Determination of Sperm Sex Ratio in Bovine Semen Using Multiplex Real-time Polymerase Chain Reaction
Article information
Abstract
Gender selection is important in livestock industries; for example, female calves are required in the dairy industry. Sex-sorted semen is commonly used for the production of calves of the desired gender. However, assessment of the sex ratio of the sorted semen is tedious and expensive. In this study, a rapid, cost effective and reliable method for determining the sex ratio was developed using a multiplex real-time polymerase chain reaction (PCR) assay. In this assay, the X and Y chromosome-specific markers, i.e., bovine proteolipid protein (PLP) gene and sex-determining region Y (SRY) were simultaneously quantified in a single tube. The multiplex real-time PCR assay was shown to have high amplification efficiencies (97% to 99%) comparable to the separated-tube simplex real-time PCR assay. The results obtained from both assays were not significantly different (p>0.05). The multiplex assay was validated using reference DNA of known X ratio (10%, 50%, and 90%) as templates. The measured %X in semen samples were the same within 95% confidence intervals as the expected values, i.e., >90% in X-sorted semen, <10% in Y-sorted semen and close to 50% in the unsorted semen. The multiplex real-time PCR assay as shown in this study can thus be used to assess purity of sex-sorted semen.
INTRODUCTION
Sexual pre-selection plays an important economic role in animal production industries. For example, female cattle are required for the dairy industry while males are preferred in the beef cattle industry (Seidel Jr, 2007). Sorting of X (female) or Y (male) chromosome-bearing spermatozoa in semen is a popular method for sex-selection. Several techniques for sperm sorting technology have been developed; for example, based on different immunological properties of the sperm (Hendriksen et al., 1993; Blecher et al., 1999; Sang et al., 2011), different swimming ability (Madrid-Bury et al., 2003), differential separation in percoll (Machado et al., 2009) and differential separation in albumin gradients (Wolf et al., 2008; Machado et al., 2009). Currently, separation of X- and Y- bearing spermatozoa based on differences in their chromosomal content by flow cytometry is considered to be the most reliable and effective method, in which spermatozoa of the desired sex can be obtained in greater than 90% purity (Morrell et al., 1988; Garner and Seidel Jr, 2003). In order to quantify the purity of sorted semen, flow cytometry reanalysis is usually carried out. However, the accuracy of the estimate of purity is compromised by the systematic error inherent in the technique. Therefore, a different technique should be used to confirm the sex ratio of the flow-sorted semen to reduce the error. Multicolor fluorescent in situ hybridization is considered to be highly reliable method for identifying sex of the spermatozoa, and this method can be used to evaluate the sex ratio of sorted semen (Piumi et al., 2001; Rens et al., 2001; Lee et al., 2004; Habermann et al., 2005). However, this technique is complicated, laborious, time-consuming, and requires highly skilled technicians, which limits its use. Polymerase chain reaction (PCR) techniques for detecting DNA sequences on sex chromosomes have been reported which can be used to identify the sex of individual spermatozoa and sex ratios of spermatozoa in semen (Colley et al., 2008; Wang et al., 2011). However, accurate determination of the sex ratio by single-sperm PCR requires analysis of a large number of individual sperm, which is laborious and expensive. In contrast, the sex ratio of semen can be determined accurately and more simply by quantitative real-time PCR (Parati et al., 2006). In this technique, fluorescent labeled-Taqman probes are used to detect DNA sequences on each sex chromosome in two separate reaction tubes, one specific for bovine proteolipid protein (PLP) gene located on the X chromosome and another for sex-determining region Y (SRY) located on the Y chromosome. The result obtained from the real-time PCR was proven to be not significantly different from that obtained by flow cytometry reanalysis. However, since the quantification of both genes was done in separate tubes, imprecision in dispensing a small amount of the DNA template into each reaction tube may compound error in estimation of the sex ratio in the sample. Therefore, in this study, we aimed to develop a multiplex real-time PCR technique to simultaneously quantify both genes in a single tube to eliminate bias due to pipetting error and thus provide better estimation of the sperm sex ratio. Furthermore, combining the reaction into one tube is also more efficient in terms of the reagent used, and time and labor needed to conduct the test.
