Genetic Diversity of Myanmar and Indonesia Native Chickens Together with Two Jungle Fowl Species by Using 102 Indels Polymorphisms
Article information
Abstract
The efficiency of insertion and/or deletion (indels) polymorphisms as genetic markers was evaluated by genotyping 102 indels loci in native chicken populations from Myanmar and Indonesia as well as Red jungle fowls and Green jungle fowls from Java Island. Out of the 102 indel markers, 97 were polymorphic. The average observed and expected heterozygosities were 0.206 to 0.268 and 0.229 to 0.284 in native chicken populations and 0.003 to 0.101 and 0.012 to 0.078 in jungle fowl populations. The coefficients of genetic differentiation (Gst) of the native chicken populations from Myanmar and Indonesia were 0.041 and 0.098 respectively. The genetic variability is higher among native chicken populations than jungle fowl populations. The high Gst value was found between native chicken populations and jungle fowl populations. Neighbor-joining tree using genetic distance revealed that the native chickens from two countries were genetically close to each other and remote from Red and Green jungle fowls of Java Island.
INTRODUCTION
Nowadays, it is increasingly recognized that insertion and deletion (indels) polymorphisms are an important source of genetic as well as phenotypic diversity (Brandström and Ellegren, 2007). Indels can be genotyped with simple procedures based on size separation as compared to complex and heterogenous mutation patterns of microsatellites and without the necessity of applying special equipment for high-throughput genotyping that is relatively costly for small or medium scale studies using microsatellite and SNP markers (Väli et al., 2008). Moreover, indels markers have many genetic advantages for analytical use: they are widely spread throughout the genome, all of the polymorphisms derive from a single mutation event and they have reduced mutation rates (Natalle et al., 2010). It was reported that the draft sequence of the chicken genome was composed of approximately 1.2 Gb which included 447,388 indels polymorphisms (2011, November) in the National Center for Biotechnology Information (NCBI) Entrez SNP database (www.ncbi.nih.gov/sites/entrez?db=snp). This clearly indicated that indels could form a very common class of polymorphism over the chicken genome and might be important genetic markers. The present study focused on indels polymorphisms as genetic markers for studying genetic diversity in chicken population.
Chickens are good models for studying the genetic basis of phenotypic traits because of the extensive diversity among domestic chickens (Wong et al., 2004). There are more than 18 million stocks of chickens being raised in the world (FAO Statistics, 2009). Native chickens, as the most adaptable and geographically widespread livestock species, form an integral part of the Myanmar and Indonesian ecosystem. Native chickens possess unique adaptive traits that permit them to survive and reproduce under harsh climatic, nutritional and management conditions typically associated with low input-output production systems (Mwacharo et al., 2006).
Myanmar is the largest country on main land South East Asia with a total land area of 676,577 sq Km. Myanmar possesses tropical and sub tropical climates and rich with diverse species of animal and wild relatives of livestock. Latest estimates indicate that there are 105 million domesticated birds raised in Myanmar, out of which 94 million are chicken, mostly 78.7 million birds (84%) kept under backyard production systems (Burgos et al., 2009).
The land area of Indonesia covers 1,919,445 sq Km, spreading from Aceh Province in the north west of Sumatra to the Western part of Papua. It is a large tropical island in South East Asia, which shelters a large number and variety of wild as well as domesticated animals. According to livestock statistic from 2007, Indonesia has an estimated standing population of 620 million chickens which include 317 million native/village chicken (51%) kept by rural farmers (Ministry of Agriculture -2007 cited by Bambang et al., 2009).
Although these native chickens are not available for commercial use, they are raised as dual- purpose for meat and eggs, providing cash income in times of need. It is interesting to examine the genetic variability among native chicken from two south East Asia countries using molecular markers. Therefore, the present study was conducted to evaluate the indels polymorphisms as genetic markers to characterize the genetic variability of the native chicken populations from Myanmar and Indonesia and to determine the genetic relationship among Myanmar and Indonesia native chicken populations and two jungle fowl species.
