 |
 |
|
|
Abstract
Taoyuan pig is a native Taiwan breed. According to the historical record, the breed was first introduced to Taiwan from Guangdong province, Southern China, around 1877. The breed played an important role in Taiwan’s early swine industry. It was classified as an indigenous breed in 1986. After 1987, a conserved population of Taoyuan pig was collected and reared in isolation. In this study, mitochondrial DNA sequences and 18 microsatellite markers were used to investigate maternal lineage and genetic diversity within the Taoyuan pig population. Population differentiation among Taoyuan, Asian type, and European type pig breeds was also evaluated using differentiation indices. Only one D-loop haplotype of the Taoyuan pig was found. It clustered with Lower Changjiang River Basin and Central China Type pig breeds. Based on the polymorphism of microsatellite markers, a positive fixation index value (FIS) indicates that the conserved Taoyuan population suffers from inbreeding. In addition, high FST values (>0.2105) were obtained, revealing high differentiation among these breeds. Non-metric multi-dimensional scaling showed a clear geometric structure among 7 breeds. Together these results indicate that maternally Taoyuan pig originated in the Lower Changjiang River Basin and Central China; however, since being introduced to Taiwan differentiation has occurred. In addition, Taoyuan pig has lost genetic diversity in both its mitochondrial and nuclear genomes.
China possesses the largest number of pig breeds in the world. It has been suggested that many Asian domestic pig breeds originated in China (Cheng, 1986; Larson et al., 2005). Based on evolutionary origin, geographic distribution, phenotypes, coat color, and production performance, Chinese indigenous pig breeds have been classified into six types, including: i) North China Type; ii) Lower Changjiang River Basin Type; iii) Central China Type; iv) South China Type; v) Southwest Type; and vi) Plateau Type (Cheng, 1986; Yang et al., 2003). Taiwan is located off the south-east coast of China, and its domestic pig breeds most likely arrived with human migration from mainland China to Taiwan.
In 1986, Taoyuan pig was well documented and recognized as an indigenous Taiwan breed (Cheng, 1986). Based on the historical record, Taoyuan pig possibly arrived in Taiwan from Guangdong province, Southern China, between 1877 and 1887 (Chyr et al., 2001; Zheng and Zhu, 2013). Its phenotypic characteristics, however, are similar to Lower Changjiang River Basin Type breeds (especially Taihu pig breeds, such as Meishan pig) and Central China Type pig breeds. These breeds are characterized by a docile nature, black coats, broad flat faces, large floppy ears, and extensive wrinkling over their entire bodies, and prolific traits (Supplementary Figure S1; Chyr et al., 2001). Typically, domestic pigs from the provinces of Guangdong and Fujian in south-eastern coastal China have smallish ears and fewer wrinkles than Lower Changjiang River Basin Type pigs (Cheng, 1986). At present, the genetic data supporting the historical record that Taoyuan pig was transported from Guangdong province is sparse. However, the breed may have arrived from Guangdong province after internal migration.
Because of its prolific performance, Taoyuan breed played an important role in Taiwan’s early swine industry. The population spread throughout Taiwan until Japanese occupation in 1895. Between 1895 and 1945, the Japanese introduced the Berkshire breed and crossed it with Taoyuan pig to increase pork productivity and heterosis (Chyr et al., 2001; Zheng and Zhu, 2013). After 1958, several other exotic high productivity pig breeds such as Yorkshire, Hampshire, Duroc, and Landrace were also introduced into Taiwan. From this period on, the Taoyuan breed was at risk of extinction. To help conserve the breed, founder populations were collected. These included: 2 males and 21 females in 1974, 5 males and 35 females in 1975, 12 young males in 1984, and 2 males and 11 females in 1986. The staggered collection process was designed to avoid inbreeding. Collections occurred in Taoyuan County and northwestern areas of Taiwan (Chyr et al., 2001). The population was reared in isolation from 1987 through natural mating. However, due to inconsistent breeding policies, the size of the population decreased over the years to its current 10 males and 30 females. The reproductive performances of the conserved Taoyuan pig population are as follows: at birth, litter sizes were 10.6±2.8; after accounting for piglets born alive, they were 8.0±2.8, and after 21 days, 6.4±2.8 (Chyr et al., 2001). The purpose of this study is to clarify genetic diversity in the conserved population and understand the degree of genetic differentiation among Taoyuan, Asian type, and European type pig breeds to better understand the origins of the Taoyuan breed.
Two molecular markers are commonly used to study phylogenetic relationships among different pig breeds. These are the mitochondrial genome and nuclear genome. Each is used to supply different information. Polymorphism of the D-loop (control region) DNA sequence in the mitochondrial genome gives insight into maternal genetic lineages among species. This is based on high deoxynucleotide substitution rates and rare recombination (Cummins, 2001; Gongora et al., 2004). On the other hand, microsatellite DNA markers of the nuclear genome give genetic lineages among individuals and show population diversity (Li et al., 2004). Both molecular markers are used in this study to understand: i) maternal lineages among Taoyuan pigs and Asian type pig breeds; ii) genetic diversity within the conserved Taoyuan pig population; and iii) population differentiation among Taoyuan, Asian type, and European type pig breeds in Taiwan.
