 |
 |
| Anim Biosci > Volume 33(5); 2020 > Article |
|
Abstract
Objective
Cattle were some of the first animals domesticated by humans for the production of milk, meat, etc. Long noncoding RNA (lncRNA) is defined as longer than 200 bp in non-protein coding transcripts. lncRNA is known to function in regulating gene expression and is currently being studied in a variety of livestock including cattle. The purpose of this study is to analyze the characteristics of lncRNA according to sex in Hanwoo cattle.
Methods
This study was conducted using the skeletal muscles of 9 Hanwoo cattle include bulls, steers and cows. RNA was extracted from skeletal muscle of Hanwoo. Sequencing was conducted using Illumina HiSeq2000 and mapped to the Bovine Taurus genome. The expression levels of lncRNAs were measured by DEGseq and quantitative trait loci (QTL) data base was used to identify QTLs associated with lncRNA. The python script was used to match the nearby genes
Results
In this study, the expression patterns of transcripts of bulls, steers and cows were identified. And we identified significantly differentially expressed lncRNAs in bulls, steers and cows. In addition, characteristics of lncRNA which express differentially in muscles according to the sex of Hanwoo were identified. As a result, we found differentially expressed lncRNAs according to sex were related to shear force and body weight.
Cattle were some of the first animals to be domesticated by humans for agricultural purposes. About 10,000 years ago, the ancestors of cattle were tamed for the purpose of providing milk, meat, and leather to humans [1]. In 2006, a human genome sequencing center announced the sequence of the cattle genome [2]. Utilizing the genome-wide single nucleotide polymorphism panel for cattle allows for the use of quantitative trait loci (QTL) and prediction of the genetic merit of an animal without using phenotype and pedigree records [3]. Genetic studies using livestock have shown that most of the genetic variants associated with complex phenotypic traits are located outside the protein coding region [4].
Analysis of intracellular transcripts has revealed that 50% of the transcribed genomes were not aligned with known protein coding regions. Many of these have been proven to have protein coding potential [5]. Non-protein coding transcripts can potentially be noncoding RNA (ncRNA) [6]. Long noncoding RNA (lncRNA) is a molecule longer than 200 bp in a non-protein-coding transcript, or one longer than 2 kb with a coding potential of an amino acid sequence less than 100 bp [7]. LncRNA is a relatively new class of RNA molecules that are less well-characterized than microRNA (miRNA). It has been characterized only by some functional lncRNA and has been shown to regulate all levels of the gene-regulated expression pathway [8]. Previous studies have confirmed that lncRNA plays an important role in a variety of key biological processes, including translational control, RNA splicing, and chromatin modification [9]. In addition, functional studies of lncRNAs have shown that they play an important role in basic biological processes such as dose compensation, transcriptional regulation, and epigenetic regulation [10–12]. The number of studies on lncRNA has continued to increase and many databases have been constructed that include lncRNA data for domesticated animals [13]. At the time of the announcement, 12,103 pig lncRNAs, 8,923 chicken lncRNAs, and 8,250 cattle lncRNAs were included in the ALDB (domestic-animal lncRNA database) database [14].
This study was conducted to compare the expression of lncRNA in bulls, steers and cows as well as the characteristics of each sex. Samples from muscles tissues were used for each sex, and the expression of lncRNA by tissue was analyzed according to sex. We also analyzed QTL associated with lncRNA, which is significantly expressed.