MATERIALS AND METHODS
In the present study, we used 11 native chicken populations and two jungle fowl species. In Myanmar native chickens, we analyzed three populations from Yangon (Yan) and Pegu (Peg) of lower Myanmar and from Mandalay (Man) of central Myanmar. In Indonesian native chicken, we analyzed four populations of local chicken and four populations of local varieties: The local chicken populations were collected from Semarang (Sem), Kendal (Ken), Yogyakarta region (Yog) of the central Java, and Karawang region (Kar) of west Java. The populations of local varieties were Ayam Kedu (AK) from Temanggun (AK-Tem) and Solo (AK-Sol) of the central Java, Black Kedu (BK) from Semarang (BK-Sem) and Ayam Alab (AA) from Temanggun (AA-Tem). Two jungle fowls species from Java, Red jungle fowl (Gallus gallus bankiva, RJF) and Green jungle fowl (Gallus varius, GJF) were also examined. Blood samples were obtained from Yan (n = 27), Peg (n = 13), Man (n = 40), Sem (n = 24), Ken (n = 40), Yog (n = 57), Kar (n = 15), AK-Tem (n = 10), AK-Sol (n = 19), BK-Sem (n = 23), AA-Tem (n = 20), RJF (n = 3), GJF (n = 6) respectively.
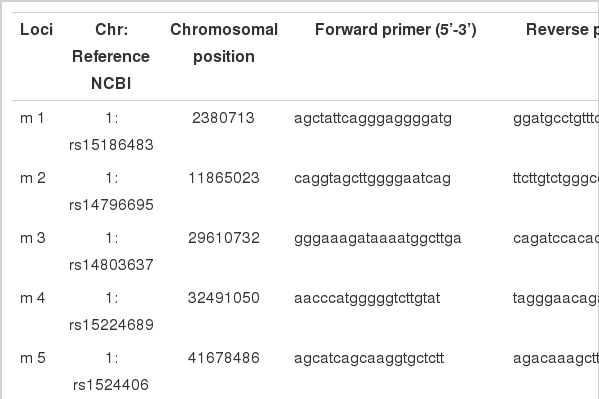
Genomic DNA was extracted from blood samples using standard phenol-chloroform extraction protocols (Sambrook, 1989). Suitable indels markers were selected by using the NCBI Entrez SNP database. The primer sequences used in this study were listed in Table 1.
In primer design, indels sequences of 20 to 30 bp along the every 10 Mb of chromosome length were selected. The PCR amplifications were performed in a 10 μl reaction volume, which included 6.15 μl of distilled water, 1 μl of 10× reaction buffer, 0.8 μl of dNTP (2.5 mM), 0.25 units of Ex Taq™ (Takara Bio Inc, Otsu, Japan), 10 ng of template DNA, 25 μM each of forward and reverse primers. The PCR reaction was performed with an initial denaturation for 2 min at 94°C followed by 28 to 32 cycles of denaturation for 30 s at 94°C, annealing for 30 s at 48 to 65°C and extension for 30 s at 72°C and final extension for 7 min at 72°C in Takara PCR thermal cycler - TP650 (Takara Bio Inc). The PCR products and a 100 bp DNA ladder (Sib Enzyme Ltd, Russia) were electrophoresed on 1 to 2% agarose gel electrophoresis in 1×TBE. The indel polymorphisms were identified through a Toyobo FAS III UV-Transilluminator (Toyobo, Osaka, Japan) after ethidium bromide staining for 15 to 30 min. Genotypes of indel polymorphisms were determined by size difference between the PCR fragments.
Statistical analysis
We calculated allele frequencies from genotyping data by direct counting. The proportion of polymorphic loci (Ppoly) was calculated as the ratio of polymorphic loci to the total number of loci analyzed. Chi square (χ2) approximation was used to test Hardy-Weinberg Equilibrium (HWE) at each locus (Weir, 1996). To estimate the genetic variability, we calculated the average observed heterozygosity
RESULTS
Indels polymorphisms
The mean (±SE) Ppoly value of each population is listed in Table 2. Out of 102 indels markers, 97 markers (Ppoly = 95%) were polymorphic in native chicken populations, all of which were in HWE (Table 1). The Ppoly was higher in Indonesian native chicken populations (Ppoly = 94%) than Myanmar native chicken populations (Ppoly = 92%) (Table 2). One marker (m 66) was monomorphic only in Indonesian native chicken populations whereas three markers (m 11, m 33 and m 92) were monomorphic only in Myanmar native chicken populations.