Blood samples of conserved Taoyuan pigs were collected from 3 pigs at the Kaohsiung propagation station, and 30 pigs from the Taiwan Livestock Research Institute (TLRI). The Lanyu pig is an indigenous miniature pig breed with a black coat. Previous research has shown remote genetic distance through mitochondrial DNA (mtDNA) between the Lanyu and Taoyuan breeds (Wu et al., 2007). Blood samples from 44 head of conserved Lanyu pigs were also collected, 5 from National Taiwan University and 39 from Taitung Animal Propagation Station. Further, exotic pig blood samples from 35 Meishan, 32 Landrace, 31 Yorkshire, 32 Duroc, and 30 Berkshire animals were obtained from the TLRI. Purified genomic DNA was extracted from collected blood samples with a QIAamp DNA Blood Maxi kit (Qiagen, Valencia, CA, USA) while mtDNA was extracted from samples and purified using Qiagen’s QIAamp DNA mini kit (Qiagen, USA). All sampling protocols were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) of National Taiwan University, and followed the Guide for the Care and Use of Laboratory Animals and the guidelines of the Animal Welfare Act.
Entire sequences of the mtDNA control region were amplified by polymerase chain reaction (PCR) in a MJ thermal cycler using the following primers: L1, 5′-CCAAGACTCAAGGAAGGAGA-3′ (sequence of position 16542–16561 of pig mtDNA, GenBank accession number AF034253) and H1, 5′-GGCGCGGATACTTGCATGTG-3′ (position 1290–1309). Thermal cycling was conducted in 50 μL volumes using the FastStart High Fidelity PCR system (Roche, Penzberg, Germany). Each contained 1 ng of mtDNA, 10 mM Tris-HCl pH 8.3, 1.8 mM MgCl2, 0.4 μM of each primer, 200 μM of each dNTP, and 2 units of FastStart polymerase. Thermal cycling parameters were as follows: 95°C for 5 min; 30 cycles of 95°C for 30 s, 60°C for 30 s, and 72°C for 80 s. At the end, an extension at 72°C for 4 min was given. Complete control regions were sequenced in both directions using the following primers: L1; H1; L2, 5′-CCTATGTACGTCGTGCATTA-3′ (position 160–179); L3, 5′-TACTTCAGGACCATCTCACC-3′ (position 434–453); H2, 5′-AGTGTA AGTTAGGCTTATTG-3′ (position 963–982); and H3, 5′-TTGTGGTAGATTGGCGTAAA-3′ (position 1072–1091). All sequences were determined using an Applied Biosystems 3730 DNA sequencer and analyzed with SeqEd software (Perkin-Elmer, Waltham, MA, USA). Full sequences of the control region were generated by overlapping forward and reverse sequences with EditSeq software (DNASTAR Inc., Madison, WI, USA; Hein and Støvlbæk, 1996). The complete D-loop sequences used in this study of Type I Lanyu (EF375877), Type II Lanyu (DQ972936), Taoyuan, Meishan, Yorkshire, Berkshire, Duroc, and Landrace (GQ169775-GQ169780) have been deposited into the NCBI GenBank.
The following reference mtDNA control region sequences were obtained from NCBI GenBank: 1 Ryukyu wild boar (AB015087), 1 Japanese wild boar (AB015085), 1 Hainan Wuzhishan (EF590146), 1 Zhejiang Jinhua (EF590190), 1 Guangdong Lantang (EF590161), 1 Hainan Lingao (EF590159), 1 Guangxi Bama (EF590178), 1 Fujian Huai (EF590143), 1 Satsuma (AB015091), 1 Guizhou Xiang (EF590172), 1 Sichuan Chenghua (EF590175), 1 Guizhou Guanling (EF590169), 1 Shaanxi Bamei (EF590165), 1 Hunan Daweizi (EF590176), 1 Jiangsu Jiangquhai (EF590195), 1 Anhui Wei (EF590193), 1 Hainan Tunchang (EF590148), 1 Hubei Jianli (EF590163), 1 Okinawa native pig (AB015092), 1 Anhui Wanan (AF276924), 1 Anhui Wanhua (AF276932), 1 Anhui Wannanhua (AF276925), 1 Jiangsu Shawutou (EF590153), 1 Zhejiang Jiashan (EF590186), 1 Jiangsu Dongchuan (EF590171), 1 Hubei Tongcheng (EF590149), 1 Zhejiang Chalu (EF590188), 1 Jiangsu Meishan (D17739), 1 Jiangxi Leping (EF590141), 4 Landrace (AM040613-AM040616), 6 Duroc (AM040623-AM040628), 1 Hampshire (AY574046), and 1 Italian wild boar (AB015094). Chinese indigenous pig classification followed the criteria of Cheng (1986) and Yang et al. (2003). Twenty six Chinese pig breeds are categorized into 5 Chinese pig breed types: i) North China Type; ii) Lower Changjiang River Basin Type; iii) Central China Type; iv) South China Type; and v) Southwest Type. Note that previous research has shown that Tibetan pig belongs to the Plateau Type, domesticated in isolation on the Tibetan highlands and not in the middle and downstream regions of the Yangtze River. This is based on mtDNA polymorphism (Yang et al., 2011). Given this knowledge of independent domestication, Plateau Type pigs were excluded from this study.
Eighteen microsatellite paired primers randomly located on fifteen chromosomes, as recommended by the Domestic Animal Diversity Information System of the Food and Agriculture Organization of the United Nations (ISAG-FAO), were chosen for genotyping (FAO, 2004). The primers used for genomic DNA amplification were fluorescent end-labeled with 6-carboxyfluorescein, hexachlorofluorescein, or carboxytetramethylrhodamine fluorescent dye (Supplementary Table S1). The fragments of microsatellite DNA in the genome were amplified by PCR in 15 μL reaction volumes with 50 ng of genomic DNA, 1.5 μL of 10× PCR buffer, 0.375 μL of 8 mM dNTP, 4.5 pmole sense and antisense primers, and 0.6 units of DNA Taq polymerase (Amersham Biosciences, Piscataway, NJ, USA). The thermal cycling conditions were: an initial 95°C for 5 min, then 37 cycles at 95°C for 30 s, 48°C to 58°C for 30 s (depending on the locus), and 72°C for 45 s with a final extension at 72°C for 7 min. PCR products were resolved by electrophoresis and sequenced with a MegaBACE 1000 DNA sequencer (Amersham Biosciences, USA). The fluorescent-labeled marker ET-400 (Amersham Biosciences, USA) was used as an internal size standard for length calibration. Allele sizes were determined using Genetic-Profiler software version 2.2 (Amersham Biosciences, USA).
In the analysis of pairwise distance of control regions, the tandem repeat motif ‘CGTGCGTACA’ with a variable number of repeats in individuals, and Type I and II Lanyu-specific repeated motifs (ACACAAACC and TAAAACACTTA, respectively) of the mtDNA control region were excluded from analysis (Wu et al., 2007). Sequence alignment of obtained sequences was performed using MegAlign multiple alignment software (DNASTAR Inc., USA; Hein and Støvlbæk, 1996). Haplotype and nucleotide diversities within the conserved Taoyuan pigs, Lanyu pigs, and exotic pigs were obtained with DNA sequence polymorphism (DnaSP) software version 4.20.2 (Rozas et al., 2003). The PHYLIP program package version 3.66 was used to generate the neighbor-joining (NJ) and maximum likelihood (ML) phylogenetic tree (Felsenstein, 1989). For ML analysis, MODELTEST version 3.7 was used to determine the best-fit model for the data, including nucleotide composition, substitution matrix among nucleotides, and proportion of invariant sites (Posada and Crandall, 1998). Nodal supports of the NJ tree and ML trees were evaluated by bootstrap resampling (1,000 replications) using the PHYLIP program package (Felsenstein, 1989).