All analysis was conducted with data reprocessed from the law data from a previous study [15]. The animals and sample preparation were as follows: A total of nine (three bulls, three steers and three cows) Hanwoo cattle (Bos taurus coreanae) were used in this study. They were fed the same diet and managed at the same location, Hanwoo Experimental Station in National Institute of Animal Science, throughout the experiment. The average (±standard deviation) carcass weight of the cattle at slaughter was 430.2 (±40.66) kg. Immediately after slaughter, muscle tissues were separated and sampled. All of the tissue samples were immediately frozen using liquid nitrogen and stored at −80°C until analysis. All animal use, care, and experimental protocols for this experiment were reviewed and preapproved by the Institutional Animal Care and Use Committee of the National Institute of Animal Science (number 2010-042). Total RNAs of tissues were isolated using TRIzol (Invitrogen, Karlsruhe, Germany) and an RNeasy RNA purification kit with DNase treatment (Qiagen, Hilden, Germany). The messenger RNA (mRNA) was isolated from the total RNA using oligo-dT beads and was reverse transcribed into double-stranded cDNA fragments. Constructing and sequencing of the RNA-seq library for each sample was carried out based on the protocols of Illumina HiSeq2000 in order to generate 101 pair-end reads. The qualities of the RNA-seq reads from all of the tissues were checked using FastQC.
Filtering was conducted in order to remove the low-quality sequences. The filtered sequences were mapped to Bovine Taurus genome (bosTau6) using STAR v2.4.0b [16]. Expression levels were calculated using Cufflinks v2.2.1. Bovine gene information was used to measure expression levels [17]. Multi read correction and frag bias correct options were additionally used to increase the accuracy of expression measurement, and other options were used as defaults. The bovine lncRNA analysis was used to conduct mapping with reference to annotated bos taurus ensemble ID results. We refer to the lncRNA list of cattle identified in the study of Koufariotis et al [18]. We matched the ensemble ID of the transcripts we found and identified lncRNA in the muscle of Hanwoo (ensembl.org).
Differences in the expression level of each tissue were expressed by Heatmap using R package gplots (v3.0.1). LncRNA principal component analysis (PCA) plots were used to identify the differences between bulls, steers and cows muscle tissues using Mev (http://mev.tm4.org/). The DEGseq R package was used to identify differential expression of lncRNA between the bulls, steers and cows muscle tissues. Significant lncRNAs were identified using cut-off of: |fold difference| ≥1 and p-value ≤0.05. Sex-specific differentially expressed lncRNAs statistics analysis were using Prism 5 program (San Diego, CA, USA). The Venn diagram was used to identify the lncRNA assemblages that were extracted between the bulls, steers and cows. The Venn diagram was displayed using InteractiVenn (http://www.interactivenn.net/).
QTL regions for comparative analysis with lncRNAs were identified from the cattle QTL Database (http://www.animalgenome.org/cgi-bin/QTLdb/BT/index). In these cattle, the QTL associated with the quality and productivity of the meat was selected. The position of the selected QTL was compared with the lncRNA expressed in sex.
The location of lncRNA was identified by ensembl biomart (ensembl.org/biomart) using transcript ID. The databases used included ensemble genes 92 cow genes (UMD3.1). We found for by nearby gene predicted to be affected by lncRNA. The position information of lncRNA in the genome and the gene position information of the bovine were matched using Python script. Transcription directions of lncRNA and nearby genes were used transcript information by ensembl (ensembl.org).
All analyses were conducted by reprocessing data from previous studies (Lee et al [15]). The RNA-seq raw data information is as follows: 34.2 Mb of raw readings were averaged on the muscle. More than 99.5% readings were retained after being filtered by quality control, and over 95.9% of these were mapped to the reference genome. The average length of the lncRNA was 866 bp, the minimum length was 257 bp, and the maximum length was 1,911 bp.