In two jungle fowl species, RJF showed more polymorphic loci (22 markers, Ppoly = 21%) than GJF (4 markers, Ppoly = 3.9%) (Table 2).
Genetic variability
The
Genetic distance and phylogenic analysis
The Ds distances are shown in Table 4. The Ds distances between the native chicken populations ranged from 0.034 to 0.078 within Myanmar, from 0.009 to 0.147 within Indonesia and from 0.063 to 0.200 between the two countries.
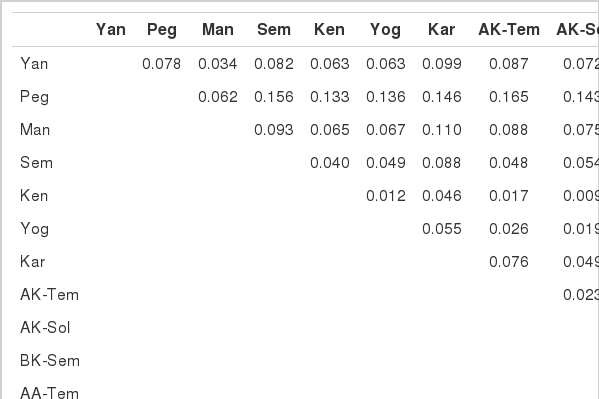
Pair wise genetic distance matrix between 13 populations of native chickens and two jungle fowl species
The smallest (0.063) was observed between the Ken and Yan population and between the Yan and Yog populations. The largest (0.200) was obtained from between BK-Sem and Peg populations. The genetic distances between native chicken populations and the two jungle fowl species ranged from 0.247 to 0.405.
The NJ tree constructed from the Ds distances between the 13 populations gave two major clades as shown in Figure 1. The first clade was composed of Myanmar and Indonesian native chicken populations. The second clade was composed of RJF and GJF, which was located outside the first clade.

Neighbor-joining (NJ) tree constructed by genetic distance matrix from 13 populations of native chickens and two jungle fowl species. Yan = Yangon; Peg = Pegu; Man = Mandalay; Sem = Semarang; Ken = Kendal; Yog = Yogyakarta; Kar = Karawang; AK-Tem = Ayam Kedu from Temmanggun; AK-Sol = Ayam Kedu from Solo; BK-Sem = Black Kedu from Semarang; AA-Tem = Ayam Alab from Temmanggun; RJF = Red jungle fowl; GJF = Green jungle fowl.
DISCUSSION
Indels polymorphisms
The results revealed both of the native chicken populations showed polymorphisms in most of the indels loci (greater than 90%). It is larger than the previous study of Väli et al. (2008) and their study stated that 81 and 76 out of 94 indels markers (86.2% and 80.9%) could be validated as polymorphic loci in dogs (n = 7) and wolves (n = 18). Jungle fowl populations especially in GJF showed much fewer polymorphic indels loci (4%) than native chicken populations. These results suggest that the indels polymorphisms examined here have been acquired in the chickens after the separation of ancestral species of the GJF and chickens.
Genetic variability
In this study, the
In the present study, the heterozygosities of native chickens were higher than the heterozygosities of ancestral species. This finding was similar to the earlier report of Väli et al. (2008). They reported that the values of
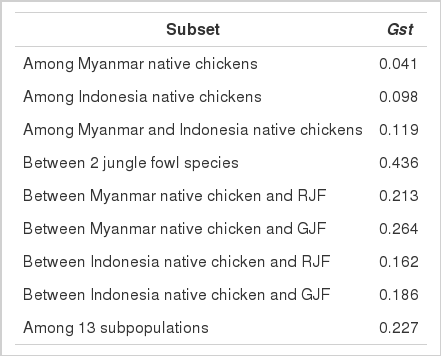
The Gst values of Myanmar and Indonesian native chicken populations were 0.041 and 0.098, indicating that the degree of genetic differentiation is higher in Indonesian native chicken than Myanmar native chicken. This might be related to the sample population of Indonesian native chicken, which contained local chicken varieties like BK, AA, and AK, and there may be some degree of genetic differentiation between them.