The genetic distances within the 33 conserved Taoyuan pig individuals and 203 individuals of 6 exotic breeds were determined by MSA software (Dieringer and Schlötterer, 2003). Allele frequencies at each locus, expected heterozygosity (HE), observed heterozygosity (HO), and polymorphic information content (PIC) in conserved Taoyuan pigs were calculated using CERVUS version 2.0 (Marshall et al., 1998). The GENEPOP software package version 3.4b was used to calculate allele frequencies, Wright’s F-statistic (FIS), and test for deviation from the Hardy-Weinberg equilibrium (Raymond and Rousset, 1995). FIS was used to determine the reduction in heterozygosity among individuals within the conserved Taoyuan pig population. The fixation indices (FST) as determined by MSA were used to determine genetic diversity among the 7 breeds. The effective number of alleles was estimated according to Kimura and Crow’s formula (Kimura and Crow, 1964).
The geometric picture among the individuals from 7 pig breeds based on the proportion of shared alleles (POSA) to establish genetic distances was done with non-metric multi-dimensional scaling (MDS) technique using Polymouth Routines in Multivariate Ecological Research (PRIMER) program package (Carr, 1996). The distance Aij between populations i and j in the MDS ordination plot is calculated as
A i j = [ ( X 1 i - X 1 j ) 2 + ( X 2 i - X 2 j ) 2 ] S T R E S S = ∑ i < j ( D i j - A i j ) 2 ∑ i < j D i j 2
To understand the genetic lineages of mtDNA D-loop between Taoyuan pigs and Asian type breeds, mitochondrial genomes of 33 head of conserved Taoyuan pigs were purified from blood samples and mtDNA D-loop regions were sequenced. Asian and European type pigs’ D-loop sequences, including 25 Chinese pigs, 4 Japanese pigs, 11 European pigs and 1 Italian wild pig were obtained from NCBI Genbank. In the present study, only 1 mtDNA D-loop haplotype was obtained from conserved populations of Taoyuan pigs, indicating a loss of mtDNA heterozygosity in conserved Taoyuan pigs. All of the obtained D-loop (1,044 bp) sequences were aligned by MegAlign multiple alignment software, and were defined as 22 haplotypes. The D-loop sequence of Taoyuan pig is identical to some of the Lower Changjiang River Basin Type breeds (located in eastern China), including the Jiangsu Jiangquhai, Anhui Wei, Hubei Jianli, and Anhui Wanan pig breeds, and to Central China Type pigs, including the Hunan Daweizi pig breed (Figure 1). The mtDNA D-loop sequences from the Hainan Tunchang pig (southern China) and Okinawa native pig (Okinawa, Japan) are also identical to the Taoyuan pig sequence. In the present study, the mtDNA control region was found to have 51 nucleotide polymorphic sites including 44 transitions, 5 tranversions, and 2 deletions in the above haplotype sequences. No unique nucleotide substitution in the Taoyuan pig D-loop sequence was identified (Figure 1).
The NJ and ML phylogenetic trees based on the polymorphic sequences of the D-loop region were constructed to analyze genetic lineage between Taoyuan pig, Asian, and European Type pig breeds. The NJ tree (Figure 2) and ML tree (Supplementary Figure S2) possessed a similar tree topology. The European type pigs (except Yorkshire* and Berkshire* breeds, whereby the asterisk [*] represents pig breeds reared in Taiwan) and Asian type breeds formed two major clades (Figure 2). The Asian type breeds were then divided into two subclades (SUB I and SUB II), not including: Type I Lanyu*, Ryukyu wild boar, Japanese wild boar, Hainan Wuzhishan and Yorkshire* pigs. The SUB I clade contained 12 breeds: 1 North China Type, 1 Central China Type, 5 South China Type, 2 Southwest Type pig breeds, 1 indigenous Taiwan breed (Type II Lanyu pig), 1 Japanese breed (Satsuma pig), and 1 Meishan*. The D-loop sequence of Meishan* was totally identical to Zhejiang Jinhua. The SUB II clade was separated into subclades SUBII-I and SUBII-II. Nine breeds were classified into SUBII-I clade, including Taoyuan* (1 haplotype), 5 Lower Changjiang River Basin Type breeds, 1 Central China Type breed, 1 South China Type breed (Hainan Tunchang), and 1 Japanese native breed (Okinawa native pig). The SUB II-II clade contained 9 pig breeds: 5 Lower Changjiang River Basin Type breeds, 3 Central China Type breeds, and 1 Berkshire* breed. The majority of breeds classified in the SUBII clade are Lower Changjiang River Basin Type breeds (Figure 2).
Only 1 haplotype was identified in the mtDNA D-loop sequences from the 33 conserved Taoyuan pigs. To understand whether their nuclear DNA suffered from a loss of heterozygosity similar to their mtDNA, 18 microsatellite markers were used to determine nuclear genetic diversity within Taoyuan pig. In total, 62 alleles were identified with an average allele number for 18 loci of 3.44. The mean PIC was 0.445, ranging from 0.248 to 0.682, indicating the 18 markers were informative of conserved Taoyuan pigs (Table 1). The correlation between the PIC value and number of alleles per locus was analyzed, and the correlation coefficient was 0.748 (p<0.01). Mean HO (0.467) in the Taoyuan population was less than HE (0.516). In a total of 18 microsatellite markers, the mean FIS value was positive (+0.096) and FIS values of eight microsatellite markers were higher than 0. Nevertheless, three markers (S0005, S0228, and S0386) showed significant deviation from Hardy-Weinberg equilibrium (p<0.01; Table 1). These data indicate that the nuclear genome of conserved Taoyuan pigs has suffered from inbreeding and gene drift.