A hierarchical clustering analysis of the transcripts of sex transcripts in muscle tissue was performed in Hanwoo cattle. Clustering analysis of steer expressions did not show any significant differences (Figure 1). Clustering analysis using PCA analysis showed that the distances between the three groups differed by sex. The distance between the bull and the steer was relatively close to the distance to the cow (Figure 2). An analysis of the differential expression of lncRNA in each sex was conducted and found seven lncRNAs in bulls, nine lncRNAs in steers, and five lncRNAs in cows (Table 1, Figure 3A). The results of Venn diagram analysis for the found lncRNA revealed that 5 lncRNAs were found commonly in bulls and steers. We found one common lncRNA that was differentially expressed in bulls and cows. It was also identified that there was no overlapped lncRNA in the steers and cows (Figure 3B).
Functional traits are defined as characteristics of animals that increase efficiency by reducing the cost of breeding [19]. Livestock species have long been selected to improve their characteristics that are associated with economic benefits. These traits are generally affected by genetic and environmental factors [20]. The detection of genetic markers closely linked to loci affecting quantitative traits (QTL) will provide a tool for the application of marker-assisted selection and is a prerequisite for the detailed molecular genetic analysis of traits [21]. Previous studies have identified 7188 lncRNAs from cows and have identified 2753 loci on chromosomal regions of livestock QTL related to muscle development. The function of QTL was also related to intramuscular fat, lean meat yield, longissimus muscle area, shear force [22].
In this study, we conducted QTL analysis according to sex in Hanwoo muscle. QTL loci related to meat quality and productivity of cattle were selected with reference to a cattle QTL database. We identified 33 QTLs with overlapping loci with lncRNAs. Ten QTLs were identified in the bulls, 16 QTLs were identified in the steers, and seven QTLs were identified in the cows. In all sexes, lncRNA was found to be mainly associated with shear force QTL (Figure 4, Table 2).
Most lncRNAs have lower conservation and expression levels than mRNA [23]. It has been reported that lncRNA can regulate the expression of adjacent protein coding genes through transcriptional activation and inhibition [24,25]. LncRNAs can be targeted by miRNAs and it has been reported that they can regulate the expression of mRNA [26,27]. The miRNAs consist of a length of about 22 nucleotides and function like silencing RNA and regulating gene expression after transcription [28,29]. In this study, three mRNAs were found to overlap with three lncRNAs. In addition, we identified and classified the transcriptional direction of lncRNA and mRNA (Table 3).
This study profiled lncRNA, which differentially expresses the difference of sex in Hanwoo cattle. We identified the expression patterns of transcripts in each sex by comparing the bull, steer and cow strains and identified 21 lncRNAs. We identified lncRNA related to meat quality and productivity according to sex in Hanwoo through QTL analysis. Many lncRNAs were found to overlap with QTL loci associated with shear force and body weight. We also identified adjacent genes that overlap with the three lncRNAs. It will be helpful to study the genetic characteristics of Hanwoo in the future.
ACKNOWLEDGMENTS
This This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (2019R1A2C1089807) and the Next-Generation BioGreen 21 Program (PJ01315101, PJ01321901), Rural Development Administration, Republic of Korea.
Figure 1
Comparison of sex transcripts expression in Hanwoo muscles. Expression of transcripts by muscle tissue of bull, steer and cow were identified. Expression of each sex was expressed as z-score, with red as negative and blue as positive.
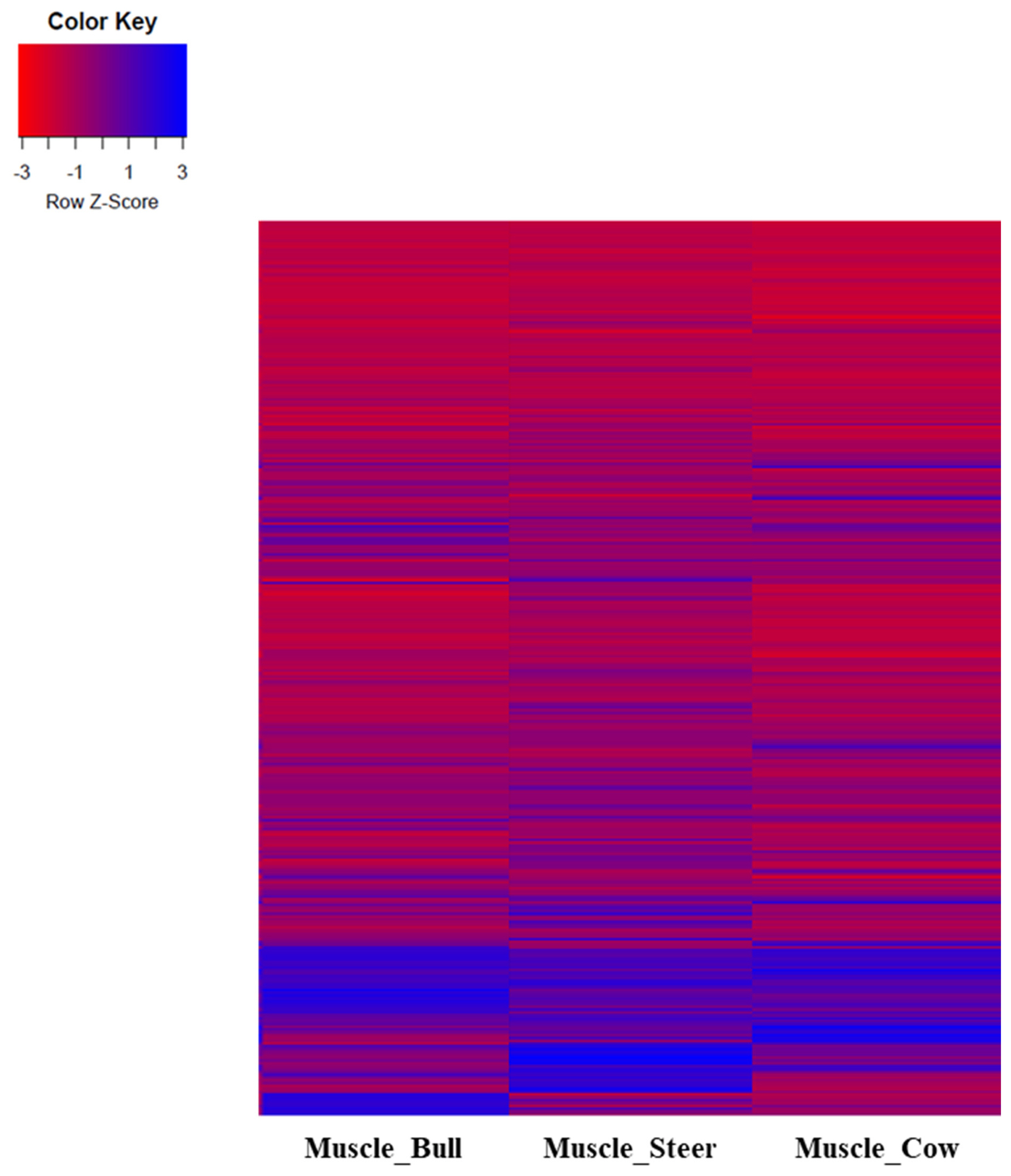
Figure 2
lncRNA principal component analysis plot of bulls, steers and cows in muscle tissues. Clustering analysis was conducted to determine the differences between the three sex. The distance between bull and steer is relatively close compared to cow. Red color is bull, blue color is steer and green color is cow.