MATERIALS AND METHODS
Primers and probes
Primer and probe sequences for amplification and detection of the X chromosome-specific (PLP) and Y chromosome-specific (SRY) regions were as previously reported (Parati et al., 2006) (Table 1). The X- and Y-Taqman probes used in the multiplex PCR assay were labeled at the 5′ end with different fluorophores, i.e., CAL Fluor Orange 560 (Orange 560) for the X-specific probe and 6-Carboxyfluorescein (FAM) for the Y-specific probe. For the simplex PCR assay both probes were labeled with the same fluorophore, FAM. BHQ-1 quencher was incorporated at the 3′ end of all probes to permit fluorescent detection of specific product. All primers and probes were synthesized from Biosearch Technologies, Inc., Novato, CA, USA.
Construction of reference DNA
Two recombinant plasmids containing the amplified section of the PLP gene (p-PLP) and SRY sequence (p-SRY) were constructed and used as reference templates for quantification of the sex-related DNA sequences. Each DNA fragment was amplified using primers as described in Table 1. The fragments were then purified and cloned into pDrive cloning vector using a QIAGEN PCR Cloning PlusKit (Qiagen GmbH, Hilden, Germany) according to the manufacturer’s instruction. The recombinant plasmids were extracted using a QIAprep Spin Miniprep Kit (Qiagen GmbH, Germany). Plasmid concentrations were quantified spectrophotometrically at 260 nm using NanoDrop1000 spectrophotometer (Thermo Fisher Scientific, Inc., Wilmington, DE, USA). Both plasmids were mixed at a ratio of 1:1, and ten-fold serial dilutions ranging from 10 pg to 1 fg (approximately 2.33E+06 to 2.33E+02 plasmid copies) were used to construct a standard curve for quantification. Different ratios of p-PLP and p-SRY, i.e., 1:9 (X10), 1:1 (X50), and 9:1 (X90) at a final concentration of 100 fg were used as known ratio templates to validate the method.
Reaction conditions for real-time polymerase chain reaction
The real-time PCR reaction condition was carried out as described by Parati et al. (2006). The 25 μL reaction mixture of both simplex and multiplex real-time PCR contained 20 mM Tris-HCl (pH 8.4), 50 mM KCl, 0.4 mM dNTPs, 6 mM MgCl2 and 0.625 U Platinum Taq DNA Polymerase (Life Technologies Co., Sao Paulo, Brazil). For the simplex PCR, a final concentration of 900 nM of the primers and 200 nM of the corresponding probes were added into two separate reaction tubes, one for PLP and one for SRY detection. For the multiplex PCR, both set of primers were added into the same reaction tube at a final concentration of 400 nM each, and 200 nM of each probe (labeled with different fluorophores) were added into the same tube. An iCycler iQ real-time PCR detection system (Bio-Rad Laboratories, Inc., Hercules, CA, USA) was used for real-time PCR. The amplification conditions consisted of initial denaturation and enzyme activation at 95°C for 10 min, followed by 40 cycles of 95°C for 15 s and 60°C for 60 s. The fluorescent signals were measured at the end of each amplification cycle. Analysis of the signals for determination of the threshold cycles (Ct) was done using CFX manager software version 1.1 (Bio-Rad Laboratories, Inc., USA).
Semen samples and DNA extraction
The samples used in this study comprised two X-sorted semen samples, one Y-sorted semen sample, and three unsorted semen samples. The sorted semen samples containing spermatozoa of one sex at greater than 90% purity was purchased from Accelerated Genetics, Inc., Baraboo, WI, USA. Three unsorted semen were purchased from the Dairy Farming Promotion Organization of Thailand.
Each frozen semen sample was thawed and centrifuged at 500×g for 10 min to collect the sperm pellet. The pellet was then washed with 500 μL phosphate buffered saline, centrifuged at 500×g for 5 min, and the supernatant was discarded. Then, 10 μL of 1 M dithiothreitol, 10 μL of 1 M proteinase K and 100 μL of 5% w/v chelex 100 resin (Bio-Rad Laboratories, Inc., USA) were added to the sperm pellet. All reagents were mixed and incubated overnight at 56°C. The cell debris was removed by centrifugation at 1,000×g for 5 min. Then, the crude DNA extract was further purified using a Wizard genomic DNA purification Kit (Promega, USA) following the manufacturer’s protocol. The DNA concentration was measured spectrophotometrically at 260 nm.