The maternal ancestor of Taoyuan pig originated from the Lower Changjiang River Basin and Central China. However, the Taoyuan pig has been isolated on Taiwan for a long time and its conserved population is small. Potentially, isolation effect and a small population enhance genetic differentiation. To test this possibility, genetic differentiation among conserved Taoyuan, Asian type, and European type pig breeds reared in Taiwan was evaluated. FST values based on pairwise distances according to the polymorphism of microsatellite markers from 7 breeds were estimated by the MSA program. High differential FST values (>0.2105) were detected between breeds (Table 2). The FST values among European breeds (Landrace, Yorkshire, Duroc, and Berkshire, ranging from 0.2105 to 0.3730) were less than those between Asian breeds (Taoyuan, Meishan, and Lanyu, ranging from 0.3889 to 0.4215). The FST value between Taoyuan and Lanyu is 0.3957, and between Taoyuan and Meishan is 0.4215. Very high genetic differentiation was identified between Taoyuan and Berkshire breeds (0.5142). The result indicates genetic differentiation among Taoyuan, Meishan, Lanyu, and exotic pig breeds. To demonstrate clear geometric structure among individuals of conserved Taoyuan, Lanyu, and other exotic pig breeds, POSA genetic distances based on 18 microsatellite loci of the above 7 pig breeds were established with a non-metric MDS technique using the PRIMER5 program package. The analysis shows that individuals of the Taoyuan pig breed could be clearly assigned into a distinctive group (Table 2; Figure 3). The matrix distances among European breeds were closer than between Asian breeds (Figure 3). The STRESS value was less than 0.18 after 10 restarts.
In Taiwan, Taoyuan pig was recorded apocryphally to have been introduced from Guangdong province of China to Taiwan by emigrants in the period 1877 to 1887 (Chyr et al., 2001; Zheng and Zhu, 2013). Ambiguously, Taoyuan pig was classified as a Lower Changjiang River Basin Type pig breed in the Book of ‘Pig Breeds in China’ published by Shanghai Scientific and Technical Publishers, China (Cheng, 1986). No genetic study has been undertaken to clarify this apparent contradiction until this present study. Here we showed that the D-loop sequence of Taoyuan pig clustered together with the Lower Changjiang River Basin and Central China Type breeds. Further, the D-loop sequence of Taoyuan pig was identical to some of Lower Changjiang River Basin and Central China Type breeds, including the Hunan Daweizi, Jiangsu Jiangquhai, Anhui Wei, Hubei Jianli, and Anhui Wanna breeds. The pig breeds from Guangdong, Guangxi, and Fujian provinces (South China Type breeds), including the Guangdong Lantang, Hainan Lingao, Guangxi Bama, and Fujian Huai clustered in another clade (SUB I clade, Figure 2). The Taoyuan pig did not cluster with any pig breed from Fujian and Guangdong provinces, for example Fujian Huai and Guangdong Lantang breeds. Furthermore, cytosine at position 278 and thymidine at position 451 were identified in most of the D-loop sequences from South China and Southwest China Type pig breeds, while thymidine at position 278 and cytosine at position 451 were identified in most of the Lower Changjiang River Basin and Central China Type pig breeds (Figure 1; SUB II clade, Figure 2). Remote genetic distances of mtDNAs were determined between the Taoyuan and an indigenous breed, the Lanyu pig. This result supports the assignment of the Taoyuan pig to Lower Changjiang River Basin and Central China Type pigs. Because no pig from Guangdong and Fujian provinces clustered with Taoyuan pig, our data did not support the stories that the Taoyuan pig originated from Guangdong province (Chyr et al., 2001); however, we could not rule out the possibility that the ancestor of Taoyuan pig may have been transported from Lower Changjiang River Basin and Central China through Guangdong province rather than being transported into Taiwan directly. The sequential order of domestication and breeding history of Lower Changjiang River Basin and Central China Type pigs still needs clarification. This could give greater understanding of human mediated Taoyuan pig dispersal. The low bootstrap values determined in the present study can be attributed to lower genetic distance among the haplotypes chosen. This is influenced by similar sequence characterization and a large number of haplotypes (Wróbel, 2008).
The control region haplotype of the Taoyuan pig breed was found to be identical to the Okinawa native pig breed. The study of morphological characteristics and the phylogenetic relationship of S. scrofa bone remains obtained from archaeological sites on the Okinawa islands suggests that the Okinawa native pig breed might have already existed on Okinawa some 1,700 to 2,000 years ago (Matsui, 1997; Watanobe et al., 2002). Evidence exists that the Okinawa native pig breed may have been introduced from Kyushu island (Japan) and mainland Asia to the Okinawa islands through trade (Watanobe et al., 2002). This is based on: i) analyzed morphological changes of prehistoric S. scrofa on mainland Japan, suggesting that domestic pigs were introduced from the Asian continent to mainland Japan during the Yayoi period (Nishimoto, 1993); ii) mainland Japan Yayoi pottery shards, shell artifacts and coins from China found in archaeological sites for the Yayoi-Heian period on the Okinawa islands; and iii) a lack of any artifacts from Taiwan identified in archaeological sites in the Okinawa islands. This paper, however, provides important evidence supporting the possibility of gene flow between Okinawa native pig and Taoyuan pig. This is reflected in genetic information that links control region sequences. It is possible that the prehistorical Okinawa native pig was introduced from Taiwan as geographically Taiwan and Okinawa are close. More archaeological information from sites in Taiwan and Okinawa is needed to clarify the route and timing of the Taoyuan pig’s introduction onto Taiwan and the direction of any gene flow between Taiwan and Okinawa.
The Tunchang pig of Hainan Island in Southern China belongs to a group of small eared miniature pigs. Its coat is black from its head down the top of its back to its rump while its body is predominantly white. This is in contrast to Taoyuan pig, which is solid black with large floppy ears. According to polymorphism of microsatellite markers, the Tunchang pig is genetically clustered with the Wuzhishan miniature pig, which also originates from Hainan Island (Wang et al., 2006). In this paper, however, both showed different mtDNA control region haplotypes, indicating their mitochondrial lineages have different origins (Figure 2). Because no other pig’s control region haplotype except Tunchang pig of Hainan Island was identical to the Taoyuan pig, or Lower Changjiang River Basin and Central China Type breeds, these data suggest that the control region haplotype of Tunchang pig originated in the Lower Changjiang River Basin and Central China Type pig breeds via genetic introgression.