Figure 3
Number of lncRNA found between each sex. (A) Number of lncRNAs by sex in muscle. (B) Venn diagram showing the co-expression of lncRNAs in bulls, steers and cows. Many of the lncRNAs from Bull and Steer were found to overlap. Red color is bull, blue color is steer and green color is cow.
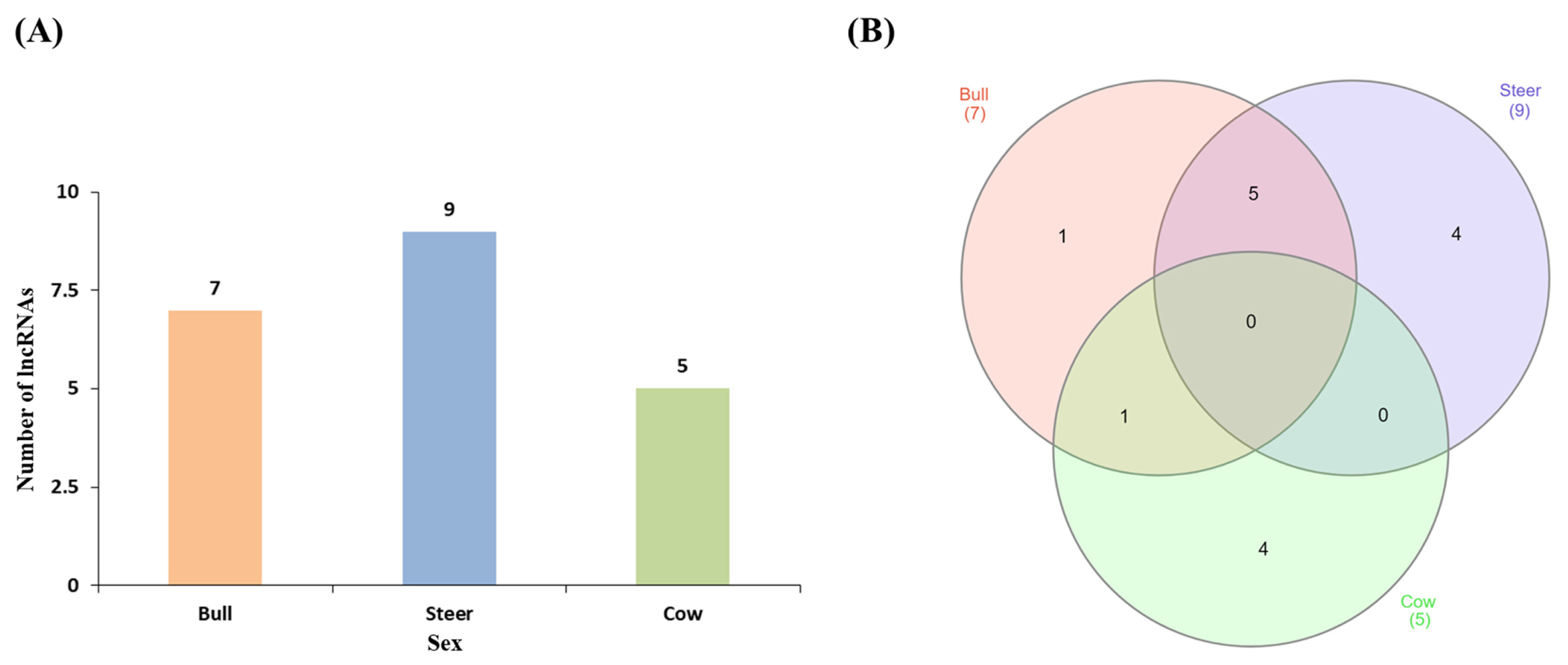
Figure 4
Number of quantitative trait loci (QTLs) associated with Hanwoo cattle economic traits by sex in muscle. (A) Number of QTL loci associated with lncRNA by each sex. (B) QTL locus associated with lncRNA found in bull. (C) QTL locus associated with lncRNA found in steer. (D) QTL locus associated