The Gst of present study was higher than those of Myanmar and Indonesian native chicken (0.024 and 0.020) estimated in the genetic constitution of four calpain gene polymorphisms (Okumura et al., 2006). However Gst of Myanmar native chicken was close to the range of 0.001 to 0.039 (Kinoshita et al., 2004) estimated from egg white protein polymorphism of local populations in Asian countries (Myanmar, Indonesia, China, Nepal, Vietnam and Laos). The Gst of Indonesian native chicken was close to blood protein polymorphisms found in four chicken breeds from Yunnan Province of China (Gst = 0.075, Okamoto et al., 2003) and Nepal (Gst = 0.093, Maeda et al., 1992).
The Gst between Myanmar and Indonesian native chickens was 0.119, indicating that the genetic differentiation between them was not large. Therefore, Myanmar and Indonesia native chickens can be regarded as genetically close populations. The Gst values between Myanmar native chicken populations and two jungle fowl species (0.213 to RJF and 0.264 to GJF) are greater than between Indonesian native chicken populations and two jungle fowl species (0.162 to RJF and 0.186 to GJF). It may be due to the fact that the small Gst was obtained between populations in a close geographic area. However, highest Gst (0.436) was observed between RJF and GJF from Java Island.
The Gst value in this study may be higher than the value of other studies. According to the review of Theresa et al. (2002), a wide range of Gst values usually results from uneven allele frequency distributions across populations at some loci. In the present study, the major allele was the same in all populations at 87 indels loci. Whereas uneven allele frequency distributions were observed in the remaining 15 loci: the major allele in most populations was minor in some populations, which may contribute to high Gst value.
Genetic distance and phylogenic analysis
The low genetic distance observed among native chicken populations (0.009 to 0.200), may be reflecting the fact that genetically these populations are not greatly isolated from each other. In addition, the average genetic distances among native chicken populations (0.088) observed in present study is close to the findings of Yamashita et al. (1994) by DNA fingerprinting analysis among the stock of domestic fowls (0.104). The larger genetic distances were found between native chicken populations and two jungle fowl species from Java Island.
The topology of the NJ tree showed that Myanmar native chickens and Indonesian native chickens form a respective cluster in one clade whereas RJF and GJF from Java Island formed another clade, suggesting that native chickens are genetically closely related to each other and remote from jungle fowls of Java Island.
In the previous studies of Yamashita et al. (1994) and Okumura et al. (2006), the GJF was located far away from native chicken populations and it is consistent with the hypothesis that domestication of the chicken may have started from RJF. However, RJF from Java Island comprises a different clade that was far away from native chicken populations, which agreed with the un-rooted neighbor joining (NJ) population tree of Niu et al. (2002). In their NJ tree, domestic fowls belonged to the same cluster as G. g. gallus and G. g. spadiceus in Thailand and its adjacent areas, whereas G. g. bankiva from Indonesian Islands formed a separate cluster. Furthermore Akishinonomiya et al. (1994) stated that the domestic fowl from Indonesian Island had large genetic differences compared with G.g.bankiva from the same region.
The present study examined the genetic characteristics of the Myanmar and Indonesia native chickens and two jungle fowl species from Java Island by using indels polymorphisms as genetic marker. The genetic variability is higher among native chicken populations and lower in the two jungle fowl species. The high genetic differentiation occurred between native chicken populations and two jungle fowl species from Java Island. The native chickens from two countries were genetically close to each other and remote from jungle fowls of Java Island. Although the indels markers showed low heterozygosity compared to microsatellite markers, they can demonstrate close genetic variability and phylogenic topology to other studies cited above. Therefore, indels polymorphisms are efficient for studying genetic diversity of populations.