Determination of sperm sex ratio by simplex real-time polymerase chain reaction
For the simplex real-time PCR, the fluorescent signals from both X and Y amplifications were measured from the same channel (FAM). A single threshold line for determination of the Ct can be defined for both X and Y amplification. The threshold was set approximately in the middle of the exponential increase of the signal from positive samples. A standard curve showing relationship between Ct and copy number of the template for each sex-specific DNA was constructed using the reference plasmid DNA. Each sex-specific DNA from the test samples was then quantified using the corresponding standard curves, i.e., X from X-standard curve and Y from Y-standard curve. The percentage of X content (%X) and Y content (%Y) were estimated using the assumption that the sum of X and Y contents in each sample equals 100%.
Determination of sperm sex ratio by multiplex real-time polymerase chain reaction
For the multiplex real-time PCR, two fluorescent signals were generated from each reaction tube, namely the Orange 560 signal from X amplification and the FAM signal from Y amplification. A standard curve was constructed from each fluorescent channel and a threshold line of each fluorescent channel was defined using the same criteria as the simplex PCR. Quantification of X and Y contents as well as the sperm sex ratio were calculated in a similar fashion to those from the simplex PCR.
Validation of the multiplex real-time polymerase chain reaction assay
Sperm sex ratios presented as percentage of X chromosome-bearing spermatozoa (%X) obtained from the multiplex real-time PCR were compared with those obtained from the simplex real-time PCR. A total of nine samples used in this experiment included three samples of known %X reference plasmids (X10%, X50%, and X90%), two X-sorted semen samples, one Y-sorted semen sample, and three unsorted semen samples. Each sample was tested in triplicate independent PCRs. For each PCR, the samples were tested in duplicate reaction tubes and the average values were used to calculate the mean, standard deviation (SD), and 95% confidence interval (95% CI) of the triplicate experiments. Validity of the multiplex real-time PCR assay was assessed by comparing the expected values (from the reference plasmids) to those obtained from the measurement. The assay was considered valid when all expected values fell within 95% CI of the measured values. Agreement between the sex ratio estimated from the multiplex real-time PCR and the simplex real-time PCR assay was assessed using a two-tailed paired t-test with significance level of p<0.05. The equality of variances between the methods was tested using Levene’s test.
RESULTS
Standard curves and amplification efficiency
Standard curves showing relationship between the amount of template (copy number, logC) and Ct for each sex-related DNA from the simplex and multiplex assays are shown in Figure 1 and 2, respectively. All curves fitted to linear regression models with correlation coefficient (r)>0.99. For the simplex PCR, the general linear equations obtained from the three runs can be represented as: Ct_X = 35.40–3.34 logC for X amplification and Ct_Y = 36.12–3.39 logC for Y amplification. For the multiplex PCR, the equations for X and Y were: Ct_X = 40.56.16–3.39 logC and Ct_Y = 41.01–3.35 logC, respectively.
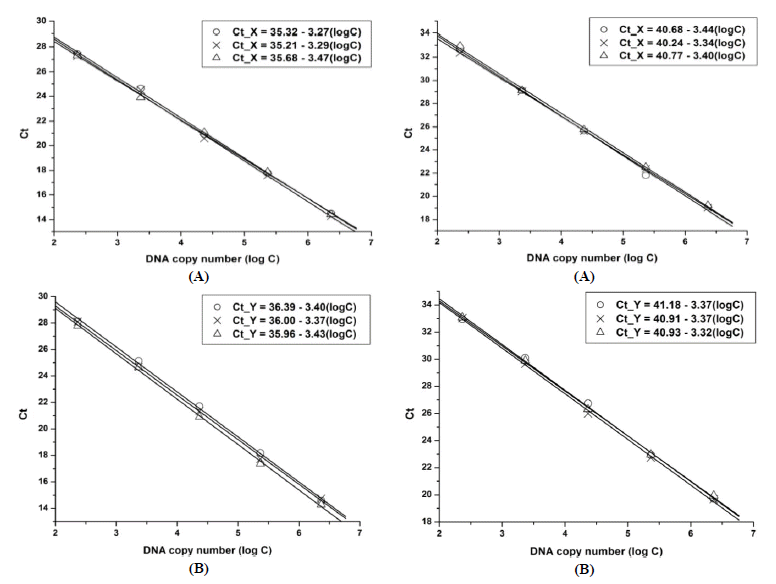
Standard curves and linear equations obtained from the simplex real-time polymerase chain reaction assay showing relationship between threshold cycle (Ct) and DNA copy number (logC) from triplicate experiments. (A) X chromosome-specific amplification, (B) Y chromosome-specific amplification.