with lncRNA found in cow. Compared to other sex, many QTL loci were identified in steer. All three sex were found to be associated with shear force. Red color is bull, blue color is steer and green color is cow.
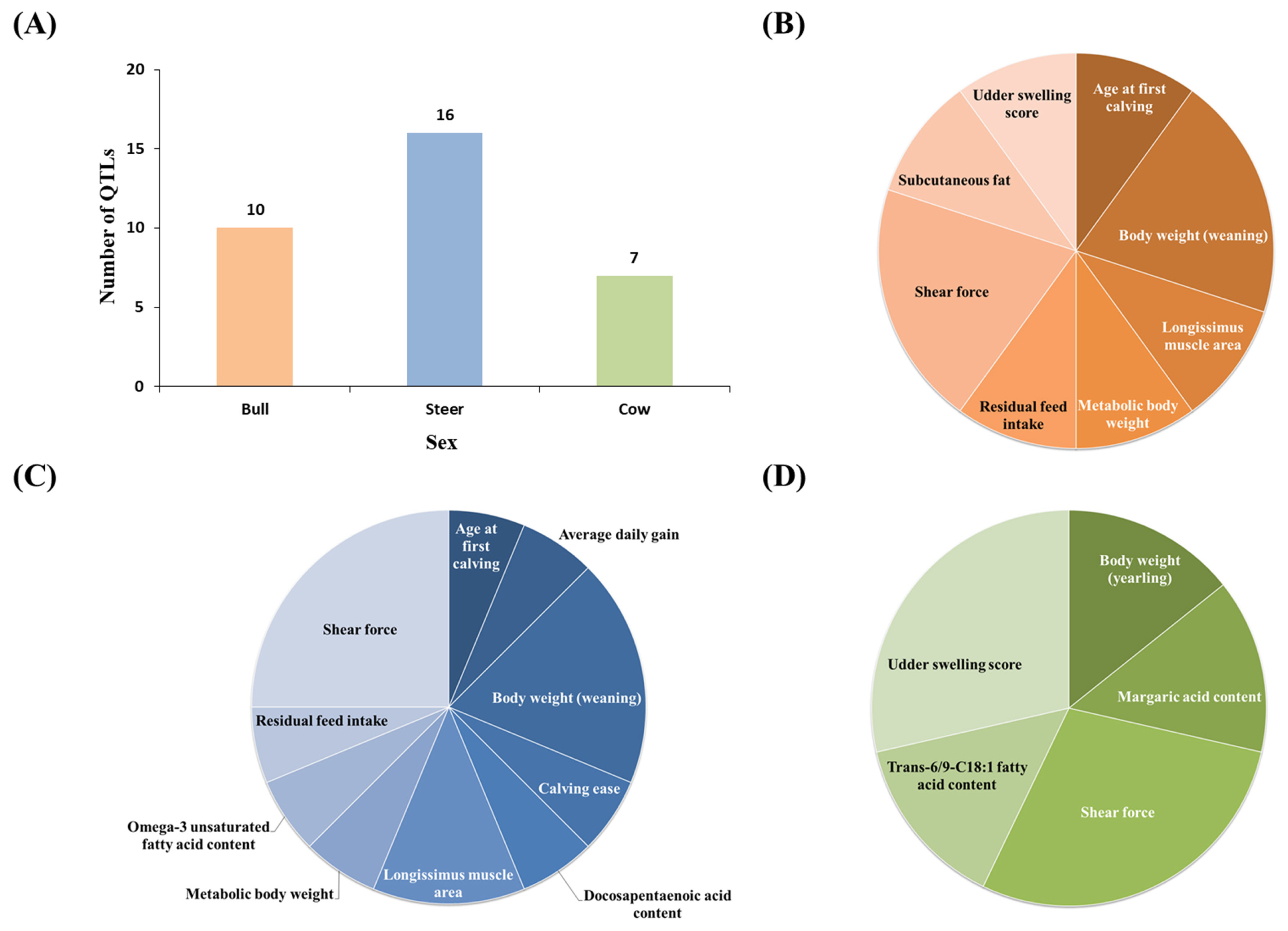
Table 1
List of differentially expressed lncRNA of Hanwoo cattle in muscle
Table 2
List of QTLs associated with Hanwoo cattle economic traits
Table 3
Information on lncRNAs with nearby genes
REFERENCES
1. Edwards CJ, MacHugh DE, Dobney KM, et al. Ancient DNA analysis of 101 cattle remains: limits and prospects. J Archaeol Sci 2004; 31:695–710.
https://doi.org/10.1016/j.jas.2003.11.001