Only 1 haplotype of mtDNA D-loop was identified in the conserved Taoyuan pig population. In addition, the mean FIS value of 18 microsatellite markers for their polymorphism in nuclear DNA was positive (>0.096), with 8 microsatellite marker FIS values being higher than 0. Three microsatellite markers showed significant deviation from the Hardy-Weinberg equilibrium (p<0.01). These data indicate the conserved Taoyuan pig population has suffered a severe loss of heterozygosity in both mitochondrial and nuclear genomes, and suggest that the population is suffering from inbreeding. The relatively smaller litter size of the conserved Taoyuan pig population might be due to a loss of genetic diversity from inbreeding. The founder population’s collection history for conserved Taoyuan pigs shows that 21 males and 67 females were collected before 1986. In theory, the founder population should possess enough genetic variation to prevent a severe loss of heterozygosity; however, a breeding scheme reliant on natural mating has possibly led to a major loss of genetic diversity. Unfortunately, the conserved population is now very small (only 10 males and 30 females) and a genetic quality control and population management scheme is urgently needed to maintain genetic diversity.
Previous studies showed no gene introgression between Taiwan’s two indigenous pigs — the conserved Taoyuan and Lanyu pig breeds. This was proved by constructing a neighbor-joining tree based on the -ln(POSA) distance for 240 individuals from 7 breeds (Jiang et al., 2008; Chang et al., 2009). Fixation index (FST) and principal component factor score analyses were conducted to understand genetic structure and differentiation among Taoyuan, Asian type, and European type pig breeds. High FST values were detected between some breeds. Relatively, lower FST values (0.2105 to 0.3730) were seen between European breeds (Landrace, Yorkshire, Duroc, and Berkshire breeds) compared to Asian breeds (Taoyuan, Meishan, and Lanyu breeds). Further, our data show the genetic distance of Taoyuan pig mtDNA is close to Lower Changjiang River Basin Type breed; nevertheless, polymorphism of microsatellite markers gives a high FST (0.4215) value between Taoyuan and Meishan (Lower Changjiang River Basin Type breed) pigs.
Records clearly show the Berkshire breed was introduced from Japan onto Taiwan and crossed with the Taoyuan pig to improve productivity during Japanese occupation of Taiwan. Therefore, before conservation in 1987, the Taoyuan pig ancestor genome likely experienced genetic introgression from the Berkshire breed. In this paper, a high differentiation index (FST = 0.5142) between the Taoyuan and Berkshire breeds was discovered. Additionally, two-dimensional MDS plotting, showing geometric structure among individuals of each breed, clusters conserved Taoyuan pig in a group closer to the Meishan population than the Berkshire population. To understand any lack of agreement in the statistical results of MDS plotting, the STRESS value was calculated. After 10 restarts, a value of 0.18 was achieved. Since STRESS values of <0.2 indicate a potentially useful picture of the situation, this MDS plot has some analytical merit (Bond et al., 2002). We think there are 3 good potential reasons for genetic differentiation among Taoyuan, Meishan, and Berkshire pig breeds: i) founder population effect; ii) natural mating causing significant gene drift; and iii) a lack of an effective breeding program to maintain genetic diversity and enhance genetic differentiation.
This work used the mitochondrial and nuclear genomes of a conserved population of Taoyuan pig (an indigenous Taiwan breed) and compared it with that of other Asian and European breeds to trace Taoyuan pig’s introduction to Taiwan from Southern China. The data do not directly support the historical record that the Taoyuan pig was introduced to Taiwan from Guangdong province around 1887. The mtDNA data show that the Taoyuan pig is maternally related to the Lower Changjiang River Basin and Central China Type pig breeds. It is possible, however, that the Taoyuan pig arrived on Taiwan from Guangdong province through migration within Southern China and then onto Taiwan. The study also indicates that the conserved Taoyuan pig population is suffering from inbreeding and a consequent loss of genetic diversity in both mitochondrial and nuclear genomes. This likely relates to breeding practices, and founder population effect. On the other hand, the conserved breed shows high specificity and is differentiated from other breeds by isolation. It is evident from this study that an effective breeding program for population management and genetic quality is needed to maintain genetic diversity among the Taoyuan breed.
ACKNOWLEDGMENTS
This work was supported by a grant from the National Science Council of Taiwan, Republic of China (NSC 102-2313-B-002-024-MY3), Council of Agriculture of Taiwan, Republic of China (103AS-2.1.1-AD-U1[8]), Chiang Ching-Kuo Foundation for International Scholarly Exchange (RG008-D-07), and Yushan National Park (98-1185). We thank Daniel Flynn for editing the manuscript.
Figure 1
Nucleotide substitution sites of control region sequences in Taoyuan pig and 40 Asian and European type pigs. Sequence position numbers (given in the first row) follow those in the Type I Lanyu sequence (accession number EF375877). Only variable sites in the mtDNA control region of these animals, with sequence positions given above, are shown. Abbreviations in the leftmost column indicate the geographical origins of these animals: T, Taiwan; J, Japan; I, North China Type; II, Lower Changjiang River Basin Type; III, Central China Type; IV, South China Type; V, Southwest Type; E, Europe. The former abbreviations in taxa’ names indicate provinces: HN, Hainan; ZJ, Zhejiang; GD, Guangdong; GX, Guangxi; FJ, Fujian; GZ, Guizhou; SC, Sichuan; SX, Shaanxi; HU, Hunan; JS, Jiangsu; AH, Anhui; HB, Hubei, and JX, Jiangxi. The nucleotides identical to the mtDNA control region consensus sequence are denoted by a dash (-). Dot signs (.) indicate gaps in or compared to the consensus sequence. Asterisk signs (*) followed by taxa name represent the pig breeds reared in Taiwan.