Standard curves and linear equations obtained from the multiplex real-time polymerase chain reaction assay showing relationship between threshold cycle (Ct) and DNA copy number (logC) from triplicate experiments. (A) X chromosome-specific amplification, (B) Y chromosome-specific amplification.
The mean and SD of the slope from the simplex PCR assay were −3.34±0.11 and −3.39±0.03 which corresponded to the PCR amplification efficiency of 99% and 97% for X and Y amplification respectively; and from the multiplex PCR, the slopes were −3.39±0.05 and −3.35±0.03 which corresponded to efficiencies of 97% and 99% for X and Y amplification, respectively.
Estimation of %X in samples and comparison between the simplex and multiplex real-time polymerase chain reaction assay
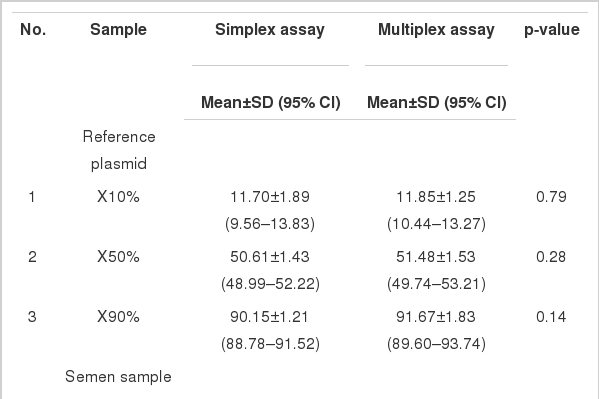
The simplex PCR and multiplex PCR assays were used to measure ratios of the sex-specific sequences in the reference plasmids and semen samples. The results expressed as %X in the samples are shown in Table 2. The expected values from the reference plasmid samples fell within the 95% CI of the measured values obtained from the assays in all test ranges indicating validity of the assays for accurately assessing the sex ratio of the semen samples. Statistical analysis showed that the variances and mean of the sex ratio estimates obtained from the simplex and multiplex assays were not significantly different (p>0.05) (Table 2).
DISCUSSION
In this study, we developed a multiplex real-time PCR technique as an alternative method for determination of sperm sex ratio in bovine semen samples. Analysis of the sperm sex ratio is important for selective reproduction of livestock by artificial insemination. This rapid and cost effective assay can also facilitate development of new sperm sex-sorting technologies. The assay simultaneously detects both X- and Y-specific determinants in a single reaction tube. Quantification of each sex-specific DNA was done by the use of differently labeled Taqman probe, and comparing the Ct obtained from the sample to that obtained from the corresponding standard curve. The standard curve in this multiplex assay was also done in multiplex format, i.e., both X- and Y- specific recombinant plasmids were mixed at the ratio of 1:1 and a series of ten-fold dilutions of the known-quantity reference DNA was used to construct the standard curve for each sex-specific DNA. In addition, for reference DNA, genomic DNA may be used to construct the standard curve (Yun et al., 2006). Although the primers used in this multiplex assay were only half of the concentration used in the simplex assay, the amplification efficiencies were the same, i.e., they fell in the range between 97% to 99% efficiency for both assays. The multiplex assay was also proven to be as effective as the simplex assay in determination of the sperm sex ratio in both known-ratio reference plasmids and actual semen samples. For the sorted semen samples, the assay showed that they contained greater than 90% of the declared sex-specific DNA, which was in agreement with the semen specification guaranteed from the manufacturer (>90% purity). For the unsorted semen in which the ratio of 1:1 was expected, the multiplex real-time PCR assay yielded approximately 50% of each sex-specific DNA, which is in agreement with the expected value. The multiplex real-time PCR technique as developed in this study yielded accurate assessment of the sperm sex ratio in both sorted and unsorted semen. Although the accurate assessment of the ratio can be obtained with other techniques such as flow cytometry analysis (Garner et al., 2013), individual sperm-PCR (Colley et al., 2008; Wang et al., 2011), simplex real-time PCR (Parati et al., 2006), the multiplex method as reported in this study is simpler, faster, and more cost-effective to perform with only a single reaction tube required for each semen sample. In conclusion, the multiplex real-time PCR assay could be used as an alternative method to identify the sex ratio of the sperm in bovine semen.
ACKNOWLEDGMENTS
This work was funded by National Science and Technology Development Agency (NSTDA), Thailand. The first author was supported by Thailand Graduate Institute of Science and Technology scholarship (TG-22-10-51-005D) granted from NSTDA. We thank Dr. Philip Shaw for manuscript proofreading.