2. Bovine Genome S, Analysis C, Elsik CG, et al. The genome sequence of taurine cattle: a window to ruminant biology and evolution. Science 2009; 324:522–8.
https://doi.org/10.1126/science.1169588



3. Goddard ME, Hayes BJ. Mapping genes for complex traits in domestic animals and their use in breeding programmes. Nat Rev Genet 2009; 10:381–91.
https://doi.org/10.1038/nrg2575



4. Weikard R, Demasius W, Kuehn C. Mining long noncoding RNA in livestock. Anim Genet 2017; 48:3–18.
https://doi.org/10.1111/age.12493


5. Hung T, Chang HY. Long noncoding RNA in genome regulation: prospects and mechanisms. RNA Biol 2010; 7:582–5.
https://doi.org/10.4161/rna.7.5.13216



6. Kapranov P, St Laurent G, Raz T, et al. The majority of total nuclear-encoded non-ribosomal RNA in a human cell is ‘dark matter’ un-annotated RNA. BMC Biol 2010; 8:149
https://doi.org/10.1186/1741-7007-8-149



7. Guttman M, Amit I, Garber M, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 2009; 458:223–7.
https://doi.org/10.1038/nature07672




8. Wapinski O, Chang HY. Long noncoding RNAs and human disease. Trends Cell Biol 2011; 21:354–61.
https://doi.org/10.1016/j.tcb.2011.04.001


9. Wutz A, Gribnau J. X inactivation Xplained. Curr Opin Genet Dev 2007; 17:387–93.
https://doi.org/10.1016/j.gde.2007.08.001


10. Duret L, Chureau C, Samain S, Weissenbach J, Avner P. The Xist RNA gene evolved in eutherians by pseudogenization of a protein-coding gene. Science 2006; 312:1653–5.
https://doi.org/10.1126/science.1126316