Figure 2
Neighbor-joining phylogenetic tree based on the Taoyuan, Asian, and European pig control region sequences. Right brackets indicate the sequences clustered in every clade. Numbers on the branches are bootstrap values based on bootstrap resampling (1,000 replications). Only values higher than 50% are shown. The numerals and the nucleotide abbreviations in the center of horizontal lines indicate nucleotide substitution sites of control region sequences and the characteristics of substitution nucleotide in every clade. The abbreviations of geographical origins and located provinces are identical to Figure 1. The asterisk (*) represents pig breeds reared in Taiwan.

Figure 3
Scatter diagram showing relative position of 7 pig breeds based on the genetic distance of the 18 microsatellite loci by non-metric multi-dimensional scaling (MDS) technique. Solid circles (●) represent Taoyuan pigs; solid triangles (▲) represent Meishan pigs; open triangles (△) represent Lanyu pigs; inverted solid triangles (▼) represent Landrace pigs; inverted open triangles (▽) represent Yorkshire pigs; solid squares (■) represent Duroc pigs; open squares (□) represent Berkshire pigs. Stress (Standardized residual sum of squares) value is 0.18.
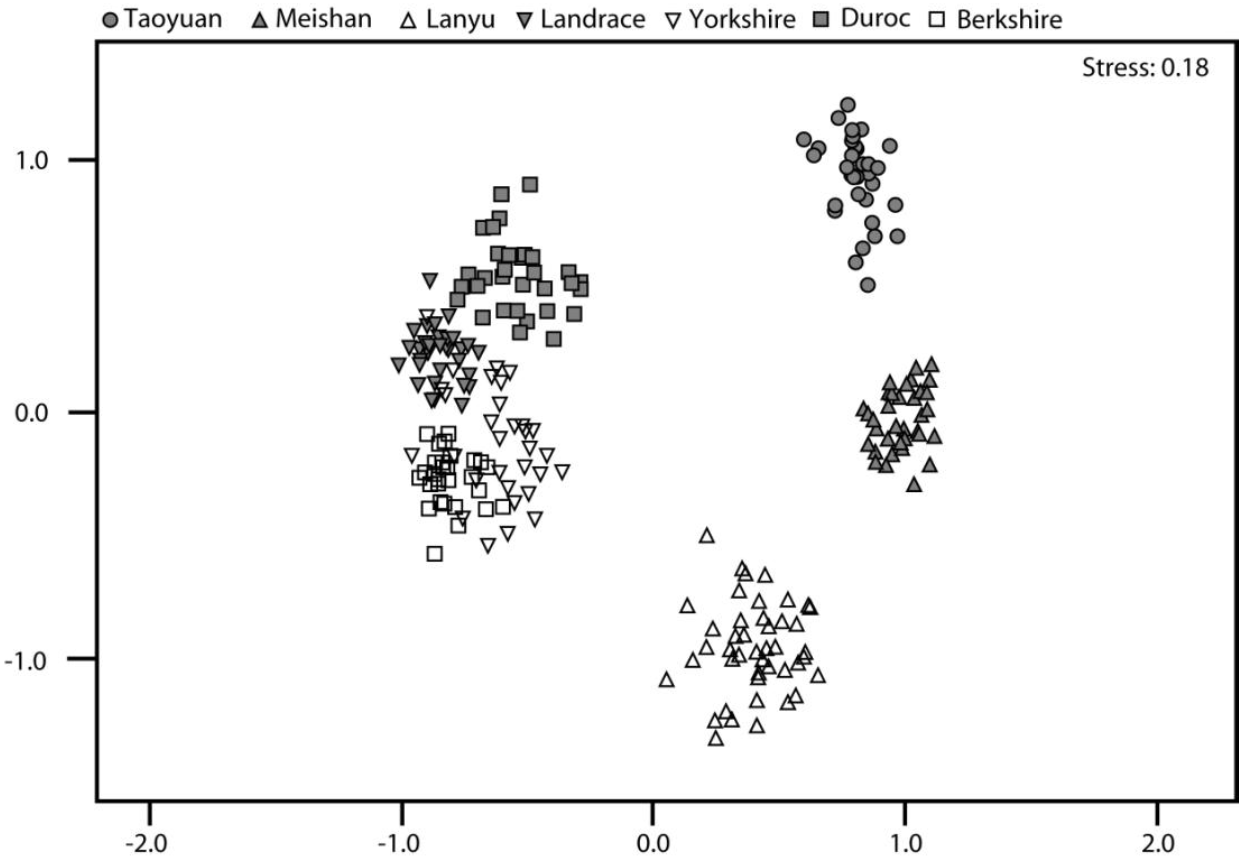
Table 1
Summary statistics for number of alleles (NA), effective number of alleles (NE), number of heterozygotes (Hets) and homozygotes (Homs), observed (HO), and expected (HE) heterozygosities, polymorphism information content (PIC), tests for deviation from Hardy-Weinberg equilibrium (HW) and fixation indices (FIS) of 18 microsatellite loci in conserved Taoyuan pigs
| Locus | NA | NE | Hets | Homs | HO | HE | PIC | HW | FIS |
|---|---|---|---|---|---|---|---|---|---|
| SW857 | 5 | 2.900 | 22 | 11 | 0.667 | 0.665 | 0.594 | NS | 0.094 |
| IGF1 | 4 | 1.784 | 17 | 16 | 0.515 | 0.446 | 0.367 | NS | −0.065 |
| S0155 | 2 | 1.619 | 9 | 24 | 0.273 | 0.388 | 0.309 | NS | 0.307 |
| S0005 | 6 | 3.290 | 11 | 22 | 0.333 | 0.707 | 0.655 | ** | 0.383 |
| SW911 | 3 | 2.248 | 17 | 16 | 0.515 | 0.564 | 0.455 | NA | 0.032 |
| S0068 | 3 | 1.364 | 10 | 23 | 0.303 | 0.271 | 0.248 | NA | −0.075 |
| S0228 | 4 | 1.811 | 3 | 26 | 0.103 | 0.456 | 0.395 | ** | 0.933 |
| SW24 | 4 | 3.718 | 29 | 2 | 0.935 | 0.743 | 0.682 | NS | −0.222 |
| S0227 | 3 | 2.396 | 22 | 11 | 0.667 | 0.592 | 0.503 | NS | −0.060 |
| SW72 | 4 | 3.175 | 24 | 9 | 0.727 | 0.696 | 0.624 | NS | 0.069 |
| S0218 | 2 | 1.581 | 8 | 25 | 0.242 | 0.373 | 0.300 | NS | 0.361 |
| S0355 | 2 | 1.766 | 15 | 18 | 0.455 | 0.441 | 0.340 | NS | −0.033 |
| SW122 | 4 | 2.578 | 20 | 13 | 0.606 | 0.621 | 0.531 | NS | −0.014 |
| S0225 | 4 | 1.799 | 17 | 16 | 0.515 | 0.451 | 0.394 | NS | −0.092 |
| S0226 | 3 | 1.929 | 15 | 18 | 0.455 | 0.489 | 0.431 | NS | −0.030 |
| SW951 | 2 | 1.619 | 17 | 16 | 0.515 | 0.388 | 0.309 | NS | −0.337 |
| S0215 | 3 | 1.638 | 16 | 17 | 0.485 | 0.395 | 0.347 | NS | −0.147 |
| S0386 | 4 | 2.478 | 3 | 30 | 0.091 | 0.606 | 0.529 | ** | 0.620 |
| Mean | 3.44 | 2.205 | 15.28 | 17.39 | 0.467 | 0.516 | 0.445 | 0.096 |
Table 2
The genetic differential FST values among conserved Taoyuan, Asian type, and European type pig breeds
REFERENCES
Bond JM, Veenendaal EM, Hornby DD, Gray AJ. 2002. Looking for progenitors: A molecular approach to finding the origins of an invasive weed. Biol Invasions 4:349–357.

