such a population should be in HWE (Falconer, 1996). The population of LHK was randomly mated, and there were neither migration nor artificial selection. Natural selection has occurred over a long period of time in South-East Asia, and this may be the reason why the SNPs deviated from the HWE. Moreover, natural selection has also led to Thai indigenous chickens having disease resistance.
A low, negative correlation between body weight and NDV titre was found in this study. This did not agree with the studies of Lamont (1998), Rauw et al. (1998) and Masilamani (2003), which found a relationship between growth performance and the efficiency of the immune system in chickens. By contrast, Van der Most et al. (2011) found that in a line selected for immune function, the effect on growth was close to zero, which suggested that it might be possible to breed animals for increased growth without loss of immune function. Different genotypes of each breed or line may explain the inconsistency of these results. Moreover, in the current study, which used LHK chicken, low growth performance was observed: even the fastest-growing LHK chickens had a lower growth performance than that of commercial broilers, which may be another explanation for why there is a low, negative correlation in the present study. Therefore, genetic selection for growth may be conducted with concern for a negative effect on disease resistance, particularly when the selection continues for several generations, and growth performance is increased.
Significant associations between the SNPs and NDV titre were detected. The results support those of Du et al. (2011), Guo et al. (2012), Li et al. (2012), and Liu et al. (2009), who studied the association between various indigenous chickens. Different SNPs, however, were observed using these different populations. The different genetic structures of various breeds of chicken are the main cause of the differences in relevant SNPs. These results are not a new finding, but they confirm the association between the MHC gene and the immune system, and could be used as a benchmark for conducting selection for growth in Thai indigenous chickens.
Associations between SNPs and body weight were also detected. Previous review articles (Bacon, 1987; Lamont, 1998) reported that the immune response of chickens has a role in growth performance and other production traits. In theory, the immune system could have a connection to the central nervous system that may involve the release of hormones such as Adrenocorticotropic hormone and Thyrotropin. These hormones have a role in the process of gluconeogenesis, which affects protein synthesis and protein accumulation in muscle, and may also have an effect on the body weight of animals (Jepson et al., 1986). Furthermore, Pinard-van der Laan (2002) found that selection for improvement of the immune system affected growth performance. Thus, we speculated that growth performance and resistance to disease are associated with one another.
In the present study, significant associations between MHC class II SNPs and NDV titre and body weight were detected in Thai indigenous chicken LHK. Additionally, even when the correlation was not strong, it was still statistically significant. However, when growth traits are the main purpose of breeding, the impact on disease resistance should be monitored. Moreover, the results suggested that MHC class II has a pleiotropic effect on the titre and growth performance. Therefore, the mechanism of this effect should be investigated in a future study.
ACKNOWLEDGMENTS
The authors are grateful to the Suranaree University of Technology for financial support. We also thank the Department of Livestock Development and the project Establishment of “Korat Meat Chicken” Strain for Small and Micro Community Enterprise (SMCE) Production, phase II (financial support by Thailand Research fund), for supplying the Thai indigenous chickens for research.