11. Orom UA, Derrien T, Beringer M, et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 2010; 143:46–58.
https://doi.org/10.1016/j.cell.2010.09.001



12. Lee JT. Epigenetic regulation by long noncoding RNAs. Science 2012; 338:1435–9.
https://doi.org/10.1126/science.1231776


13. Yoon J, Kim H. Multi-tissue observation of the long non-coding RNA effects on sexually biased gene expression in cattle. Asian-Australas J Anim Sci 2019; 32:1044–51.
https://doi.org/10.5713/ajas.18.0516



14. Li A, Zhang J, Zhou Z, Wang L, Liu Y, Liu Y. ALDB: a domestic-animal long noncoding RNA database. PLoS One 2015; 10:e0124003
https://doi.org/10.1371/journal.pone.0124003



15. Lee H-J, Park H-S, Kim W, Yoon D, Seo S. Comparison of metabolic network between muscle and intramuscular adipose tissues in Hanwoo beef cattle using a systems biology approach. Int J Genomics 2014; 2014:Article ID 679437. https://doi.org/10.1155/2014/679437


16. Dobin A, Davis CA, Schlesinger F, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013; 29:15–21.
https://doi.org/10.1093/bioinformatics/bts635



17. Trapnell C, Williams BA, Pertea G, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 2010; 28:511–5.
https://doi.org/10.1038/nbt.1621




18. Koufariotis LT, Chen YP, Chamberlain A, Vander Jagt C, Hayes BJ. A catalogue of novel bovine long noncoding RNA across 18 tissues. PLoS One 2015; 10:e0141225
https://doi.org/10.1371/journal.pone.0141225



19. Groen AF, Steine T, Colleau J-J, Pedersen J, Pribyl J, Reinsch N. Economic values in dairy cattle breeding, with special reference to functional traits. Report of an EAAP-working group. Livest Sci 1997; 49:1–21.
https://doi.org/10.1016/S0301-6226(97)00041-9

20. Boichard D, Grohs C, Bourgeois F, et al. Detection of genes influencing economic traits in three French dairy cattle breeds. Genet Sel Evol 2003; 35:77
https://doi.org/10.1186/1297-9686-35-1-77




21. Kuhn C, Freyer G, Weikard R, Goldammer T, Schwerin M. Detection of QTL for milk production traits in cattle by application of a specifically developed marker map of BTA6. Anim Genet 1999; 30:333–9.
https://doi.org/10.1046/j.1365-2052.1999.00487.x


22. Liu XF, Ding XB, Li X, et al. An atlas and analysis of bovine skeletal muscle long noncoding RNAs. Anim Genet 2017; 48:278–86.
https://doi.org/10.1111/age.12539


23. Ulitsky I. Evolution to the rescue: using comparative genomics to understand long non-coding RNAs. Nat Rev Genet 2016; 17:601–14.
https://doi.org/10.1038/nrg.2016.85



24. Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet 2009; 10:155–9.
https://doi.org/10.1038/nrg2521



25. Rinn JL, Kertesz M, Wang JK, et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007; 129:1311–23.
https://doi.org/10.1016/j.cell.2007.05.022



26. Fan C, Hao Z, Yan J, Li G. Genome-wide identification and functional analysis of lincRNAs acting as miRNA targets or decoys in maize. BMC Genomics 2015; 16:793
https://doi.org/10.1186/s12864-015-2024-0



27. Gong Z, Yang Q, Zeng Z, et al. An integrative transcriptomic analysis reveals p53 regulated miRNA, mRNA, and lncRNA networks in nasopharyngeal carcinoma. Tumour Biol 2016; 37:3683–95.
https://doi.org/10.1007/s13277-015-4156-x



28. Ambros V. The functions of animal microRNAs. Nature 2004; 431:350–5.
https://doi.org/10.1038/nature02871



29. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116:281–97.
https://doi.org/10.1016/S0092-8674(04)00045-5


- TOOLS
-
METRICS

-
- 0 Crossref
- 0 Scopus
- 4,516 View
- 110 Download
- Related articles
-
Comparative study of fiber characteristics of rice and wheat straws1996 February;9(1)






 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Download Citation
Download Citation Supplement
Supplement Print
Print