Carr MR. 1996. PRIMER User Manual. Ver 4.0. Plymouth Routines in Multivariate Ecological Research. Plymouth Marine Laboratory; Plymouth, UK:
Chang WH, Chu HP, Jiang YN, Li SH, Wang Y, Chen CH, Chen KJ, Lin CY, Ju YT. 2009. Genetic variation and phylogenetics of Lanyu and exotic pig breeds in Taiwan analyzed by nineteen microsatellite markers. J Anim Sci 87:1–8.

Cheng PL. 1986. Description of Chinese pig breeds. Pig Breeds in China. Zhang Z, editorShanghai Scientific and Technical Publishers; Shanghai: p. 25–178.
Chyr SC, Lin KJ, Chang HL, Yang TS, Yen HT. 2001. Breeds and genetic improvement. Guidelines in Animal Science-Swine Production. Chyr SC, editorChinese Society for Animal Science Press; Taipei: p. 29–95.
Cummins J. 2001. Mitochondrial DNA and the Y chromosome: Parallels and paradoxes. Reprod Fertil Dev 13:533–542.


Dieringer D, Schlötterer C. 2003. MICROSATELLITE ANALYSER (MSA): A platform independent analysis tool for large microsatellite data sets. Mol Ecol Notes 3:167–169.

Felsenstein J. 1989. PHYLIP – phylogeny inference package (version 3.2). Cladistics 5:164–166.
Food and Agriculture Organization of the United Nations (FAO). 2004. Secondary Guidelines for Development of National Farm Animal Genetic Resources Management Plans. Measurement of Domestic Animal Diversity (MoDAD): Recommended Microsatellite Markers. FAO; Rome:
Gongora J, Fleming P, Spencer PBS, Mason R, Garkavenko O, Meyer JN, Droegemueller C, Lee JH, Moran C. 2004. Phylogenetic relationships of Australian and New Zealand feral pigs assessed by mitochondrial control region sequence and nuclear GPIP genotype. Mol Phylogenet Evol 33:339–348.


Jiang YN, Wu CY, Huang CY, Chu HP, Ke MW, Kung MS, Li KY, Wang CH, Li SH, Wang Y, Ju YT. 2008. Interpopulation and intrapopulation maternal lineage genetics of the Lanyu pig (Sus scrofa) by analysis of mitochondrial cytochrome b and control region sequences. J Anim Sci 86:2461–2470.


Kimura M, Crow JF. 1964. The number of alleles that can be maintained in a finite population. Genetics 49:725–738.




Krzanowski WJ. 1988. Principles of Multivariate Analysis: A User’s Perspective. Oxford University Press; Oxford:
Larson G, Dobney K, Albarella U, Fang M, Matisoo-Smith E, Robins J, Lowden S, Finlayson H, Brand T, Willerslev E, Rowley-Conwy P, Andersson L, Cooper A. 2005. Worldwide phylogeography of wild boar reveals multiple centers of pig domestication. Science 307:1618–1621.


Li SJ, Yang SH, Zhao SH, Fan B, Yu M, Wang HS, Li MH, Liu B, Xiong TA, Li K. 2004. Genetic diversity analyses of 10 indigenous Chinese pig populations based on 20 microsatellites. J Anim Sci 82:368–374.


Marshall TC, Slate J, Kruuk LEB, Pemberton JM. 1998. Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7:639–655.


Matsui A. 1997. Animal remains in Gushibaru shellmidden. Report on Cultural Assets of Okinawa Prefecture. 130:Okinawa Prefectural Board of Education. Okinawa Prefectural Board of Education; Naha, Japan: p. 159–187.
Nishimoto T. 1993. The physical character of the pig in the Yayoi Period. B Natl Mus Jpn Hist 50:49–70.
Posada D, Crandall KA. 1998. MODELTEST: Testing the model of DNA substitution. Bioinformatics 14:817–818.


Raymond M, Rousset F. 1995. GENEPOP (version 1.2): Population genetics software for exact tests and ecumenicism. J Hered 86:248–249.

Rozas J, Sánchez-DelBarrio JC, Messeguer X, Rozas R. 2003. DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497.


Wang X, Ou J, Huang L, Nishihara M, Li J, Manabe N, Zhang Y. 2006. Genetic characteristics of inbred Wuzhishan miniature pigs, a native Chinese breed. J Reprod Develop 52:639–643.

Watanobe T, Ishiguro N, Nakano M, Takamiya H, Matsui A, Hongo H. 2002. Prehistoric introduction of domestic pigs onto the Okinawa Islands: ancient mitochondrial DNA evidence. J Mol Evol 55:222–231.


Wróbel B. 2008. Statistical measures of uncertainty for branches in phylogenetic trees inferred from molecular sequences by using model-based methods. J Appl Genet 49:49–67.


Wu CY, Jiang YN, Chu HP, Li SH, Wang Y, Li YH, Chang Y, Ju YT. 2007. The type I Lanyu pig has a maternal genetic lineage distinct from Asian and European pigs. Anim Genet 38:499–505.


Yang SL, Wang ZG, Liu B, Zhang GX, Zhao SH, Yu M, Fan B, Li MH, Xiong TA, Li K. 2003. Genetic variation and relationships of eighteen Chinese indigenous pig breeds. Genet Sel Evol 35:657–671.



Yang SL, Zhang H, Mao HM, Yan DW, Lu SX, Lian LS, Zhao GY, Yan YL, Deng WD, Shi XW, Han SX, Li S, Wang XJ, Gou X. 2011. The local origin of the Tibetan pig and additional insights into the origin of Asian pigs. PLoS ONE 6:12e28215



Zheng CC, Zhu YT. 2013. On pig breeding culture in Taiwan rural areas during the period of Japanese occupation. Taiwan Res J 125:55–64.





 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Full text via PMC
Full text via PMC Download Citation
Download Citation Supplement
Supplement Print
Print





